( 1)
Contents
Mean square variation in path length
Calculation of atomic correlations
Muffin-tin Radii and related Parameters
Whole Sectrum Refinement ofCopper Foil
Tetrakis imidazole Copper(II) Nitrate
9. Fourier Transform parameters
12. Restrained refinement parameters
1. The crystallographic cell parameters
6a. Spectrum parameters - beam polarisation parameters
7a. Theory parameters - multiple scattering parameters
7b. Theory parameters - energy parameters
7c. Theory parameters - whole-spectrum parameters
9. Fourier transform parameters
10a. Surface parameters - surface displacement parameters
10b. Surface parameters - the surface relaxation parameter SURREL
12. Restrained refinement parameters
EXCURVE was written in 1982 as an alternative to the programs available at the time. It attempted to provide an integrated environment for the analysis of EXAFS spectra while providing a platform for the newly developed fast spherical wave method, later published by Gurman, Binsted and Ross (1984). The current version is based on this method for single scattering, but uses the method of Lee and Pendry (1975) for the exact polarisation dependent theory. Multiple scattering has options to use the methods of Lee and Pendry (1975), Gurman, Binsted and Ross (1986), and Rehr and Albers (1990). It allows fitting of both background-subtracted, and normalised total absorbance spectra. In the latter case the program calculates the atomic contribution of the spectrum. This is referred to as whole-spectrum fitting.
The purpose of the program is to find a structural model of a material which agrees with the available XAFS spectra.
In order to evaluate a structure, a model must first be defined in terms of one or more clusters of atoms, weighted according to the average composition of the material. Clusters may represent different atomic sites in a single phase, or multiple phases. A full description of each cluster requires :
1. the radial or cartesian coordinates of scattering atoms about each absorbing atom for which spectra are available
2. a pair distribution function for each symmetrically unique excited-atom/scattering atom pair
3. the point group of each cluster
The parameters used to define the model may be refined until optimum agreement with the XAFS data is obtained.
Refinement may use additional data to that given by XAFS spectra, for example distance and angle restraints using bond distances obtained by other techniques.
For crystalline solids, cluster information may readily be obtained from a crystallographic model. In this instance, the point symmetry of each site is defined and therefore all the distances and angles within the structure. A full multiple scattering calculation may then be performed.
For amorphous solids, only the partial radial distribution functions for pairs of atoms are defined, and the calculation is in general limited to single scattering.
In other instances, a model of the solid may include certain bond angles, or the presence of well-defined groups, which permit a limited treatment of multiple scattering. Such an approach is typical of metallo-proteins.
The manual is organised into a Theory section, a Program Guide, an Examples section, and Reference sections on parameters and commands, terminated by Tables. The program guide is largely a sequential guide, terminated by discussion of topics which might initially only be used optionally. Most of the background knowledge required is included in the theory section, but a basic knowledge of quantum mechanics is assumed, as is a knowledge of Unix or DOS commands, which can be used within the program to supplement the built-in commands. The reference sections contain definitions of the parameters and commands needed to analyse data, with information essential to their use but with a minimum of background material. References quoted in the text, together with others that might be useful, are given in the bibliography.
Acknowledgements.
We are grateful to the University of Washington, WA, and especially to Prof. J.Rehr for permission to use code derived from FEFF in calculating the Hedin-Lundqvist excited state exchange and correlation potential.
The refinement routine VA05A is used under licence from UKAEA, Harwell Laboratory.
This section discusses the calculation of the theoretical spectra, the way in which differences between theory and experiment are minimised, and the criteria used to define the best-fitting parameters.
Refinement involves minimising the quantity given by:
( 1)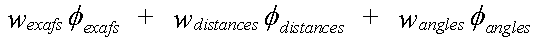
The weightings wexafs etc. determine the relative significance of the EXAFS, distance, and angle contributions. The sum wexafs + wdistances + wangles must equal one.
Often, only the XAFS contribution is used. The other two terms are restraints, used in special circumstances where a model includes well characterised groups of atoms.
The EXAFS contribution is given by:
( 2)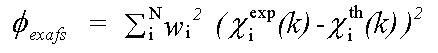
χexp(k) and χth(k) are experimental and theoretical XAFS:
( 3)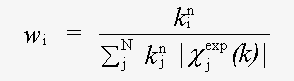
Here k is the magnitude of the photoelectron wavevector.
The distance and angle contributions to the refinement are:
( 4)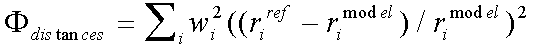
with:
( 5)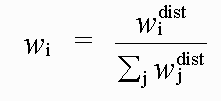
and:
( 6)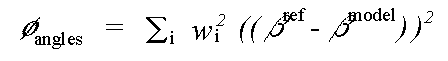
with:
( 7)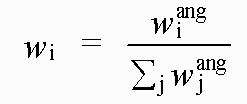
Where rref and rmodel refer to refined and model distances respectively, and where βref and βmodel refer to refined and model angles.
Calculation of the EXAFS theory
Analysis of XAFS spectra is greatly simplified by the fact that in a single particle approximation, provided coupling between final states is ignored, the atomic contribution (μ0) and the scattering contribution (χ) to the total absorption (μ) may be separated, as in:
( 8)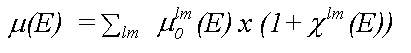
where the sum is over all allowed final states lm and the atomic contribution to the cross section, μ0lm (E) is given in the dipole approximation by the Golden rule expression:
( 9)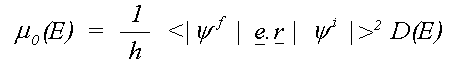
Here e is the electric vector of the photon, r the position with respect to the atomic nucleus and D(E) the energy density of final states. D(E) depends on the way the wavefunctions are normalised and is usually 1 for Rydberg states below the ionisation threshold and proportional to k, the magnitude of the photo-electron wave vector for a free electron.
Unless otherwise stated, Hartree atomic units (h=e=m=1) are used to avoid unnecessary constants, hence the unit of μ is here the square of the Bohr radius.
Many-body effects may be included approximately within this basic formalism, for example by including additional terms for two-electron transitions, and by using effective one-electron potentials calculated for an embedded atom with a complex energy to account for inelastic losses to represent the true many body potential.
It is normal to separate the oscillatory part of the experimental spectrum, χ, from the atomic contribution by means of a background subtraction, in which the atomic background is approximated by polynomial or spline functions. Here it is assumed that this has been done, and it is only necessary to calculate the oscillatory part. This is not the case with XANES or whole-spectrum fitting.
For an amorphous or polycrystalline sample, the oscillatory part of the spectrum due to a transition to a specific final state l is:
(10)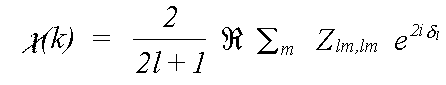
The final exponential factor is a phase term due to the outward and inward passage of the photoelectron through the central atom potential. It is described in terms of the atomic phaseshift δl, which is described later. The m sum is over all allowed quantum numbers -l ≥ m ≥ l.
Z is expanded as a scattering series:
(11)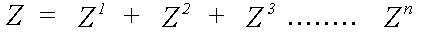
Z1 includes a sum over all atoms, excluding the central atom, Z2 a sum over all pairs of atoms i ≠ j, etc.
