- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2013
- Structural biology
- How the ribosome decodes a modified ‘wobble’ position anticodon for accurate protein translation
How the ribosome decodes a modified ‘wobble’ position anticodon for accurate protein translation
Ribosomes are the macromolecular machines required for translating genetic information into proteins using a messenger RNA (mRNA) template and aminoacylated transfer RNA (aa-tRNA) substrates. One of the most remarkable characteristics of the ribosome is its ability to synthesise proteins at rates of between 10 and 20 amino acid residues per second with error frequencies as low as 1 in 10,000. This feat is mainly achieved by correctly pairing the three bases encompassing a ‘codon’ sequence on the mRNA with the anticodon of cognate aa-tRNA. In theory at least 61 different tRNAs would be required for the canonical Watson-Crick base pairing of all codons possible in mRNA. However, some mismatch base-pairing (i.e. non-Watson-Crick) must be allowed as there are less than 50 different tRNA anticodons observed in nature. Such mispairing occurring only at the third codon position (tRNA residue 34) led to the ‘wobble hypothesis’ proposed by Crick in 1966 [1], and is fundamentally important to understand because nearly 40% of all codons are decoded using this mechanism.
However, a growing number of post-translational modifications of tRNA anticodons have been reported, some of which have been implicated in human disease. The most common modifications are the methylation of uridine, deamination of adenine to inosine and the attachment of lysine to cytosine to form lysidine (k2C), which was first reported in Escherichia coli tRNA2Ile [2]. The unmodified tRNA2Ile has a CAU anticodon that is perfectly complementary to the AUG methionine codon. By modifying C34 to k2C34, a minor AUA isoleucine codon is recognised. A similar modification, the addition of decarboxy arginine to cytosine to form agmatidine (agm2C), is found in archaea. This suggests that the ability to discriminate between purines in the wobble position (AUA versus AUG) of codons is conserved in nature. To determine structurally how this is achieved, the crystal structure of the 70s ribosome in complex with the AUA codon and tRNA2Ile was determined using diffraction data collected at beamline ID14-4.
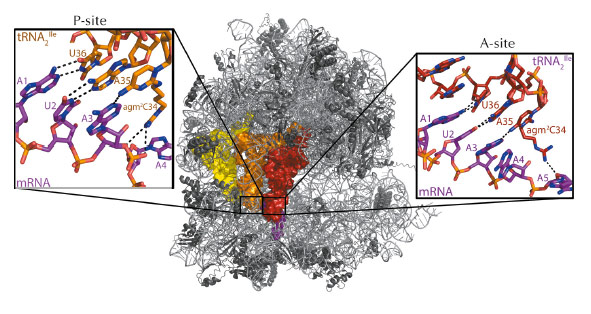
Archaeal tRNA2Ile was observed in both the amino-acyl (A) and peptidyl (P) sites of the 70s ribosome (Figure 59), providing an insight into how agmatidine confers specificity for AUA over AUG codons. As expected the ribosome was observed in a ‘closed’ conformation, with the first and second codon-anticodon positions mediating canonical Watson-Crick interactions. Surprisingly, the A3-agm2C34 is observed in a non-standard geometry in the A-site, forming a single hydrogen bond between the exocyclic amine of agm2C34 and the N1 of A3 (Figure 59). This is mainly due to the extended modification of C34 that hinders the adoption of a more stable tautomer that could potentially form two hydrogen bonds. A similar geometry was observed in the P-site resulting in a shift of both the mRNA and tRNA, which contrasts with previous studies showing the mRNA remaining stationary.
 |
|
Fig. 59: The crystal structure of the 70s ribosome with mRNA coding for two tRNA2Ile. The mRNA is coloured magenta with the tRNA2Ile coloured red for the A-site, orange for the P site and yellow for the E-site. The inserts show detailed views of the A and P-site codon-anticodon binding sites and highlight the interactions of the extended agmatidine modification with the sugar-phosphate backbone of downstream mRNA residues. Image credit: A.A. McCarthy, EMBL. |
Only a single hydrogen bond is observed in the wobble position, which suggests a thermodynamically unfavourable binding of tRNA2Ile. However, in the crystal structure, a hydrogen bond is formed between the terminal amine of agmatine and the O4’ atom of the ribose of a downstream mRNA nucleotide (Figure 59). A similar interaction is also observed in the P-site, this time between agmatine and the backbone phosphate of a downstream mRNA residue (Figure 59). These hydrogen bonds would also be possible with a lysidine modification of C34, indicating this might be a broadly conserved mechanism of binding tRNA2Ile, compensating for the weaker interaction in the wobble position. A steric clash between the exocyclic amine of agmatidine and a G modelled at the third mRNA codon would prevent a canonical Watson-Crick base pairing, providing insights into how the presence of agmatidine and lysidine would prevent binding of tRNA2Ile to the near cognate AUG codon. So, even 40 years after the genetic code was first elucidated, we are still unravelling the mysteries of how the ribosome accurately codes protein synthesis.
Principal publication and authors
R.M. Voorhees (a), D. Mandal (b), C. Neubauer (a), C. Köhrer (b), U.L. RajBhandary (b) and V. Ramakrishnan (a), Nat Struct Mol Biol 20, 641-643 (2013).
(a) Molecular Research Council Laboratory of Molecular Biology, Cambridge (UK)
(b) Department of Biology, Massachusetts Institute of Technology, Cambridge, Massachusetts (USA)
References
[1] F.H. Crick, J. Mol. Biol. 19, 548-555 (1966).
[2] T. Muramatsu et al., J. Biol. Chem. 263, 9261-9267 (1988).



