- Home
- News
- Spotlight on Science
- Towards atomic resolution...
Towards atomic resolution of the plant photosystem I supercomplex
14-05-2007
Thanks to diffraction data collected at the ESRF, the structure of the intact plant photosystem I supercomplex from pea plants, containing 17 protein subunits and 178 cofactors, has been revealed at 3.4 Å resolution thus providing near atomic details of this extremely complex membrane supercomplex.
Share
All higher organisms on earth receive energy directly or indirectly from oxygenic photosynthesis performed by plants, green algae and cyanobacteria. Plant photosystem I (PSI) is arguably one of the most intricate membrane supercomplexes in Nature. It consists of a reaction centre complex (RC) and a light-harvesting complex (LHCI). Each of these sub-complexes is composed of many protein subunits, most of which transverse the chloroplast membrane several times. Plant PSI has about 200 pigments and every captured photon is trapped and results in electron translocation. PSI thus exhibits a remarkable quantum yield value of nearly 1, and is the most efficient nano-photoelectric machine in Nature.
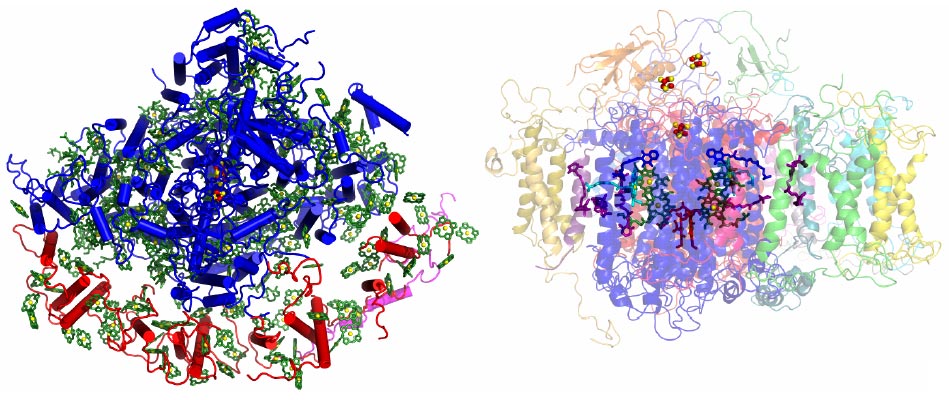
A structural study of PSI supercomplex was initiated seven years ago. In 2003 the structure of plant PSI was solved at 4.4 Å resolution, revealing a β-carbon model of 16 proteins, 45 transmembrane helices, 2 phylloquinones, 3 FeS clusters and 167 chlorophylls [1]. Continuous and progressive improvement of crystal quality, during which it was found that a dehydration procedure significantly altered the diffraction properties of the crystals, enabled the recent determination of the crystal structure of PSI at 3.4 Å resolution [2]. This provides a picture at near atomic detail of 16 of the 17 protein subunits with an additional subunit (PsaN) being identified for the first time on the luminal side of the supercomplex. The positions of 3038 out of 3443 predicted amino acids were assigned as were those of 168 chlorophylls (65 revealing the orientation of the Qx and Qy transition dipolar moments), 2 phyloquinones, 3 FeS clusters and 5 carotenoids (Figure 1).
One of the most important factors that allowed the higher resolution data to be obtained was the availability of high intensity ESRF MX end-stations ID29 and ID23-2. The latter microcrystallography beamline was especially effective, since the better diffracting crystals were of much smaller dimensions than those previously obtained. Although the intensity of the X-ray beam meant that, due to radiation damage, no more than 10 diffraction images could be collected from any one part of a crystal, the small beam available on ID23-2 allowed the collection of data from several areas of the best crystals.
The structural information on proteins, co-factors and the interactions between them described in this work provides a significant step towards our understanding of how the unprecedented high quantum-yield of plant PSI in light capturing and electron transfer is achieved [2].
References
[1] A. Ben Shem, F. Frolow & N. Nelson, Crystal structure of plant Photosystem I, Nature, 426, 630-5 (2003).
[2] A. Amunts, O. Drory & N. Nelson, The structure of a plant Photosystem I supercomplex at 3.4 Å resolution, Nature, 447, 58-63 (2007).
Authors
A. Amunts, O. Drory and N. Nelson.
Department of Biochemistry, Tel Aviv University (Israel)