- Home
- News
- Spotlight on Science
- Iodide as a universal...
Iodide as a universal tool for solving new structures of membrane proteins
31-05-2017
A simple protocol of soaking the crystal of a membrane protein in a solution of iodide followed by freezing led to the incorporation of sufficient iodide ions to solve its structure by iodine-SAD. The general utility of this technique for novel structures has been demonstrated for several other membrane proteins.
Iodide has turned out to be a panacea for the resolution of the phase problem – a well-known obstacle in protein crystallography – when applied to the particular case of membrane protein crystals. This has been demonstrated in experiments at beamline ID23-1 whereby four crystal structures of membrane proteins from different species were solved through the incorporation of iodide ions. Although, the method has been in use since the early 2000s [1], iodide incorporation into the protein crystal structure via cryo‑soaking (soak-and-freeze) of protein crystals in iodide-containing solution is now experiencing a renaissance with particular application to membrane proteins.
Membrane proteins are very important pieces of cell machinery located within or close to the lipid membrane. Since their malfunctioning is associated with a wide range of diseases, structural studies of membrane proteins are a major research direction in applied medicine. To date, X-ray crystallography has been the foremost method for the determination of three-dimensional protein structures of the highest resolution. In a typical experiment, a protein crystal is placed in an X-ray beam and the diffraction intensities of the crystal are recorded. The information on the diffraction intensities can then be used to reconstruct the electron density of the protein molecule via a three-dimensional Fourier transform. The usual problem in crystallography is that the phases of Fourier coefficients are not known, since they are not measurable in the experiment. The phase information has to be retrieved computationally which can be achieved either by using data from a similar protein structure that has already been determined or by capturing the anomalies in the diffraction caused by heavy atom scatterers in the structure. The availability of known similar structures is limited, and the method employing them, named molecular replacement, does not always work well. The alternative, recording anomalous diffraction signal from heavy atoms in techniques such as SAD/SIRAS/MAD does not guarantee the successful solution either. Sometimes the signal is too low and requires highly-redundant measurements for it to increase above the level of measurement error, as is the case for the sulfur-SAD technique. The highest signal is provided by the heaviest elements, often metal complexes that are toxic and dangerous to work with and do not necessarily bind well to the protein (and also require long soaking time or labour-consuming co-crystallisation). In comparison, iodide has both sufficient anomalous signal strength at the X-ray wavelengths available at most synchrotron crystallographic beamlines and an intrinsic ability to bind to membrane protein molecules within a short time frame practical for cryo-soaking.
The idea of using iodide came from the real case of a membrane protein crystal structure that could not be solved by molecular replacement. The trial of iodide-soaking before other approaches saved a lot of time, effort and material. The protein structure solved by iodide-SAD contained clusters of iodide ions near the surface of the membrane, which led to our hypothesis of the universality of this method.
The general feature of all membrane proteins that makes the method reliable is positively charged amino acid residues specifically distributed in the protein structure. Commonly known as the “positive-inside rule”, this principle describes the existence of positive charge carried by amino acid residues such as lysine or arginine on a protein surface at or close to the inner surface of a cell membrane, which is essential for proper folding and insertion of a membrane protein into the membrane [2]. Iodide has been hypothesised to interact particularly strongly with the positive charges near the surface of a lipidic membrane.
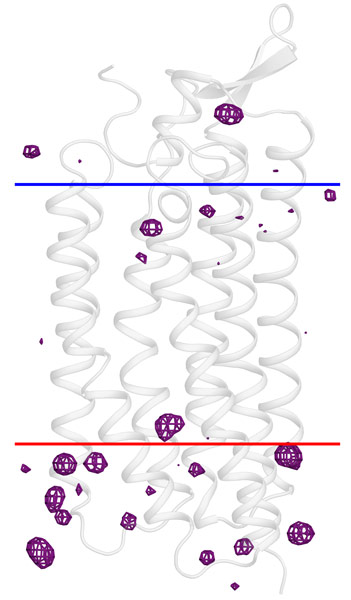
The hypothesis of the general applicability of this technique to membrane proteins has now been tested for four membrane proteins, the original case included. All four structures were solved straightforwardly by iodide-SAD on crystals that were soaked in sodium iodide-containing solution. Further investigation of the crystal structures revealed that clustering of iodide ions near the membrane surface (Figure 1), especially in the vicinity of positively charged arginine and lysine, is a common feature of the four target structures. These results clearly demonstrate the high affinity of iodide to membrane proteins, which coupled with its strong anomalously diffracting properties makes the method of iodide-SAD via crystal cryo-soaking a universal solution to address the phase problem in membrane protein crystallography.
Principal publication and authors
Fast iodide-SAD phasing for high-throughput membrane protein structure determination, I. Melnikov (a), V. Polovinkin (b,c,e), K. Kovalev (c,e), I. Gushchin (c,e), M. Shevtsov (e), V. Shevchenko (c,d,e), A. Mishin (e), A. Alekseev (c,e), F. Rodriguez-Valera (f), V. Borshchevskiy (e), V. Cherezov (e,g), G.A. Leonard (a), V. Gordeliy (b,c,e), A. Popov (a), Science Advances 3, e1602952 (2017); doi: 10.1126/sciadv.1602952.
(a) ESRF
(b) Institut de Biologie Structurale J.-P. Ebel, Université Grenoble Alpes-CEA-CNRS, Grenoble (France)
(c) Institute of Complex Systems (ICS), ICS-6: Structural Biochemistry, Research Centre Jülich (Germany)
(d) Institute of Crystallography, University of Aachen (RWTH) (Germany)
(e) Moscow Institute of Physics and Technology, Dolgoprudny (Russia)
(f) Department of Vegetal Production and Microbiology, Universidad Miguel Hernández de Elche, San Juan de Alicante (Spain)
(g) Departments of Chemistry, Biological Sciences, Physics & Astronomy, Bridge Institute, University of Southern California, Los Angeles (USA)
References
[1] Z. Dauter, M. Dauter, K.R. Rajashankar, Novel approach to phasing proteins: derivatization by short cryo-soaking with halides, Acta Crystallographica Section D Biological Crystallography 56, 232-237 (2000).
[2] G. von Heijne, Control of topology and mode of assembly of a polytopic membrane protein by positively charged residues, Nature 341, 456-458 (1989).