For single scattering from an atom at (r,Ω):
(12)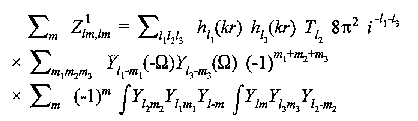
Simple algebra leads to the well known expression of Gurman, Binsted and Ross:
(13)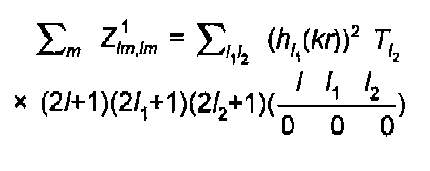
The l sums are strongly restricted by the rules on coupling of angular momentum: (l1+l2+l) even, l2≤l1l, l1≥l2-l. This results in just two terms for a K-edge, 3 for an L3, etc. The 3J coefficient above is that of Brink and Satchler (1968), also given by Zare (1988).
The equation above refers to atoms in fixed positions r. Disorder is expressed as a pair distribution function g(r) describing the motion of a scattering atom relative to the central atom. In the case of static disorder, g(r) may also describe the distribution of an atom in a disordered site, again relative to the excited atom:
(14)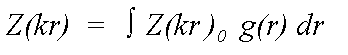
Where Z0 is calculated neglecting all thermal or static disorder.
Traditionally this has been approximated by:
(15)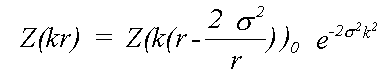
Here, even if Z is calculated exactly using a spherical wave theory, disorder is treated in a plane wave approximation, which ignores terms due to spherical-wave effects (see Rennert, 1992/3). This approximation gives rise to the familiar exponential Debye-Waller factor. This expression also ignores anharmonic contributions, that is the cumulant expansion of g(r) involves no terms of higher order than two. In this event, g(r) is a Gaussian characterised by σ:
(16)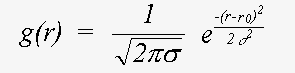
Also ignored here are terms arising from motion of the atoms in the plane normal to the interatomic bond. The inclusion of these would give rise to multi-dimensional distribution function g(r). The treatment of disorder is discussed at greater length below.
For the single-scattering, polarisation independent expression above [13], all the structural information is included in the Hankel functions h(kr). Similarly all the information on the scattering atoms is contained within the T matrix, whose l'th element is given by:
(17)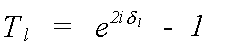
Here it is assumed that T is diagonal, that is, that the scattering potential is spherically symmetric. The atomic scattering phaseshifts which define Tl have already been encountered in the description of the central atom phase. The scattering phaseshift, δl, for each partial wave of angular momentum l contains all the information that is needed to completely describe the scattering properties of an atom.
Before discussing the calculation of the phaseshifts it is necessary to introduce a model for the atomic potentials in a solid.
In a free atom, the electron density near the nucleus is associated with tightly bound, relatively localised orbitals. Valence orbitals are less localised, and there is finite electron density well beyond what would normally be considered the atomic radius. An example is shown in figure 1 (top). This is the radial charge density, 4/3πr3ρ, calculated for Cu, using a relativistic Hartree-Fock program. This was used to generate the charge-density tables for the program. The program calculates potentials and charge densities self-consistently in the one-electron approximation, in which the complex many-body interactions between electrons are approximated by an effective one-electron potential. A potential is the best way of representing the force on an electron due to its electro-static interaction with the nucleus and other electrons. The potential also includes the quantum effects of exchange and correlation, which lower the potential further relative to a purely electrostatic model. The one electron potential can be used to calculate the Schrödinger wave functions ( or their relativistic equivalents, the Dirac wave functions) either for an electron in the atom, or a photo-electron passing through it.
It is not necessary to tabulate both the charge-densities and the potentials - the effective one-electron potential can be recovered from the charge density using Poisson's equation:
(18)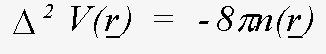
In XAFS however, what is usually required, is a potential for a solid or molecule rather than a free atom. The muffin-tin approximation is a simple model of a solid. The solid is defined as an array of touching spheres, using a regular lattice, such as FCC, BCC. Inside the spheres the potential is atom-like (though with spherical symmetry). Between the spheres (26% of the crystal volume for FCC), the potential is a constant value, obtained from averaging the potential in this region. The justification for this model is that the potential due to overlapping those of individual atoms, is lowered, and flattened relative to the free atom in this region. The Muffin-tin potential is further lowered relative to the free-atom potential by a term for the exchange and correlation energy of the photoelectron. An example is shown in figure 1 (bottom). This is for FCC Cu, looking along the (110) direction of the crystal. The solid lines are the overlapped free atom potentials. The dotted line is the muffin-tin potential for the solid, including the ground-state exchange contribution. An excited state exchange contribution (a +ve correction which raises the potential), is added later. It depends on the energy of the
photoelectron.
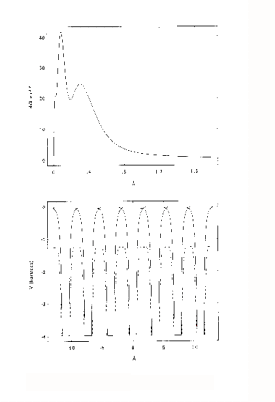 Figure 1: Atomic Charge Density (above) and Muffin-tin
potential (below) for Cu.
Figure 1: Atomic Charge Density (above) and Muffin-tin
potential (below) for Cu.Ground state Exchange and Correlation potential
The correct treatment of exchange is controversial. Indeed, in view of the simplicity of the model that is used, and the wide range of compounds to which the method is applied, it is unlikely that a single equation will give the best results in all cases. The program has two options for the ground state exchange term, and two for the energy dependent excited state term. Evaluating the four resulting options can provide weeks of amusement. For ground state, the two schemes are the X-α and the von-Bart and Hedin terms. These are both functions of the local electron density ρ(r). The X-α formula, in Rydberg atomic units is (Clarke, 1984):
(19)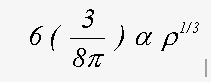
where α normally takes a value of 2/3 and with α=1 is equivalent to the Slater free electron formula.
The von Barth and Hedin (1972) formula is:
(20)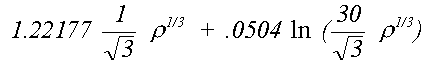
The appropriate term is used both in the Hartree-Fock calculations used to derive the atomic charge-densities and in the photo-electron exchange and correlation. There is no consistent preference for either of the two approaches, either in terms of quality of fit or agreement with crystallographic results. Where a significant difference occurs, it is usually due to a small energy shift in a resonance where one of the phaseshifts goes through π/2. In applying the Mattheis method to compounds other than the metals or van der Waals solids for which it was designed it might be expected that some flexibility needs to be introduced to ensure alignment of these features.
Within the muffin-tin model, the electrons outside the spheres can easily be described as complex spherical Bessel functions, with the photoelectron described as a sum over many partial waves of given angular momentum when it is necessary to describe it with reference to a point other than the centre of the excited atom. It is never actually necessary to know what happens inside the spheres. It is only necessary to know 1. the phase-change when passing in and out of the central atom and 2. how an electron is scattered by the sphere.
