- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2013
- Soft condensed matter
- A viral protein nanocage assembly probed in vitro
A viral protein nanocage assembly probed in vitro
Noroviruses are the first cause of non- bacterial gastroenteritis in humans, as well as in animals, causing about 250 million cases of gastrointestinal distress and over 200,000 deaths each year. They are made up of a protein nanocage called the capsid (~20-40 nm in diameter), enclosing the genetic material, or genome, in the form of RNA. For noroviruses and many other viruses, the capsid consists of 180 copies of a single structural protein arrayed in an icosahedron, i.e. a solid with 20 triangular faces and 12 vertices. Remarkably, the norovirus capsid proteins can self-assemble reversibly in vitro, without the assistance of the genome or any cellular components, solely by the interplay of solution pH and ionic strength [1]. The molecular mechanisms of such efficient and precise capsid protein self-assembly still remain unknown to date and the nature of the intermediate species is much debated.
The self-assembly kinetics of capsid proteins is a multiscale process that requires heterogeneous molecular species with nanometre sizes to be probed over a time scale ranging from milliseconds to hours. The classical techniques used to probe self-assembly kinetics such as time-course light scattering provide temporal traces from which characteristic time scales can be inferred, but no structural information can be extracted. In contrast, time-resolved small-angle X-ray scattering (TR-SAXS) permits a determination of the shape and the organisation of particles with a spatial scale in the range of a few angstroms to thousands of angstroms, and with time resolution below 100 ms [2].
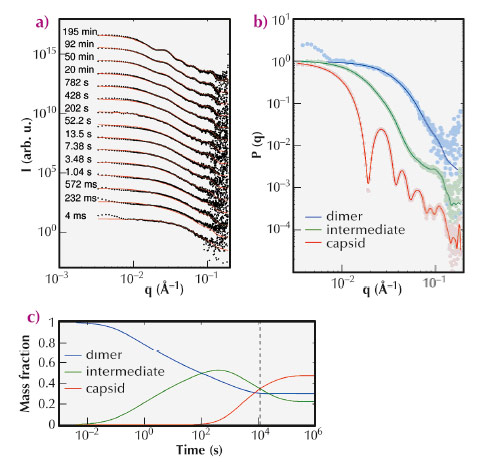
We have thereby investigated the self-assembly of capsid proteins derived from a norovirus, by exploiting the exceptional quality and time resolution of beamline ID02. Via a stopped-flow device, protein dimers were rapidly mixed with a buffer solution triggering the self-assembly and injected into a scattering cell. Figure 87a shows TR-SAXS patterns collected during the capsid formation. We devised a kinetic model with three main species – dimers, intermediates and capsids – exchanging matter at reaction rates that were determined by fitting the experimental data.
 |
|
Fig. 87: (a) TR-SAXS patterns of self-assembling norovirus capsid proteins collected from 4 ms to 11,721 s. Black dots are the experimental data and solid lines are reconstructed patterns from a kinetic model. (b) Form factors P(q) of the three main species in solution. (c) Mass fractions of the three main species as a function of time estimated by a kinetic model. The vertical dashed line marks the end of the experimental data. |
An original global fitting procedure applied to the experimental data allowed us to extract the form factor of the intermediate species (Figure 87b) as well as to estimate the mass fraction of each species over time (Figure 87c). The three-dimensional structure, at nanometre resolution, of the intermediate species was then obtained by ab initio shape determination from its extracted form factor.
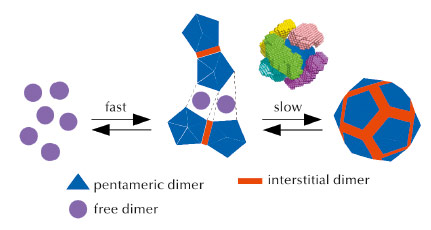
The analyses thus revealed that in the first step, some ten dimers combine to form a stave-shaped intermediate, possibly made of two pentamers of dimers connected by an interstitial dimer (Figure 88). In the subsequent, slower step, which takes hours, these intermediates interlock into a capsid. In contrast, capsids form by sequential addition of dimers in many other viruses such as the hepatitis B virus.
 |
|
Fig. 88: Kinetic scheme of norovirus capsid assembly. Free dimers are represented in magenta, dimers related by five fold symmetry in the final capsid in blue, and interstitial dimers in red. Above the last assembly step, a representation of interlocking intermediates is given as a possible mechanism. Six intermediates, each in a different colour, have been positioned above the six contiguous fragments made of two pentamers of dimers connected by an interstitial dimer. |
By clarifying the kinetics involved in norovirus assembly, this study provides a better understanding of the physical processes at work in the self-assembly of a viral capsid. It could also advance efforts to treat or prevent these infections, and it could be applied to engineer viral nanoparticles to make diagnostic agents and tailored therapeutics.
Principal publication and authors
G. Tresset (a), C. Le Cœur (b), J.-F. Bryche (a), M. Tatou (a), M. Zeghal (a), A. Charpilienne (c), D. Poncet (c), D. Constantin (a) and S. Bressanelli (c), J. Am. Chem. Soc. 135, 15373-15381 (2013).
(a) Laboratoire de Physique des Solides, Université Paris-Sud, CNRS (France)
(b) Institut de Chimie et des Matériaux Paris-Est, Université Paris-Est, CNRS (France)
(c) Laboratoire de Virologie Moléculaire et Structurale, CNRS, INRA (France)
References
[1] G. Tresset, V. Decouche, J.-F. Bryche, A. Charpilienne, C. Le Cœur, C. Barbier, G. Squires, M. Zeghal, D. Poncet and S. Bressanelli, Arch. Biochem. Biophys. 537, 144-152 (2013).
[2] I. Grillo, Curr. Opin. Colloid Interface Sci. 14, 402-408 (2009).



