- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2005
- Structural Biology
- RecBCD: A Molecular Machine for DNA Repair
RecBCD: A Molecular Machine for DNA Repair
The genetic integrity of all organisms is under constant threat from a variety of internal and external sources, such as radiation, chemical carcinogens and reactive cellular species. Often, these agents damage DNA directly. As a consequence, cells have evolved elaborate mechanisms to continuously scan for and repair such damage.
One of the major DNA repair pathways is that of homologous recombination (HR). In principle, this pathway allows an error-free restoration of the original genetic information. The repair process involves the copying of an undamaged strand of identical (homologous) DNA and splicing the resultant copy into the damaged strand. Taken as a whole, the process is quite complex, but a key step is the initial processing of the damaged DNA to generate free 3’ single-stranded tails [1]. In the bacterium E. coli, the RecBCD multi-protein complex, mediated by an octameric DNA sequence known as Chi (![]() ), catalyses this process.
), catalyses this process.
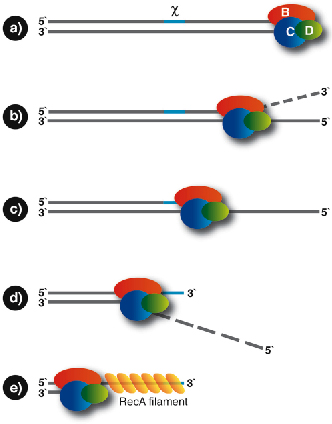
The RecBCD enzyme binds at the broken DNA end and rapidly unwinds the duplex in an ATP-dependent manner. As unwinding proceeds, the 3’ terminated strand is continuously degraded by an endonuclease activity. This unwinding and digestion continues until the Chi sequence is encountered, at which point the enzyme pauses, and the nuclease switches to the other DNA strand. The enzyme then continues unwinding the duplex, digesting the 5’ terminated strand and loading the RecA strand-exchange protein onto the 3’ strand, now capped by the Chi sequence (Figure 76) in preparation for the next stage of HR.
 |
|
Fig. 76: Schematic diagram of the reaction catalysed by RecBCD. The enzyme binds to a free duplex end (a) and starts to unwind the DNA and digest the 3’ strand (b). Upon encountering the Chi site (c), the nuclease activity is switched and the enzyme continues; now digesting the 5’ strand (d). RecBCD is also capable of loading RecA onto the exposed 3’ strand to provide the substrate for the next step of homologous recombination (e). |
Considerable amounts of biochemical and genetic data have been obtained on the RecBCD system. Recent results have shown that the enzyme has a bipolar helicase activity, with one motor translocating along each strand, each running with opposite polarity [2]. It has been shown that the 3’-5’ and 5’-3’ helicase activities reside with the RecB and RecD subunits respectively, the nuclease is associated with the C-terminal domain of RecB and Chi recognition occurs in RecC.
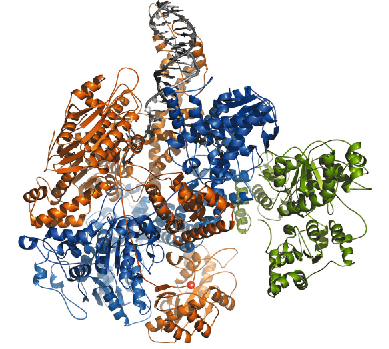
We have recently determined the crystal structure of RecBCD bound to a blunt-ended DNA hairpin using data collected on ESRF beamlines ID14-1 and ID14-4. The structure was solved in the absence of ATP and so represents an initiation complex prior to translocation (Figure 77). Despite this, the DNA substrate has been partially unwound, with four base-pairs of the duplex split.
 |
|
Fig. 77: Ribbon diagram of the RecBCD complex. RecB is shown in orange, RecC blue, and RecD green. The two strands of DNA (grey) are split at the top of the protein and enter tunnels directing the 3’ tail to the RecC Chi-recognition site and nuclease, and the 5’ tail to RecD. The active site of the nuclease is highlighted by a bound calcium ion, depicted as a red sphere. |
As expected from sequence analyses, both RecB and RecD have folds characteristic of superfamily I DNA helicases, with the C-terminal nuclease domain of RecB attached by a long flexible linker. In the crystal structure a calcium ion replaces the native magnesium ion in the nuclease active site. The RecC protein forms a plate-like scaffold around which the RecB is tightly wrapped, while RecD maintains a looser association. RecC has three channels running through it; a large central cavity responsible for binding RecB, and two smaller channels for each of the DNA tails. These channels guide the 3’ and 5’ tails towards the nuclease and RecD respectively. Intriguingly, the 3’ channel is partially formed from the N-terminal domain of RecC, which also has a characteristic helicase fold, a fact not apparent from the sequence. This observation suggests that RecC is in fact a defunct helicase, which has evolved the ability to recognise the Chi sequence. In the current crystal structure, the nuclease is ideally positioned to accept the 3’ tail of the DNA following its passage through RecC. Presumably the Chi sequence induces some conformational switch, which alters the nuclease strand specificity as well as activating the RecA loading process.
These structural studies have shown how two helicases can combine to form a highly efficient molecular machine and provide a basis for an elaborate regulated nuclease system. Unravelling the details of this control mechanism should prove to be a rewarding exercise.
References
[1] S. C. Kowalczykowski, Trends Biochem. Sci., 25, 156-165 (2000).
[2] M.S. Dillingham, M. Spies, S.C. Kowalczykowski, Nature 423, 893-897 (2003).
Principal Publication and Authors
M.R. Singleton (a), M.S. Dillingham (b), M. Gaudier (a), S. C. Kowalczykowski (c), D.B. Wigley (a), Nature 432, 187-193 (2004).
(a) Cancer Research UK, London Research Institute (UK)
(b) University of Bristol, Department of Biochemistry (UK)
(c) University of California at Davis, Center for Genetics and Development (USA)