Quantum scattering is totally unlike classical scattering. An electron is scattered whenever there is a change in potential. In the constant potential region there is no scattering. At the muffin-tin boundary, where there is a small, and rather unrealistic step, and within the sphere, there is a scattered and transmitted wave, the intensity of each of which varies continuously. In practice, most of the scattering occurs where the potential varies most strongly, which is near the nucleus. All that is required to describe scattering completely is the phaseshift evaluated at the muffin-tin boundary. This is given by δl in:
(21)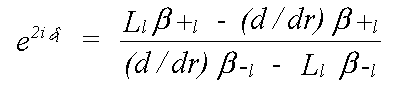
where the β are forms of complex spherical Bessel function, and the L are logarithmic derivatives of the Schrödinger (or Dirac) wave function given by:
(22)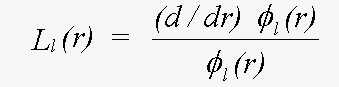
Evaluation of L involves an integration of the potential, out to the muffin-tin radius, using the boundary conditions that φ must not be singular at the origin, and must match the photoelectron wave (spherical Bessel function) at the sphere radius. The program optionally allows for scalar relativistic terms, i.e. ignoring those involving spin-orbit coupling. If necessary, the wave-function itself, and the dipole transition rate between a core state and φ may be evaluated so the atomic contribution to XAFS can be included.
Because the excited state contribution to the potential includes the effect of inelastic losses, due to core-hole lifetimes and electron inelastic scattering, the wave-function, the phaseshift and the scattered wave intensity given by exp(2iδl) are all complex, the imaginary part of δl being derived from the imaginary potential.
Thermal and static disorder have a significant effect on both XRD and EXAFS spectra, yet in neither technique is disorder treated exactly. The approximations used might be expected to give rise not only to incompatible values for the disorder parameters but also systematic errors in distances. This means that distances and disorder parameters may not be comparable between the two techniques. The treatment of disorder attempts to minimise these problems, using methods outlined in Binsted, Pack, Weller and Evans (1997).
Information on disorder in solids is often derived from structure determinations by X-ray or neutron diffraction.
For X-ray and coherent neutron methods it is usual that the thermal disorder associated with each atom is represented in the harmonic approximation by a mean square displacement <u2(r)>, with components <u2x>, <u2y>, <u2z>. In the case of powder diffraction, or low resolution single crystal refinements, it is further assumed that motion is isotropic and can be represented by an isotropic thermal factor given by:
(23)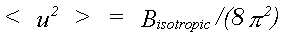
An exact treatment of disorder in both techniques requires a configurational average over all possible atomic positions. For EXAFS each path can be treated individually, giving rise to an integral over the three coordinates of each atom in the path:
(24)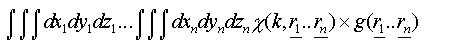
If many-body correlations are ignored these integrals are of the form:
(25)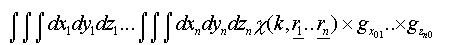
Where the g are pair distribution functions for each leg of the scattering path for each coordinate (x,y,z).
For single scattering only, if isotropic motion of each atom is assumed, equation (25) reduces to a single integral over the mean interatomic separation rm, as assumed in (14). Due to the effect of motion in three dimensions, rm differs from the equilibrium separation between atoms r0 by rm = r0+σ2/2r0 where σ2, the mean square separation in interatomic positions, is assumed small in comparison with r0. This makes the assumption that correlation is isotropic, although this is unlikely to be the case. Indeed, normal to the bond, motion is as likely to be anti-correlated as correlated. This integral can be evaluated numerically, avoiding further approximations, or else solved assuming the asymptotic form of the Hankel functions, h(kr) = 1/(kr) eikr. An approximate solution in one dimension has been given by Tranquada and Ingalls (1983), which, separating the r-dependent terms in the expression for χ(k) is:
(26)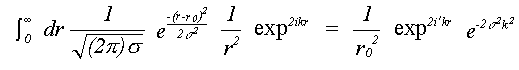
where:
(27)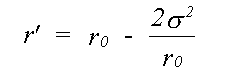
Previously only the Debye-Waller factor e-2σ2k2 , neglecting the phase term, has been used. In EXCURVE there are two options, one to perform the integral numerically, and one to use a plane-wave Debye-Waller factor but including the phase term. Using the numerical integral automatically includes spherical wave effects. Failure to include the phase term produces a small but significant apparent shortening in EXAFS distances in most cases. When disorder is large, as in inert gas solids, neglecting this term will give significant errors, such as the apparent thermal contraction noted by a number of authors ( see for example Beattie et. al. (1990)). If the term due to three dimensional motion is included, the overall distance correction will be smaller than that of Tranquada and Ingalls (1983). An option to include this term is present in the program. but due to lack of theoretical or experimental evidence for the model of 3D correlation, and the fact that it does not seem to improve agreement with crystallographic results, its use is not recommended at present.
The Debye-Waller term can be generalised to e-1/2σp2k2 where σp is the mean square variation in path-length. The same expression then describes the amplitude term for multiple scattering paths in addition to single scattering. Numerical results indicate that the phase terms are less significant for most MS paths than for single scattering, and for simplicity are neglected. The effects of disorder on bond angles can be represented by calculating the mean bond angle at each atom. This differs significantly from the equilibrium value only for angles close to 1800 when disorder will always result in smaller values. MS is particularly sensitive to changes in angles for these values hence the effects can be important and are included in the calculations.
Third and fourth order cumulant terms can now be used in the program both in the direct integrals and when using the plane-wave Debye Waller terms. If thermal expansion can be adequately represented by an isotropic coefficient of linear expansion, then only the linear expansion coefficient, α, need be entered in order to calculate the third cumulants for all the shells. This is done using an anharmonic oscillator model (see Edwards et.al.,1997). The third cumulant terms appear to make a noticeable contribution to the spectrum and their use in phases such as Cu where the model is good ( at least for shells 1,4 etc.) is being evaluated. It is not recommended that higher cumulants are refined - they are so strongly correlated with other variables (eg C3 with EF+R1) that almost any solution will appear successful.
An additional model of disorder, using a truncated exponential convoluted with a gaussian, is also included in the program. This is appropriate for certain systems of high disorder, such as in melts and ionic solids, where the series of cumulants fails to converge.
Mean square variation in path length
The mean square variation in path length can be expressed in terms of the atomic mean square displacements <ua2>, <ub2> etc. for each atom and the correlations between pairs of atoms Cab. Here anharmonic or anisotropic effects introduced by correlation and many-body correlations are ignored, although they may be important in many cases, and our intention is to include them in future versions of the program.
