Deep learning improves the quality of XRF microscopy by one order of magnitude
A self-supervised deep learning method developed at ID16A exploits independent noise across detector channels, recovering signal better than channel averaging and matching acquisitions with five to 30 times longer dwell time.
Share
The challenge
X-ray fluorescence microscopy (XRF) maps nanoscale chemical composition of samples. Achieving high spatial resolution and sensitivity to trace elements, however, often requires long acquisition times and substantial radiation doses. At synchrotrons, these constraints limit throughput and compromise correlative studies where excessive dose from one technique precludes subsequent measurements, especially in radiation-sensitive biological specimens. Samples with a large elemental dynamic range present challenges when dominant elements saturate the detector, pushing trace elements toward the noise floor.
Multi-channel detectors partially address this by collecting more photons per unit time through increased solid angle, but detector geometry, cost, and space constraints restrict how many channels can be integrated. Extending dwell times or increasing flux degrades sample integrity, creating tension between data quality and preservation. Classical denoising methods like median filtering and total variation regularisation suppress noise but compromise spatial resolution or introduce artefacts that distort quantitative analysis. The central challenge is extracting more information from existing data without additional dose or acquisition time.
The development
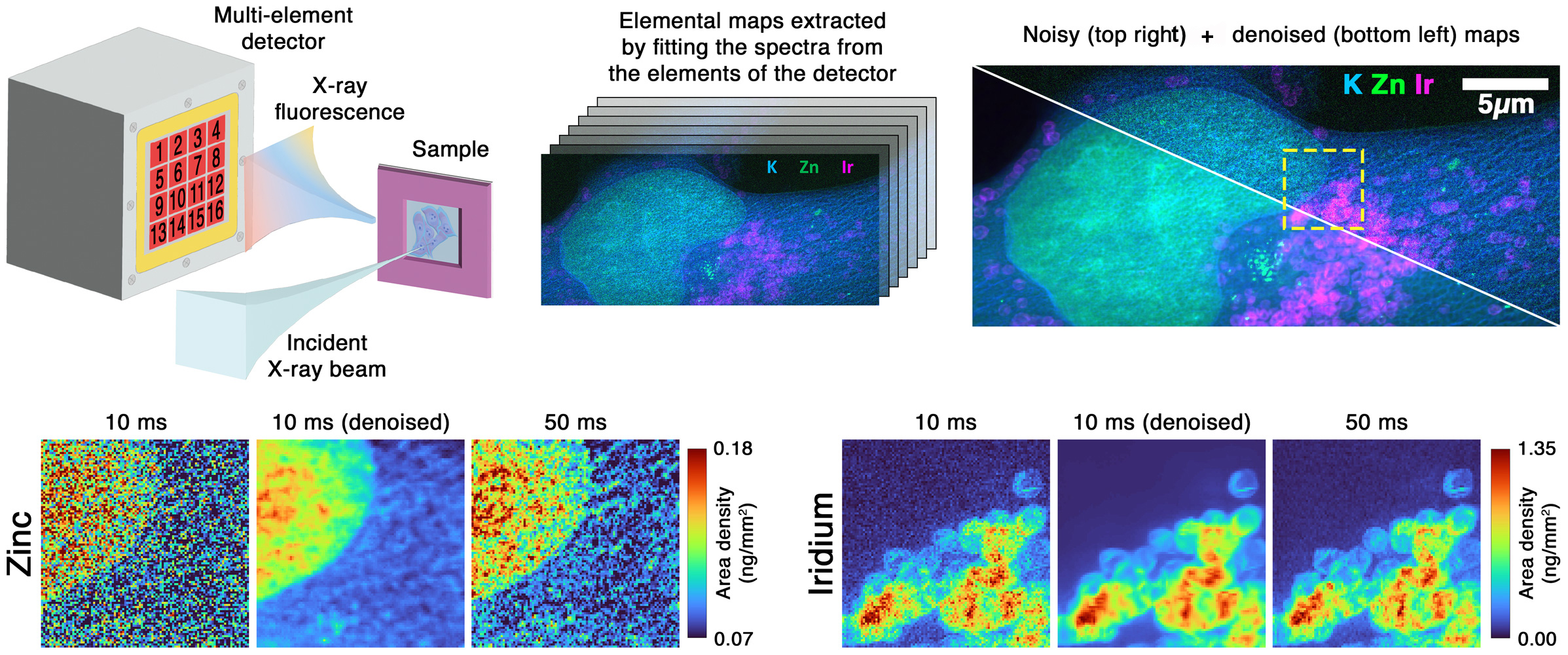
A self-supervised denoising method was developed at beamline ID16A that treats each elemental map from individual detector channels as an independent noisy observation of the same sample region (Figure 1). For instance, the 16-channel detector produces 16 separate zinc distribution maps that would normally be weighted and averaged into a single image. Instead, the approach adapts the Noise2Noise framework to train a convolutional neural network directly on pairs of these independent measurements, without requiring noise-free reference data. This works because detector channels capture statistically uncorrelated noise while viewing identical sample structures, reducing the problem to estimating the true value at each pixel given multiple independent observations.
Click figure to enlarge
Fig. 1: Schematic of the multi-element XRF detector setup at ID16A and denoising results. Comparison shows noisy data (10 ms dwell time), denoised output, and reference acquisition (50 ms dwell time).
Experiments on a Siemens star resolution target and cryogenically fixed human cancer cells showed that the method recovers signal quality matching references acquired with approximately five to 30 times higher flux or longer dwell time, depending on the sample. Denoised results surpass simple channel averaging by recovering finer structural details while preserving quantitative accuracy. The method works even with only two detector channels positioned on opposite sides of the sample, a configuration common at many XRF instruments. This conceptual shift moves denoising from a purely signal-averaging strategy to a data-driven reconstruction approach that exploits detector redundancy.
The impact
This represents one of the first application of machine learning-based denoising to X-ray fluorescence imaging. By converting detector multiplicity into a computational advantage rather than simply averaging signals, the technique allows dose-sensitive measurements and higher experimental throughput.
The approach benefits trace element mapping in radiation-sensitive biological specimens, materials characterisation requiring large-area surveys, and time-resolved studies where acquisition speed matters. Users at ID16A can maintain current data quality while scanning approximately five to 30 times faster or improve detection limits at current acquisition parameters. This enables users to obtain publishable-quality XRF maps in significantly shorter beamtime, expanding experimental possibilities for dose-sensitive and high-throughput studies. The method can be applied retrospectively to archived data, enabling previously noise-limited datasets to be reprocessed with substantially improved quality.
The method requires no hardware modifications and could, in principle, be extended to other scanning techniques that record parallel, statistically independent views of the same sample. To facilitate adoption, training tutorials and documented code are available in the SetkaFluo GitHub repository, allowing users to apply the approach to their own data and adapt it to other multi-channel imaging modalities.
Principal publication
Self-Supervised Deep-Learning Denoising for X-ray Fluorescence Microscopy with Multi-Element Detectors, R. Shishkov et al., Anal. Chem. (2026); https://doi.org/10.1021/acs.analchem.5c05552