- Home
- News
- Spotlight on Science
- Studying the biosynthesis...
Studying the biosynthesis of vitamin B6
01-03-2022
To synthesise vitamin B6 from carbohydrates provides a formidable challenge: in plants, a single enzyme performs over 10 catalytic steps. Mass spectrometry was combined with UV-vis absorption spectroscopy and X-ray crystallography carried out at beamline ID23-1 to investigate the mechanisms involved.
Share
Trapping reaction intermediates in enzymes can provide important insights into natural product chemistry. The enzyme PLP synthase [1] uses two simple sugars and an amino acid as building blocks for vitamin B6 (pyridoxal-phosphate, PLP). Arguably the most intriguing step is incorporation of ammonia into the growing vitamin. The result of this reaction is an intermediate that is held between two protein amino acids by covalent bonds.
A big step towards working out the fine detail of the intricate mechanism involved in this step has now been reported. Three key experiments made this advance possible: characterisation of a novel reaction intermediate by optical spectroscopic methods using the icOS laboratory, structure determination of the enzyme-intermediate complex using the macromolecular crystallography (MX) beamline ID23-1, both at ESRF, and mass spectrometry to derive the chemical nature of the intermediate. Combining these three approaches elucidates the stereoselectivity of catalytic steps leading to the vitamin’s biosynthesis.
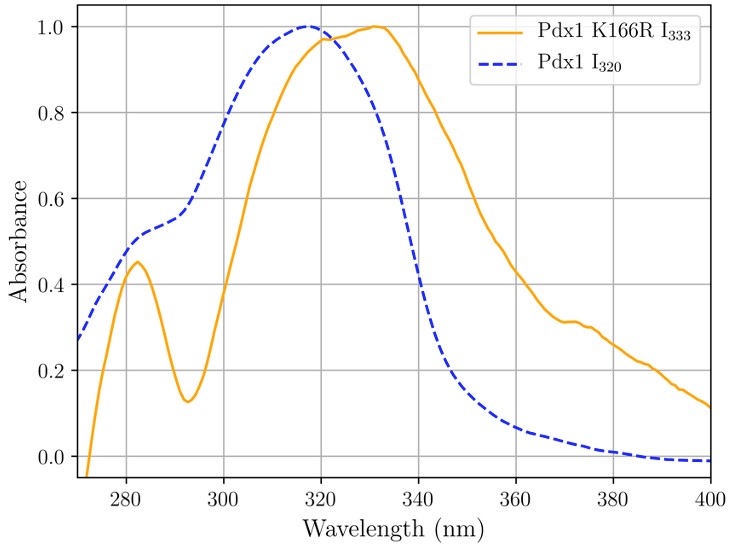
Fig. 1: UV-vis absorption spectroscopic signatures of enzyme-intermediates in PLP biosynthesis detected in crystallo: the signatures for the chromophores I320 (blue dashed line) and I333 (orange line) are shown.
The enzyme PLP synthase first binds the carbohydrate ribose 5-phosphate in a covalent bond with a protein lysine residue. When ammonia is added, a second covalent linkage to another lysine results in formation of the double imine chromophore I320 [2,3], named after its characteristic absorbance in the 320 nm region, shown as a blue dashed line in Figure 1. Modification of the enzyme by exchange of this second lysine allowed characterisation of the novel intermediate that immediately precedes I320 formation. This new species has a characteristic absorbance at 333 nm, shown as an orange line in Figure 1.
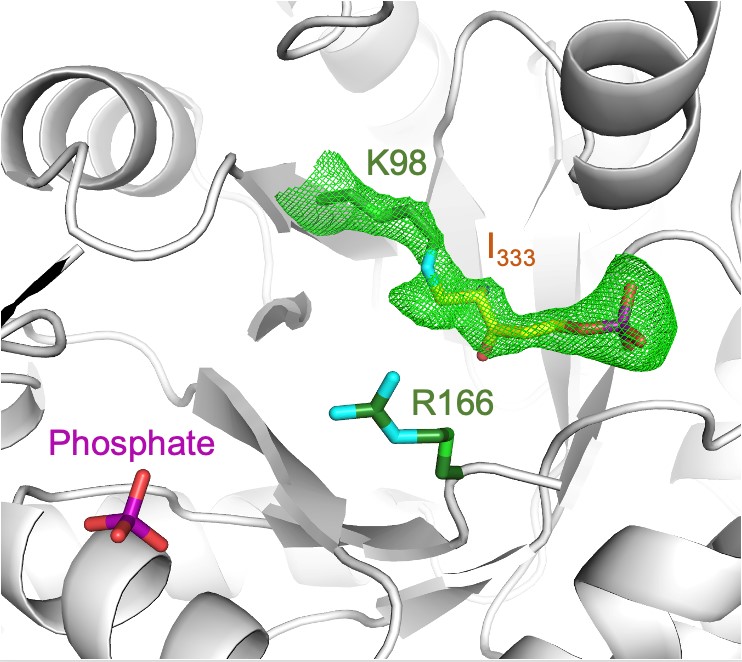
Fig. 2: The structure of the ribose tailoring site in PLP synthase (PDB:7NHE). The enzyme lysine required for I320 formation was replaced with an arginine residue in this experiment and is shown as green sticks (R166). The chromophoric intermediate I333 is seen covalently linked to the side chain of a lysine (K98). The mesh shows the simulated annealing electron density contoured at 2.5σ (calculated with Lys98–I333 residues removed). A phosphate ion is seen in the site where PLP would form.
The ability to perform these experiments on protein crystals at icOS meant that pre-characterised crystals could be selected for diffraction data collection. Indeed, the spectroscopic signature proved the integrity of the specific intermediate in protein crystals from which the enzyme-intermediate structure was determined using MX carried out at ESRF beamline ID23-1. This experiment produces an electron density map, shown as a green mesh in Figure 2. However, interpretation of this map is not unambiguous, and various tautomeric structures, in combination with determination of the sum formula of I333 by mass spectrometry, were tested to produce the final structural model. Knowledge of the detailed structure of I333 now allows a proposal as to how I320 is formed, explaining this central and rate limiting step in PLP biosynthesis.
Principal publication and authors
Trapping and structural characterisation of a covalent intermediate in vitamin B6 biosynthesis catalysed by the Pdx1 PLP synthase, M.J. Rodrigues (a, b), N. Giri (c), A. Royant (d,e), Y. Zhang (f), R. Bolton (a,b), G. Evans (b), S.E. Ealick (f), T. Begley (c), I. Tews (a), RSC Chem. Biol. 3, 227-230 (2022), https://doi.org/10.1039/d1cb00160d
(a) University of Southampton, Southampton (UK)
(b) Diamond Light Source, Didcot (UK)
(c) Texas A&M University, College Station (USA)
(d) Univ. Grenoble Alpes, CNRS, CEA, Institut de Biologie Structurale, Grenoble (France)
(e) ESRF
(f) Cornell University, Ithaca (USA)
Structures deposited with the PDB: 7NHE, 7NHF
References
[1] M. Strohmeier et al., Proc. Natl. Acad. Sci. U.S.A. 103, 19284-19289 (2006).
[2] G.C. Robinson et al., Proc. Natl. Acad. Sci. U.S.A. 113, E5821-E5829 (2016).
[3] M.J. Rodrigues et al., Nat. Chem. Biol. 13, 290-294 (2017).