The expression for the mean square variation in path length takes into account the fact that the photoelectron velocity is fast in comparison with thermal motion. If an atom is included in n legs of the scattering path, the contribution it makes to σp is n times of an atom at a 'loose end'. For σp2 the contribution is n2 times. This is an important factor leading to a reduction in the contribution of triple scattering paths involving only two or three atoms, such as paths 0-a-0-a-0 or 0-a-b-a-0 ( 0 is the central atom ). For single scattering this generates the traditional term:
(28)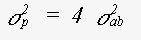
where the mean square relative displacement σab2 is:
(29)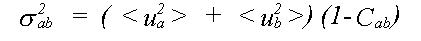
The general result for σp2 is:
(30)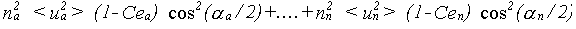
The effective correlations Ce are dependent on all the angles in the path. This can be appreciated by taking a long linear chain of atoms. The correlation effecting an atom at one end is that with the atom at the other end, not any of the atoms in between. If the chain departs from linearity, the intervening atoms will all make some contribution. No accurate solution to this problem has been obtained; it is assumed that all the correlations contribute with a relative weight determined by the same cos2(α/2) dependence as in equation 30. Ce is therefore given by:
(31)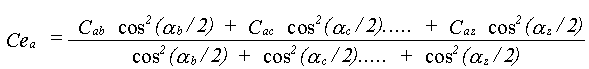
The equation only applies where there is a unique angle at each atom. Complex paths with many non-parallel legs involving the same atom are treated differently using further approximations..
Calculation of atomic correlations.
In order to obtain meaningful Debye-Waller factors it would be desirable to use just a single isotropic thermal parameter for each crystallographic site. In order to derive EXAFS Debye-Waller factors however, it is necessary to calculate the correlations between them as defined by equation (31) . The best way to do this is by means of a common theory which will generate both the atomic mean square displacements, <u2> and the correlations Cab. This can be done using Debye theory. A widely used expression for a monatomic cubic solid is given by Beni and Platzmann, 1976. The results of applying this to copper are given below. This expression has been generalised for binary metal oxides, but agreement with experiment in this case requires ad hoc expressions for the mass dependence which are still being investigated.
In many cases, for example where strongly covalent bonding occurs, Debye theory would not be expected to work. In such cases there is the option of using a single set of atomic displacements, and specifying the important correlations. Further correlations, which are principally a function of interatomic distance, are interpolated, and assumed to tend to zero for outer shells. A variant of this option which is also available is to define the correlations in terms of three refinable polynomial coefficients. This option ensures a realistic model of disorder for both methods, and reduces the number of free parameters when compared to the third method available, which is to refine XRD and EXAFS thermal parameters independently. In the latter case, because only the mean square displacements relative to the central atom are available, it is necessary to approximate σp2 for multiple scattering by:
(32)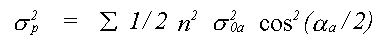
Where the sum is over all unique atoms, and σ20a is the mean square relative displacement between atom a and the central atom (except the case that a is itself the central atom when σ102 is used).
Polarisation dependence enters into the calculation, because in a dipole interaction, the momentum-vector of the emitted electron lies parallel to that of the electric field vector e of the photon. In an isolated atom, the matrix element is zero for transitions to orbitals with an angular momentum vector parallel to e. In a solid, scattering of the final state must be considered. The matrix element is then given by:
(33)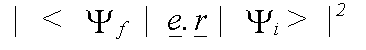
with the final state:
(34)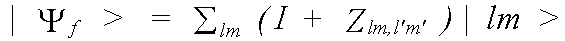
where |lm> is the unmodified final state wavefunction
Substituting this in the expression for XAFS, without the angle-averaging, only the radial part of the matrix element now cancels top and bottom, leaving:
(35)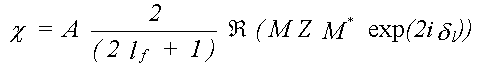
In general M is an i by j matrix where, for a specific transition, i is the number of allowed m values for the initial state, and j the number of m values for the final state.
In the case of a K-edge (i=1, j=3) the direction vector M = (A, -B, -A*) is given by:
(36)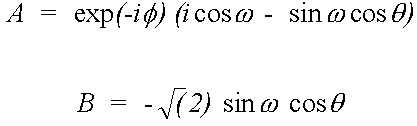
Where (θ,φ) is the beam direction and ϖ the azimuthal angle of the e vector, in the notation of Gurman (1988).
In the small atom and plane wave approximations to the scattering matrix Z, the scattered components normal to a particular leg of the scattering path actually go to zero. In this instance the expression above simplifies greatly and goes to:
(37)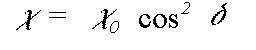
where delta is the angle between e and the leg of the scattering path. This approximation is in fact extremely good in the EXAFS region, but becomes very poor very near the edge, where the difference between the different polarisation directions is reduced. The program offers the exact polarisation dependent calculation as an option, at very considerable computational expense.
The above discussion ignores two factors which govern the EXAFS amplitudes. From equation (10) onwards it was assumed that there is one scattering atom per excited atom, with either a fixed distance, or small variations in distance governed by a pair distribution function g(r). In practice, in order to perform the sum over all atoms, it is necessary to define many shells of atoms, each with an occupation number N representing the average coordination of the excited atoms. The sum over all atoms can then be expressed as the sum:
(38)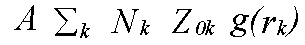
Where the sum is over all shells k with occupation number N and pdf g(rk). The factor A is the other missing term. In the program it is represented by the parameter AFAC. In other codes it is sometimes called S02. It represents the average proportion of excitations which contribute to EXAFS. That is, it provides a measure of events such two-electron transitions where the energy difference between the photon and the photoelectron is so large that the they are not seen in the spectrum. If the effect of such events are not included in the excited state contribution to the potential, then typically A should be .7 to .9, depending on the edge in question. For a Hedin-Lundqvist potential, however, multi-channel events are included, although only approximately, and only above the plasmon threshold. Although in principal A should be 1 with the HL theory, it may be necessary to use slightly lower or higher values to compensate for errors in the theory. In particular, if the edge region is fitted, a lower value may be required. Adjusting both A and the effective core-hole lifetime to obtain a good fit to a model compound, although theoretically unsound, is often the only possible procedure.
The EXAFS R-factor is defined as:
(39)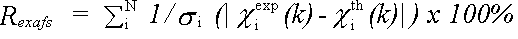
and gives a meaningful indication of the quality of fit to the EXAFS data in k-space.
A value of around 20% would normally be considered a reasonable fit, with values of 10% or less being difficult to obtain on unfiltered data.
An absolute index of goodness of fit, which takes account of the degree of overdeterminacy in the system is given by the reduced chi2 function. For EXAFS this is (Lytle et al., (1989)):
(40)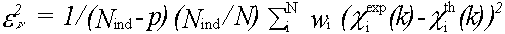
where Nind is the number of independent data points and p the number of parameters. Nind is normally less than the number of data points N , and in the case that the data from kmin to kmax is Fourier filtered using a window rmin to rmax it is given by:
(41)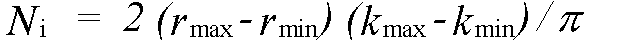
rmin and rmax should indicate the range in r-space actually fitted, not just that where structure is apparent.The variable p should include all parameters refined at any time, not just those included in the last refinement. In the program, Nindis calculated automatically, but may be overridden if the automatic value is inappropriate. p must always be entered by the user. Both these parameters should be quoted and justified along with chi2 if changes in chi2 are to be used as evidence for a fit. The absolute value of chi2 is not meaningful, unless actual experimental statistical errors σi have been read-in, and used to weight the spectra.
