- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2008
- Structural biology
- Structural characterisation of IrisFP, a biological photoswitch from the green fluorescent protein family
Structural characterisation of IrisFP, a biological photoswitch from the green fluorescent protein family
Since the successful cloning of the green fluorescent protein (GFP) from the jellyfish Aequorea victoria in 1992, fluorescent proteins (FPs) have revolutionised Life Science. The 2008 Nobel Prize in chemistry was awarded to O. Shimomura, M. Chalfie and R.Y. Tsien for having, respectively, discovered GFP, introduced it in cells as a luminous genetic tag, and refining it to a palette of ever brighter fluorescent proteins with various colours. Using the fluorescent protein toolkit, cell biologists can image a very broad range of biological processes, including gene expression, protein translocation within cells, and cell movement during development [1].
In recent years, new fluorescent proteins have been identified, notably in Anthozoa species (e.g. corals and sea anemones found in tropical reefs), which display photo-activation properties [2]. Proteins from the “Kaede-family” undergo a photo-conversion from green to red upon illumination with UV light. Proteins from the “Dronpa-family” undergo on/off photo-switching upon illumination with UV/blue light. The use of photo-activatable fluorescent proteins provided new means to dynamically track proteins within live cells [3], and to develop “nanoscopy”, a revolutionary set of techniques that defeat the diffraction barrier in light microscopy, achieving spatial resolution of ~10 nm with visible light [4]. Moreover, photo-activatable fluorescent proteins may find applications in nano-biotechnology, for example as novel data storage media [5].
All fluorescent proteins discovered so far display a so-called ß-can structure, forming an almost perfect barrel composed of eleven ß-strands. The chromophore is embedded at the heart of the ß-can, and is made of 3 amino-acids (X-Tyr-Gly) that undergo post-translational modifications to form an extended ![]() -electron system. Using X-ray crystallography, UV-vis spectroscopy and quantum chemistry, mechanistic details of the fluorescence mechanisms employed by photo-activatable fluorescent proteins can be obtained, which can be of considerable value to rationally engineer new variants with improved properties. In a collaboration between the ESRF, University of Ulm and the IBS, we have developed such a complementary approach. In the course of this project, we discovered “IrisFP” (the Greek goddess Iris personifies the rainbow), a remarkable fluorescent protein that derives from the fluorescent protein EosFP [6] and combines the properties of Kaede-like and Dronpa-like proteins, i.e. photo-conversion and photo-switching (Figure 82). Following proper laser illumination schemes, IrisFP can be switched reversibly between fluorescent and non-fluorescent forms in its green state, then photo-converted irreversibly from green to red, and finally switched reversibly between fluorescent and non-fluorescent forms in its red state.
-electron system. Using X-ray crystallography, UV-vis spectroscopy and quantum chemistry, mechanistic details of the fluorescence mechanisms employed by photo-activatable fluorescent proteins can be obtained, which can be of considerable value to rationally engineer new variants with improved properties. In a collaboration between the ESRF, University of Ulm and the IBS, we have developed such a complementary approach. In the course of this project, we discovered “IrisFP” (the Greek goddess Iris personifies the rainbow), a remarkable fluorescent protein that derives from the fluorescent protein EosFP [6] and combines the properties of Kaede-like and Dronpa-like proteins, i.e. photo-conversion and photo-switching (Figure 82). Following proper laser illumination schemes, IrisFP can be switched reversibly between fluorescent and non-fluorescent forms in its green state, then photo-converted irreversibly from green to red, and finally switched reversibly between fluorescent and non-fluorescent forms in its red state.
 |
|
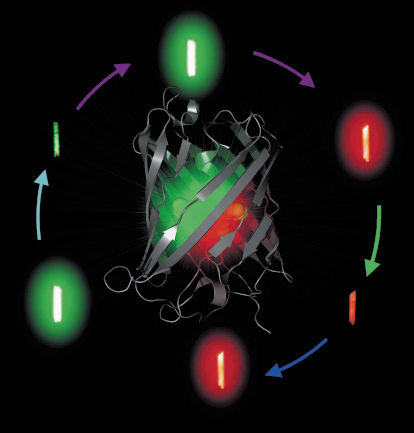
Fig. 82: The X-ray structure of IrisFP is shown at the centre. Around the structure are pictures of crystals of IrisFP in its different forms (switched on/off, green/red), recorded while fluorescing. |
We undertook a structural characterisation of IrisFP, in particular by using the Cryobench laboratory at the ESRF, which allows the illumination of crystals with a variety of lasers and the following of the induced colour changes by in crystallo spectroscopy. We discovered that photoswitching of green IrisFP involves a cis-trans isomerisation of the chromophore (Figure 83), accompanied by a major conformational change of the protein. In contrast, photo-conversion to the red-emitting state does not involve a large structural change, but results from a cleavage of the polypeptide backbone which induces an extension of the conjugated ![]() -electron system of the chromophore. Photoswitching of the red form of IrisFP appeared similar to that of the green form, involving cis-trans isomerisation of the chromophore.
-electron system of the chromophore. Photoswitching of the red form of IrisFP appeared similar to that of the green form, involving cis-trans isomerisation of the chromophore.
 |
|
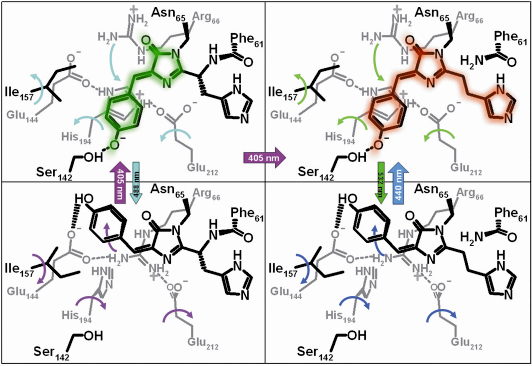
Fig. 83: Structural changes in IrisFP upon phototransformation. Structural motions induced by light are represented by curved arrows of the same colour as those used to represent light illumination at specific wavelengths. |
IrisFP holds great promises for applications in cell biology. Based on the mechanistic knowledge acquired in this work, rational engineering will be applied to further improve the behaviour of the protein. Applications taking advantage of the multiple phototransformations displayed by IrisFP will undoubtedly emerge, including two-colour nanoscopy for high-resolution spatio-temporal imaging, or sequential photo-activation schemes for unravelling complex protein-protein interactions. IrisFP also hints at the possibility of combining read-only and rewritable capabilities in future mass storage media.
Principal publication and authors
V. Adam (a), M. Lelimousin (b), S. Boehme (c), G. Desfonds (a), K. Nienhaus (c), M.J. Field (b), J. Wiedenmann (d), S. McSweeney (a), G.U. Nienhaus (c) and D. Bourgeois (a,b), PNAS 105, 18343 (2008).
(a) ESRF
(b) Institut de Biologie Structurale Jean-Pierre Ebel, Commissariat à l’Energie Atomique, Centre National de la Recherche Scientifique, Université Joseph Fourier, Grenoble (France)
(c) Institute of Biophysics, University of Ulm (Germany)
(d) National Oceanography Centre, University of Southampton (UK)
References
[1] R.Y. Tsien, Annu. Rev. Biochem. 67, 509 (1998).
[2] R. Ando et al., PNAS 99, 12651 (2002); Science 306, 1371 (2004).
[3] K.A. Lukyanov et al., Molecular Cell Biology 6, 885 (2006).
[4] E. Betzig et al., Science 313, 1642 (2006).
[5] M. Andresen et al., PNAS 102, 13070 (2005).
[6] J. Wiedenmann et al., PNAS 101, 15905 (2004).



