- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2014
- Electronic structure and magnetism
- HERFD XAS reveals a Mo(III) in the nitrogenase active site
HERFD XAS reveals a Mo(III) in the nitrogenase active site
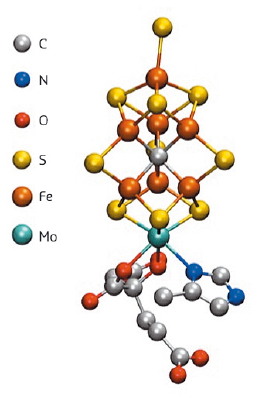
The conversion of inert nitrogen to ammonia represents a process that is fundamental for all life on earth. While both plants and animals require bioavailable ammonia for incorporation into proteins and DNA, neither plants nor animals possess enzymes that are capable of effecting this conversion. As such, we rely on diazotrophs (bacteria and archaea that are able to fix atmospheric nitrogen gas using the enzyme nitrogenase) as the biological source of ammonia. The enzyme active site that is responsible for catalysis is an MoFe7S9C cluster, known as FeMoco (Figure 23). The long-time enigmatic cluster was only recently revealed to contain a central carbon atom, and many details of the electronic structure remain unknown [1]. However, understanding the iron oxidation state, the molybdenum oxidation states, and the spin coupling within this complex biological cofactor is essential for understanding the mechanism and is a prerequisite for rational bio-inspired catalytic design.
 |
|
Fig. 23: The FeMoco site of nitrogenase. |
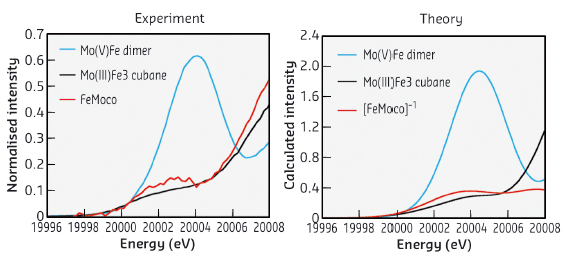
Recently, using high-energy resolution fluorescence detected Mo K-edge X-ray absorption spectroscopy (HERFD XAS) at beamline ID26, we were able to show that the nitrogenase contains a Mo(III) centre in the FeMoco active site (Figure 24). Previous Mo K-edge studies were unable to make a clear assignment of the Mo oxidation state in this cluster, due in large part to the significant core hole lifetime broadening at the Mo K-edge. By using HERFD XAS at beamline ID26, we were able to overcome many of the limitations of standard XAS measurements.
 |
|
Fig. 24: A comparison of experimental and calculated HERFD Mo K-edge XAS data for FeMoco and reference model complexes. |
Furthermore, by using a time-dependent density functional approach to calculate the Mo K-edges [2], we were able to verify the Mo(III) assignment and to also show that the Mo(III) is in an unusual spin-coupled configuration. This finding represents the first example of Mo(III) in biology. The implications that this electronic configuration has for nitrogenase reactivity are part of ongoing research efforts in our laboratory.
Principal publication and authors
R. Bjornsson (a,b), F.A. Lima (a,c), T. Spatzal (d,e), T. Weyhermüller (a), P. Glatzel (f), E. Bill (a), O. Einsle(d), F. Neese (a) and S. DeBeer (a,g). Chem. Sci. 5, 3096–3103 (2014).
(a) Max-Planck-Institut für Chemische Energiekonversion, Mülheim and der Ruhr (Germany)
(b) Present address: Science Institute, University of Iceland, Reykjavik (Iceland)
(c) Present address: Centro Nacional de Pesquisa em Energia e Materiais, Brazilian Synchrotron Light Laboratory – LNLS, Campinas (Brazil)
(d) Institute for Biochemistry, Albert-Ludwigs-Universität Freiburg (Germany)
(e) Present address: Howard Hughes Medical Institute and Division of Chemistry and Chemical Engineering, California Institute of Technology, Pasadena (USA)
(f) ESRF
(g) Department of Chemistry and Chemical Biology, Cornell University, Ithaca (USA)
References
[1] K.M. Lancaster, M. Roemelt, P. Ettenhuber, Y. Hu, M.W. Ribbe, F. Neese, U. Bergmann and S. DeBeer, Science 334, 974-977 (2011).
[2] F.A. Lima, R. Björnsson, T. Weyhermüller, P. Chandrasekaran, P. Glatzel, F. Neese and S. DeBeer, Phys. Chem. Chem. Phys. 15, 20911 (2013).