The program is started by typing EXCURVE from a Unix/Linux or DOS command prompt, or using the desktop icon provided. Generally a number of versions are provided, with different dimension limits, which involve trade-offs between, for example, the number of allowed data points, and the number of stored multiple-scattering (MS) paths (which allow rapid refinement of MS Debye-Waller factors, and experimentation with filter conditions).
The program operates using a series of commands:
command keyword option
for example:
There are also a number of special characters, which will be described before the commands are introduced.
Indicates that control is to pass from the keyboard to a command file. For example:
%abc
Will execute the commands in file abc before returning to the terminal. The normal way of starting the program is to execute a startup file in this way. The startup file may contain filenames, options and command aliases that are invariably used in connection with a particular compound. Command files may also be used for frequently used command sequences or lists of variables, as with Word-Perfect or assembly language 'macros'.
Is a comment line, usually only used within command files. The only action the program will take is to write the line in question in the logfile and at the terminal.
Accesses UNIX/Linux sh, or DOS commands - (there is no way of accessing commands unique to csh or other Unix shells, or aliases defined in csh .files). For example:
^rm -r ~/*
Will remove the whole of the current users filestore.
Gives a menu. At the command prompt ( ENTER COMMAND: ), it gives a list of commands. Following a command it gives a list of keywords, and in some cases options. For example:
REFINE ?
Gives a list of keywords for the REFINE command.
Has the same effect as a carriage return key. It may be used in order to enter several commands on one line, or at the end of a line, so as to accept a default value and avoid a prompt. e.g.:
C X1 .1;C X2 0
to change two x-coordinates to .1 and 0 respectively
PR 1120;
to output EXAFS spectra, using the default filename
Is used as an escape character. It can be used almost anywhere to terminate a command, or skip to the next part of a command. It must be followed by a carrommand. <esc> may also be used, but must again be followed be a carriagew return.
INFO SPACE;=
will terminate a listing of space group symbols after the first page
Each command can be executed using the minimum unambiguous string that will define the command. Some commands also have specific abbreviations which can be used in place of the minimum string. Keyword abbreviations are treated differently. The first entry in the list of keywords will be used (usually but not always alphabetical). Thus R P will match READ PARAMETERS and READ PHASE, but READ PARAMETERS will be used as it comes first. The names of parameters, such as AFAC, may not be abbreviated. Character options may usually be abbreviated to one letter.
Command Abbreviations Function
ALIAS AL allows you to define your own abbreviations for commands or sets of commands
ANGLE AN display an angle defined by three atoms
CALC CA used to calculate atomic potentials and phaseshifts
CCHANGE CC allows parameters to be changed using constraints
CHANGE CH C used to change the values of structural and theoretical variables.
COMPARE COM CM compares another experimental file with the current experiment and theory.
CONFIG CON configures fonts and other hardware features
COPLOT COP CP displays fourier transforms at the same time as the theory and experiment.
DEBYE DE calculates the shell Debye-Waller factors using Debye theory.
DISPLAY DI shows a table of interatomic distances and angles.
DRAW DR produces a three dimensional display of clusters, units or paths.
END END terminates the program.
EXCLUDE EXC exclude regions of the spectrum from least squares fitting or statistical analysis.
EXPAND EXP generate a cluster with C1 symmetry from higher symmetry (produce single-atom shells).
EXTRACT EXT generate a file containing a sub-set of multiple scattering paths. Also used to view information on multiple scattering paths.
FFILTER FF allows back-transformation of part of the fourier transform so that shell contributions can be isolated.
FIT FI updates the theory and displays fitting statistics.
FTSET FT selects options which control the calculation of fourier transforms.
GENERATE GE produces a new cluster using an atom in an existing cluster as the centre
GSET GS selects options which control the appearance of graphs.
IDEALISE ID convert amino acids to ideal geometry
INFO INF give tables of information used by the program
INQUIRE INQ displays information about the current state of the program control parameters.
LIST
L
lists current values of variables, and of SET, GSET and FTSET options.
MAP
M
draws a contour plot of the variation of fit-index with parameter values. PLANE
PLA
3-D geometry associated with planes of atoms. PLOT
P
displays experiment and theory, Fourier transforms, phaseshifts etc. PRINT
PR
generates files of spectra and/or parameters and variables. READ
R
reads in files of experimental data, phaseshifts and parameters. RECOVER
REC
restore parameter values, e.g. from previous session. REFINE
REF
used for the refining variables to produce the best fit. ROTATE
RO
rotate cluster or units RULE
RU
defines algebraic rules which constrain parameters SAVE
SA
saves current parameters, errors and statistics for recall, especially in conjunction with table. SET
S
allows modification of parameters which control major program options. SITE
SI
define mixed-atom sites SORT
SO
puts shells in order of distance SPARE
SPA
compresses collections of symmetry related MS spectra STATS
STA
controls the Joyner statistical tests. STRUCTURE
STR
generates shell coordinates for simple structure types. SYMMETRY
SY
calculates atomic positions from shell variables given the point symmetry. TABLE
TA
displays comparative tables of parameters, errors, and statistics. TIME
TI
displays current time of day, CPU usage and elapsed time. UNDO
U
remove the effect of the last CHANGE or ROTATE XADD
XAD
allows contributions from batch calculations to be added to the theory. XANES
XAN
calculates and displays XANES by matrix inversion method. A full list of commands can be obtained by typing ? at the command prompt. A full description of each command is given in the command reference section of the manual, chapter 5. All the keywords and
options are described there. Here only sufficient information is given to start analyzing data. A thorough knowledge of the theory
section is assumed. Menu options for each command can be obtained by typing command_name ?.This document is available
in Word Perfect, MSWord and html versions. Theory calculation, refinement and restraints are governed by a large number of parameters. These are listed and described in
the PARAMETERS section of the documentation. Parameters are changed using the command CHANGE and displayed using
the command LIST. Program control is by means of innumerable options: these are randomly distributed amongst the commands
SET, GSET, and FTSET. The keywords and options associated with these commands are fully described in the command
reference section. The command READ is used to read in one or more EXAFS spectra, structural parameters or scattering phaseshifts. EXAFS spectra can be read using: READ EXPERIMENT n where n (1, 2, 3 etc.) is incremented for each experiment read. Background-subtracted data given in terms of eV above the edge position. For whole-spectrum analysis, however, absorbance
as a function of absolute photon energy may be used. The variable E0 is then used to define the edge position (E0 must be 0 if
the energy is relative to the edge). With Unix/Linux, make sure that the data is in Unix format - terminated only by a carriage return
character. Most Linux commands now accept both Unix and DOS formats, making it difficult to determine what the format is. For
the Windows version only, it is possible to browse files, be entering *.* at the command prompt, or *.suf to restrict the search to
a particular file suffix. If your data is in k_space use the command READ KEX instead. E0 is then the difference between the edge
position and the origin of the wave vector (Ez in k^2=2.(E+Ez)). It is essential that this is correct. For background programs which
