- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2013
- Enabling technologies
- The meta-world of metadata management at the ESRF
The meta-world of metadata management at the ESRF
Meta is derived from the Greek preposition µετ’α meaning “after”, “beyond”, “adjacent”, “self” [1]. In the scientific context it is often used to mean adjacent or about. In the case of metadata it is used to mean data about data. Typical examples of metadata are file name, owner and date created. In the case of the ESRF, metadata includes the experimental context under which the data were collected such as X-ray energy, sample, experiment parameters.
Data is one of the main products produced at the ESRF. Scientific results depend on the contents of the data produced. The ESRF is a big producer of data but often without metadata. With the exception of macromolecular crystallography (MX) and certain tomography experiments, most of the data produced at the ESRF does not have a standard set of metadata describing it. This means a large number of experiments rely entirely on the users manually collecting metadata in their notebooks. Although this does not prevent scientific publications it ultimately does influence the ease with which users can analyse their data. It also seriously hampers or even prevents online data analysis from being done because data cannot be analysed automatically.
Experiments which are collecting metadata in a systematic fashion already, MX and tomography, have developed specific metadata solutions. These consist of a dedicated metadata database and macros which collect the metadata during the data acquisition process. The databases are called IspyB and TomoDB respectively. The database table structures have been designed to take into account the specific needs of the experiments involved, i.e. crystallography and tomography. The IspyB database [2] has shown the advantages of metadata by becoming a central part of the MX experiments both before and after beam time. These database tables structures are however too specific to be applied to other kinds of experiments without major modifications. IspyB is used at two sites currently (ESRF and DLS) and there is interest from other sites for MX experiments.
PaNdata [3] is a consortium of all photon and neutron sources in Europe dedicated to sharing good practices of data management. One good practice is the collecting, storing and searching of metadata. PaNdata provides a generic database for storing and searching metadata called ICAT. ICAT [4] is an open source metadata management system designed for large facilities, which has been developed by ISIS and the Scientific Computing Department of STFC. ICAT is in production at ISIS and DLS, and several laboratories (e.g. ALBA, DESY, ELETTRA and ESRF) have deployed ICAT prototype instances, which have been tested during service verification runs. ICAT can be searched across facilities.
 |
|
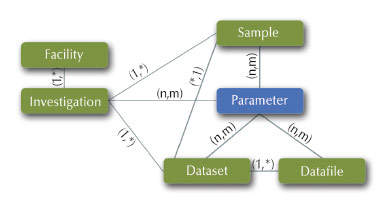
Fig. 154: Flexible ICAT metadata model. |
The ICAT database schema (Figure 154) is closely related to the standardised metadata schema implemented in NeXus. NeXus provides a complete description of the experimental conditions during an experiment, and covers essentially all of the data required for meaningful searches in a metadata catalogue. ICAT has many attractive features, such as a programmable web service interface, user authentication and authorisation, the integration of provenance information and the registration of data object identifiers. Developments at the moment include the ICAT data service (IDS) and using the Umbrella user authentication mechanisms.
 |
|
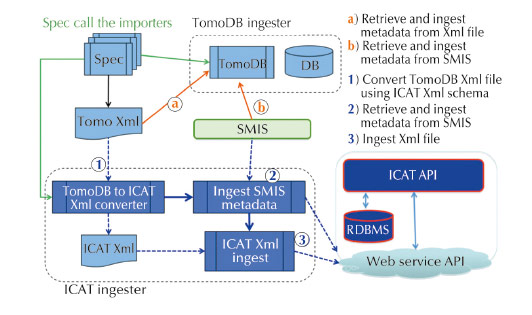
Fig. 155: Software architecture for TomoDB ICAT metadata ingester developed for ID19. |
The European funded projects Pandata-ODI and CRISP have financed resources for the ESRF to install and adapt ICAT to the ESRF beamlines. This project has been going on for 12 months and will continue to be financed for another 12 months. ICAT has been installed and an ingester for TomoDB has been written (Figure 155). Over the next 12 months an ingester will be written that can be used for all experiments to store generic and specific metadata in the database.
The goal of the metadata project at the ESRF is to collect metadata for all kinds of experiments on all beamlines automatically. Achieving this will allow metadata that is collected at the same time as the raw data to be fully exploited. Online data analysis will be facilitated and in some case made possible. With increasing data volumes online data analysis is essential for providing users with feedback on the data quality during the experiment. Another important issue addressed using metadata is data management including data archiving, exporting and searching. These issues are not managed systematically nor automatically at the ESRF today. A generalised solution for metadata will make this possible in the future.
Authors
A. Götz (a), N. Bessone (a), C. Cleva (a), D. Porte (a), J. Meyer (a) and A. Mills (b).
(a) ESRF
(b) STFC (UK)
References
[1] http://en.wikipedia.org/wiki/Meta
[2] S.Delangere et al., Bioinformatics 27, 3186-92 (2011).
[3] http://pan-data.eu/
[4] https://code.google.com/p/icatproject/



