- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2012
- Soft condensed matter
- Unravelling the protein structural dynamics of photoactive yellow protein in solution using pump-probe X-ray solution scattering
Unravelling the protein structural dynamics of photoactive yellow protein in solution using pump-probe X-ray solution scattering
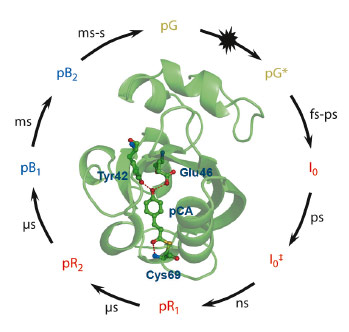
Photoreceptor proteins play crucial roles in receiving light stimuli that give rise to biological responses. The detailed time-dependent conformational transitions in their native aqueous environment have been elusive, however, even for a simple prototype photoreceptor, photoactive yellow protein (PYP). PYP is responsible for signal reception of the phototaxis response of the bacterium Halorhodospira halophila. It is one of the most studied biophysical systems and has served as a model for understanding the photoreception and the subsequent signal transduction at a molecular level [1,2]. Various experiments using spectroscopy, crystallography, and NMR have been performed to reveal the kinetic mechanism and the structure of intermediates involved in the photocycle (Figure 73). However, the global structures of intermediates in the solution phase have never been identified due to the lack of structural sensitivity, the restraint by crystal contact, and the limitation of time-resolution of the techniques used so far.
 |
|
Fig. 73: General photocycle of photoactive yellow protein with corresponding time region [1]. On illumination with visible light, the ground state (pG) is set on a photoreaction pathway with the isomerisation of the chromophore region. |
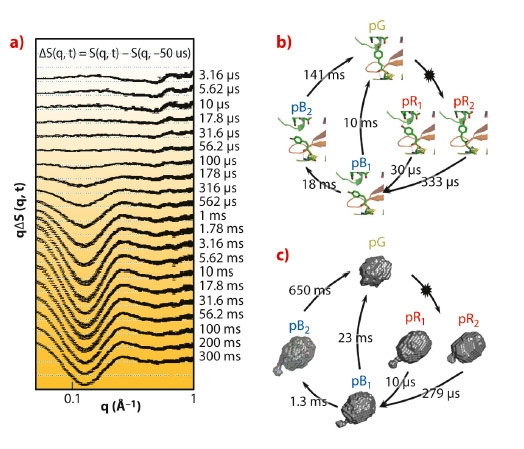
We employed the pump-probe X-ray solution scattering method established at beamline ID09B to overcome these limitations and unveil the structural dynamics of PYP in a wide time region from 3.16 μs to 300 ms (Figure 74a). The quantitative kinetic analysis was implemented to determine the number of kinetic components from the scattering curves. The results permit us to suggest the probable kinetic framework during the photocycle of PYP. Four intermediates and five time constants (10 μs, 279 μs, 1.3 ms, 23 ms, and 650 ms) during the photocycle were identified from the kinetic analysis. These rate-constants and the number of kinetic components are similar to the previous result observed with Laue crystallography [3] (Figure 74b). It implies that a common kinetic mechanism (parallel kinetics) can be operated in both the solution and crystalline phases. To describe the structural shape of an intermediate, the structural analysis, based on species-associated scattering curves determined from kinetic analysis, was performed. It permits us to visualise the global molecular shapes of all intermediates (pR1, pR2, pB1, and pB2) that show the gradual expansion of protein volume and the protrusion of the N-terminus with the progress of the photocycle (Figure 74c). The protrusion of the N-terminus becomes maximal in the pB2 state, the so-called signalling state.
 |
|
Fig. 74: a) Pump-probe X-ray difference scattering curves of the PYP photocycle. The photocycle of the photoactive yellow protein is triggered by blue light excitation (λmax = 446nm). b) The kinetic framework and the rate-constants in the crystalline phase determined by time-resolved Laue crystallography [2]. The chromophore motion is visualised with atomic-resolution. c) Reconstructed molecular shapes of the intermediates and their associated rate-constants in the solution phase determined from pump-probe X-ray solution scattering. |
Our results from pump-probe X-ray solution scattering observations provide several insights into the structural dynamics of PYP. First, the protrusion of N-terminus during the photocycle of PYP is suppressed in the crystalline phase due to the crystal contact [3]. However, in the solution phase, the unrestrained N-terminus is already elongated in the pB1 interediate, which is the precursor of the signalling state, and is maximised at the pB2. Second, the rate-constant of the intermediate accompanying large conformational change such as the N-terminus protrusion in the solution phase show a dramatic difference from those in the crystalline phase. Third, the movement of the N-terminus in the solution phase is highly correlated with that of the chromophore (p-coumaric acid, pCA).
Using pump-probe X-ray solution scattering, we can directly observe a profound effect of the molecular environment on structural dynamics and the reaction rates of the PYP photocycle. This study is the first instance where the three-dimensional molecular envelopes of the protein intermediates are reconstructed with microsecond time resolution and should serve as a cornerstone for further structural studies of protein structural dynamics in solution.
Principal publication and authors
T.W. Kim (a), J.H. Lee (a), J. Choi (a), K.H. Kim (a), L.J. van Wilderen (b), L. Guerin (c), Y. Kim (a), Y.O. Jung (a), C. Yang (a), J. Kim (a), M. Wulff (c), J.J. van Thor (b) and H. Ihee (a), J. Am. Chem. Soc. 134, 3145−3153 (2012).
(a) Institute for Basic Science, Center for Time-Resolved Diffraction, Department of Chemistry, KAIST (Republic of Korea)
(b) Division of Molecular Biosciences, Imperial College London (UK)
(c) ESRF
References
[1] K. Hellingwerf, J. Hendriks and T. Gensch, J. Phys. Chem. A 107, 1082 (2003).
[2] P. Ramachandran, J. Lovett, P. Carl, M. Cammarata, J.H. Lee, Y.O. Jung, H. Ihee, C. Timmel and J. van Thor, J. Am. Chem. Soc. 133, 9395 (2011)
[3] H. Ihee, S. Rajagopal, V. Srajer, R. Pahl, S. Anderson, M. Schmidt, F. Schotte, P.A. Anfinrud, M. Wulff and K. Moffat, Proc. Natl. Acad. Sci. USA 102, 7145 (2005).