write only absorbance versus k, Ez is often but not necessarily 0. The program will prompt for information [default values are given in brackets - type return to accept them]: Point frequency ? [1] 1 to read every point, 2 for alternate points etc. this option is useful in speeding up the early
stage of analysis or reading files where the number of points exceeds the program dimension
limits. For EXAFS spectra - determines the location of the energy (ev) and absorbance columns in
the file. e.g. 79 if energy (ev) is in column 7 and absorbance in column 9. Edge ? [CU K] defines the central atom type and the initial state. The reply should be an element symbol +
a label for the core electron excited, as two words .e.g. CU K, RB L3, W L1 Sequence-number in polarisation set [0] is set to 1, 2, 3 etc. for polarisation dependent spectra differing only in their orientation
(otherwise 0). Members of a polarisation set must use a common edge position. Number of clusters for this experiment [1] is used to relate the atomic clusters defined by the parameter table to the various spectra. It
is 1 or more unless the spectrum shares clusters with the previous spectrum. More information on using READ is given in the Command Documentation section (chapter 5), and under fitting multiple spectra
in this chapter. Having read in the data, it is useful to check it. This can be done using PLOT EXAFS. Neither the theory nor fitting statistics will
make sense at this stage, because no structural model has been defined, and no atomic phaseshifts are available. The EXAFS
plot may not show much structure because it is dominated by data below the edge. In this case either decrease the range used
by changing EMIN: C EMIN 10 S W K3 If something is wrong it might be helpful to check the correct filename was entered: LIST FILES If the problem is unresolved, check how many points were read, what energy range was used, etc. using: If the energy range is too short or too long, the spectrum may have been read using the wrong column combination, the number
of data points may have exceeded the current dimension limit or E0 may be wrong (it should normally be zero). The dimension limits may be inspected using: LIST DIMENSION If the high energy end of the spectrum does not appear, it may be necessary to change EMAX, e.g. to any high number. C EMAX 9999 A model for the structure should now be defined. This may be done in a number of ways: 1. Read a program parameter file. READ PAR Once created, a parameter file can be edited, but it is tedious to create it from scratch. If parameters are available in Brookhaven protein database format, or .car files are available from the Cerius Explorer modelling
program, these may be read using R PAR B or R PAR C. See the READ command documentation for more details. 2. For high symmetry structures, such as those with NaCl structure, use the command STRUCTURE to generate the shells. C ATOM[1,3] CA;C ATOM2 O;C T1 O;C T2 CA Sets up 7 shells for the Ca edge of CaO - only the Debye-Waller terms A[1-7], the edge position EF and various multiple scattering
parameters (see below) need then be changed in order to fit the spectrum. 3. Enter the data by hand. Although not essential, it is best to define all the atoms that are to be used before defining the structure. One or more central or
excited atoms are required (ATOM1 is always the central atom of the first cluster). Each scattering atom should also be defined
(a scattering atom for the same element as the central atom is normally required, as excited and scattering atom phaseshifts are
calculated slightly differently). For a copper protein, the atoms required might be as follows: C ATOM1 CU;C ATOM2 C;C ATOM3 N;C ATOM4 O;C ATOM5 CU ATOM variables are described more fully below. To set up the structure 'manually', first define the number of shells: Then define the characteristics of each atomic shell. C S1 You will be prompted for the following (there are no defaults in EXCURVE at present, the values in brackets are typical preset
values) N1 [4] Shell occupation number (must be the multiplicity generated by the point group if the symmetry is to be defined). T1 [2] Element to be used for EXAFS phaseshifts. If phaseshifts have been read, or ATOM parameters have been defined, use an element symbol. Otherwise, use unique numbers > 1 for each element. R1 [2.4] Shell radius in angstroms. A1 [0.005] 2 x 2nd cumulant (Gaussian term in DW factor) - 2σ2 Å2 B1 [0] 10 x 3rd cumulant (Skewness of DW factor) - Å3 C1 [0] 100 x 4th cumulant (Kurtosis of DW factor) - Å4 D1 [0] Truncated exponential term - Å UN1 [0] Unit number (for group-fitting, group MS) ANG1 [0] angle 0-n-1, where n is the shell nearest the centre or the pivotal atom of a unit. Usually undefined
for shell 1, which is itself normally the nearest shell to the centre. ANG is 0 for shells not associated with units (see below and in
the parameters section, chapter 4). TH1 [0] spherical polar coordinate θ (0 to 1800) PHI1 [0] spherical polar coordinate φ (0 to 360) Y1 [0] Cartesian coordinate Y Z1 [0] Cartesian coordinate Z Normally only the first few parameters are entered. After sufficient information has been entered, "=" will terminate the command
("=" is often used for such a purpose in many commands). If the spherical polar coordinates R, TH and PHI are used, the cartesian
coordinates X, Y and Z are not generally used. Changing any one of the polar coordinates will update all the cartesian coordinates
and vice-versa. This should be repeated for each of the atomic positions. Once the positions are defined, the defaults are the existing values.
Normally, however, CHANGE Sn will not be used again - it is usually easier to change the individual parameters: C X4 .7395 C T1 HG CHANGE S may also be used to duplicate shells for editing: C S2 1 Will copy shell 1 to shell 2. An element symbol may be used for Tn both here and when the C Sn format above is used, only once the ATOM variables have
been defined. Once a set of parameters have been entered it is wise to save them immediately: PRINT PAR The program will issue a prompt for a filename, of the form expara1.dat. If the file already existed, the a1 suffix will be incremented
to generate a unique name. If you wish to use another filename it can be entered. You will be asked to confirm whether an existing
file should be overwritten. Parameters can be listed using LIST Typical output looks like this: LMAX = 25.000 DLMAX= 6.000 TLMAX= 5.000 WP = 0.100 NS = 4.000 EF = -7.159 VPI = 0.000 AFAC = 0.914 EMIN = 20.000 EMAX =1519.880 EF1 = -7.159 VPI1 = 0.000 AFAC1= 0.914 EMIN1= -20.000 EMAX1=1519.880 Shell 0 N0 = 1.000 T0 = 1(CU) R0 = 0.000 A0 = 0.000 B0 = 0.000-1 0 Shell 1 N1 = 12.000 T1 = 2(CU) R1 = 2.556 A1 = 0.008 B1 = 0.000 1 0 Shell 2 N2 = 6.000 T2 = 2(CU) R2 = 3.615 A2 = 0.011 B2 = 0.000 1 0 Shell 3 N3 = 24.000 T3 = 2(CU) R3 = 4.427 A3 = 0.016 B3 = 0.000 1 0 Shell 4 N4 = 12.000 T4 = 2(CU) R4 = 5.112 A4 = 0.011 B4 = 0.000 1 0 Individual parameters are described in the parameters section, chapter 4. The two sets starting EF... are significant for multi-spectrum fitting. If only one spectrum has been read, only the first set need be used. The second set is still displayed however,
because they will effect the result if they have previously been changed. The final two columns in the table of shell parameters are
the cluster number (-ve for a central atom), and the unit number. UNITS, for use with molecular compounds such as enzymes,
are not often needed and are described later. If the table is very long, the first part may be displayed using, say, LIST 10 (the
default keyword SHELLS is still assumed). When changing parameters, values may be defined in terms of other parameters or arithmetic expressions C N1 4/3 is more accurate than C N1 1.3333 Similarly, for two shells at x and 2x C R2 R1*2 is better than entering a specific value, as if R1 has been refined, it will be stored with 6 or 7 decimal places. The program has to judge whether two numbers are equivalent in some situations, and therefore it is better to be unambiguous. The most difficult aspect of parameter definition involves the atom type T1 etc. T1 takes a numeric value which is associated with
a particular scattering phaseshift. Normally the cross-reference is defined by atom parameters. For example: C ATOM2 O C ATOM3 C C ATOM4 CU defines the atoms required for Cu(CO)3. ATOM1 is always the excited central atom of the first or only cluster. ATOM4 is defined
because Cu is present as a scattering atom in addition to an excited atom. If T1=2, T2=3 and T3=4, the first 3 shells are O, C and Cu respectively. Once the atom parameters are defined, it is usually
possible to forget about the numbers, and use element symbols for the atom types: C T1 O etc. If phaseshifts are calculated, they will be calculated in the correct order. If phaseshifts are read-in, it is however the users
responsibility to ensure the correct correlation between phaseshifts and ATOM or T parameters (there is as yet no automatic
checking in most situations). To check that the order is correct type: L PHASE (which will display all the atom variables, among other things) and L FILES (which will give any phaseshift files that have been read) The situation is more complicated if multiple edges are being fitted. If the core-hole lifetimes of all the edges are similar, a single
set of scattering phaseshifts may suffice. In most cases however, a separate set of scattering phaseshifts is required for each
spectrum. It is then essential to ensure that T variables refer to phaseshifts not only of the correct atomic number, but also to the
correct central atom type. The symmetry is defined using the command SYMMETRY which sets the point-group of each cluster in the material (often there
is just one cluster) A list of point group symbols (Schönflies notation) is given by: The list of available symbols is followed by a longer list including the equivalent international symbols and the multiplicities
associated with general and special positions. A few symbols differ from normal - C0v instead of C∞v etc.. Cx and Cy are settings of Ci where the principal axis is x or y rather
than z as in Ci (it is possible to transform the coordinates using ROT variables, but it is often useful to retain coordinates in a
familiar orientation). The use of Cx and Cy has been superseded by new code allowing multiple settings. SYM will request a point group SYM c4h for examples, will assign c4h to the first cluster The occupation numbers and solid angles must be consistent with the point group operator. Often several shells with the same
radial distance must be defined if atoms are not symmetrically related (i.e, the shell has a higher symmetry than the cluster as a
whole). This is most obvious with cubic lattices, where shells of 36 atoms must clearly be composed of at least two shells, one with
12 and one with 24 atoms. Sometimes one of several possible orientations must be selected. If the shell variables do not
correspond to a standard orientation, the program will attempt to find an appropriate alternative setting. If the setting number is
other than zero, the coordinates used for MS calculations, for the command DRAW, etc. will differ from those in the table of shell
parameters. The actual values used are displayed by SYMMETRY Normally the setting is defined automatically, but it is possible
to override this using for example: SYM c4h:1 For setting 1. The standard setting is 0. If STRUCTURE is used to define the shells initially, the point group will be defined at the same time (for example, Oh for NaCl
structures). Note that a subsequent use of structure will overwrite the existing coordinates, while symmetry will never change the
shell parameters. In the case above, it is assumed that there is one phase and one crystallographic site containing the excited atom. In practice, there
may be many phases and many sites containing the excited atom, either with full occupancy, or as a component of a mixed site. For each site containing one of the excited atoms, a radial cluster must be generated. Each cluster consists of an excited atom
and a number of shells. If only single scattering is being used, multiple clusters can be represented by adjusting the shell
occupation numbers, remembering to include a factor relating to the multiplicity of the sites containing a central atom. For multiple
scattering calculations this approach will not work, as occupancies must be integer, scattering between clusters must be excluded
and each shell must consist only of atoms replicated by the point symmetry of the cluster in question. The clusters in this case must
be identified by a cluster number, using the shell parameters CLUSn. CLUS parameters are the same for each atom in the
cluster. The cluster number is 1 for the first or only cluster. Excited atoms are signified by a negative value. Thus by default, CLUS0
is -1, CLUS[1-NS] is 1. The second cluster will have an excited atom with, say, CLUS5 = -2. Shells are normally arranged in blocks,
in order of cluster, thus successive excited atoms, with cluster numbers -2, -3 etc. will mark the start of a new cluster. The
occupation numbers of the excited atom shells must reflect the site multiplicity, and determine the relative significance of each
cluster (they should of course always add up to 1). Thus N0 is normally set to 1, but with two clusters, with the same frequency
in the crystal, each with the same excited atom type, its value should be .5. The situation becomes particularly complicated when
the excited atoms occur in mixed sites. See the section on multiple spectra for an example of a parameter table for multiple
clusters. Mixed sites enable the program to calculate the spectrum when either the excited atom or a scattering atom has partial occupancy
of a site. This includes the calculation of multiple scattering contributions associated with such sites. As the occupation numbers
for sites must be exactly that determined by the point group symmetry, mixed sites also provide a means of calculating the effect
of site vacancies. Mixed sites must be used if full multiple scattering calculations are to be performed for disordered systems. Mixed sites are defined using the command SITE. It is used to define a pseudo-element symbol. For example the symbol QA
may represent Cu.33Zn.67. These symbols are assigned -ve 'atomic numbers', starting at -1. The symbol, or its 'atomic number' may
be used wherever a normal element or Z-value is used within the program. To set up the example above SITE QA Will give rise to the prompt Enter element and percentage for component A CU 1/3 Enter element and percentage for component B: ZN 2/3 Enter element and percentage for component C: To check site definitions type SITE The site command will have defined three variables. Assuming this was the first use of the command, it will have defined PERCA1
(=.33333), PERCB1 (=.66667), and PERCC1 (=0.). The composition of the site may be changed by altering these variables. The
variables may also be refined. If PERCAn is refined, then PERCBn will be adjusted so the sum PERCA+PERCB+PERC is
constant, unless PERCB is also refined. This provides a mechanism for maintaining the stoichiometry of the site. In order to refine site vacancies, it is necessary to refine PERCB ( or PERCC for 3 component sites) rather than PERCA, which
should be set to 0. In order to use mixed sites in calculations, it is first necessary to define an ATOM type, as in C ATOM7 QA Atom type 7 may be then be used to define the shell variables (e.g. for shell 4). C T4 7 or C T4 QA (once ATOM7 has been defined) If phaseshifts for each central atom and each scattering atom are available, they may be read in using READ PHASE The order must correspond to that of the ATOM variables (if defined) and consistent with the atom types Tn defined in the
parameter table (as yet there is no checking in most situations). If no phaseshift files are available, they must be calculated. Phaseshifts should be calculated for: All the central atoms required for the XAFS experimental files that have been read in. All the scattering atoms Tn referred to in the list of shell parameters. The first step in a phaseshift calculation is to calculate embedded atom potentials. Before this is done, all the required ATOM parameters must be defined. For CuO: C ATOM1 CU* C ATOM2 O C ATOM3 CU The first stage is to set the method of calculating the ground state exchange energy of the excited photoelectron, as described in
the theory section. For this example we use: If the alternative X_ALPHA option is selected, the potentials are dependent on the parameters ALFn (see the theory section on
muffin-tin potentials, chapter 2). The potentials are calculated using: There is a prompt for the level of output: G (graphics), T (terminal output), M (charge densities) or C (continue without output) [C]: M is used only in order to generate charge densities for other programs. The other options are self explanatory. The next prompt requests the neighbouring atom type - phaseshifts are calculated individually using a different cluster for each.
There can be at most two atom types per cluster. Prompts are of the type: Atom: 1 (GA). Enter neighbouring atom [3 (O)] If there are several scattering atoms with the same atomic number, a number should be entered. Otherwise, a chemical symbol
may be used. If there are several different neighbours, it is often best to choose the lightest. Atomic potentials can be significantly affected by
the choice of neighbouring atom. If you are unsure of the neighbour, and do not wish to bias attempts to discover its nature, use
monatomic clusters (but where the neighbour is a ground state not an excited atom). The next prompt will occur for excited atoms only - the menu available will depend on the edge, for a K-edge it is as follows: Select Code For Exited Atom [1]: No Correction (0) 1S Core Hole (K-edge - relaxed approximation) (1) 1S Core Hole (K-edge - Z+1 approximation) (-1) A positive number will select atomic potentials calculated self-consistently in the presence of a core-hole (relaxed approximation).
A negative number will use the 'Z+1 approximation' of von Barth (sudden approximation). In general, the fully relaxed case should
be preferable at low energies. The best option at high energies is debateable. The program will then attempt to calculate the potentials. If the graphics option has been selected, the charge density and potential
functions will be plotted. If the terminal output option is selected, tables of charge densities and potentials will be displayed. In all
cases the charge density is integrated over the Wigner-Seitz sphere to give the apparent number of electrons in the atom. This
is compared to the atomic number Z. The program will also display values of V0, RHO0 and FE0 when the calculation is finished. V0 is the Muffin-tin zero, the energy
of the flat region in figure 1. This is the effective energy-origin of the theory in EXAFS. It is some way below the edge, which is itself
below the vacuum zero level. Ideally, the calculated FE0 will correspond to the Fermi energy, which should equal the difference
between the experimental edge and the vacuum zero (a negative number). It is unlikely to do so however except for metals with
free-electron-like conduction bands. RHO0 is the interstitial charge density. If several phaseshifts are calculated, the values of V0, RHO0, FE0 will differ for each two-atom cluster. This contradicts the muffin-tin model which depends on a common interstitial potential in the crystal. If the differences are large, the theory calculations for
different shells of atoms may be out of phase with one another, effecting the result. Ideally calculations should be performed to
give consistent values, although there may be instances of partial ionicity where inconsistent values give the best result, due to
their mimicking the effects of charge transfer. It is, however, possible to calculate all the potentials so as to give a common value of V0, RHO0 or FE0. It is also possible to refine
these variables. The option SET COMMON V, for example, will ensure that all potentials are calculated to a common value of V0.
(Do not type S CONST V0 - V0 is a parameter, which will be evaluated, so the command may be interpreted as S CONST -7 etc.).
The muffin-tin radii will be adjusted to give the required value. This method is to be preferred to using a cluster consisting of many
atom types. This could cause steps in the potential at the muffin-tin radius, resulting in spurious effects associated with the strong
scattering from potential steps. If whole-spectrum fitting is being used, i.e. if ATOMABS is set to ON, then a second set of excited atom potentials labelled
'ATOM0' will be generated. These are identical to the normal excited atom potentials, except that the excited state term, generated
when the phaseshifts are calculated, will be entirely real. They will be used to generate the atomic transition rates. Real potentials
are required because the trick of generating an imaginary potential to account for inelastic process in scattering, does not work
in the calculation of dipole matrix elements. Among other problems, it results in loss of charge conservation. Muffin-tin Radii and related Parameters The potential calculations make use of a table of values for muffin-tin radii (the radius of the touching spheres used in modelling
the potential in the solid). They also use parameters describing ionicity, exchange and Madelung corrections. The muffin-tin radii
actually used will be adjusted by the program if the common V0, or similar options are used (see above). They may also be
changed manually, or indeed refined against a well established model compound. Will change the value for MTR1 to 1.34 (suitable, for example, for Ga, Z=31). If it is necessary to use a different value when an atom is used as a neighbour to that used when the potential is being calculated,
either additional ATOM variables may be defined (with an MTR just used for overlaps) or the potential for each atom type may be
calculated separately: CA POT 3 For example. The values may not be refined under these circumstances. Other potential parameters that may be altered include: IONn, ALFn, CMAGn, where n is the atomic number. ALF is only used
in conjunction with the X-α ground state exchange scheme, in accordance with equation [19]. ION is the ionic charge. Generally
only +/- 1 is used as it is not possible to generate adequate anion charge densities for a free atom with a charge less than -1. Ionic
phaseshifts are not usually used - the neutral atom charge densities are more reliable, and the net effect of using ionic phaseshifts
is normally only a shift in EF. Multiple excited atoms: CALC POT assumes that ATOM 1 is an excited atom. If potentials are calculated individually, an excited
atom is indicated by using a -ve atom type (e.g. CA POT -1 would explicitly define atom type 1 as being an excited atom).
Alternatively (and preferably) an excited atom status can be indicated by, for example: C ATOM4 BR* LIST PHASE will indicate excited atoms with a '*'. Once so-defined, excited atom potentials will be calculated until scattering-atom
status is resumed after C ATOM4 BR. When multiple spectra are in use, it will be necessary to use one of these methods to ensure
that excited atom potentials are calculated properly. Phaseshifts are calculated using: this will calculate phaseshifts for all atoms for which potentials are available. Phaseshifts depend on only a small number of options,
they mostly depend on the potentials. One is the excited-state exchange term - usually the Hedin-Lundqvist method is used: SET EXCHANGE Hedin-Lundqvist (S E H) the alternative is Which uses a constant imaginary potential for the excited state. Other options include whether to use scalar relativistic corrections - that is to include corrections to energy terms of the order
of(E/c2), and corrections to the photo-electron momentum, but to ignore spin-orbit coupling and related terms. Although such
schemes were successful in LAPW band structure calculations, and it is currently fashionable to include them in XAFS, the
justification for them is poor. It may be better to ignore them by selecting: Another option is to select the contribution to inelastic losses below the plasmon threshold, as described by Quinn (1962). It is probably best to use this option as it helps reduce the discontinuity at the plasmon threshold energy. A more important parameter is the core-hole lifetime. These are normally taken from tables, compiled from PES data on half-widths
of peaks due to core excitations. There is some scope for variation however due to a number of factors: 1. Core widths are not totally immune to chemical factors. 2. Reduced experimental resolution is not explicitly accounted for, and looks rather like lifetime broadening (strictly it should be
Gaussian whereas lifetime broadening is Lorenztian). 3. The theory behind the imaginary contribution to the excited state potential is valid only for a rather limited set of situations (a
free electron gas in the plasmon pole approximation), and altering the core lifetime provides a good fudge if the electron inelastic
scattering term is wrong. 4. The accuracy of the tables used in the program is not guaranteed. With these factors in mind, tinkering with the core lifetime, or even refining it has some appeal. This may either be done, in
response to a prompt, when the phaseshifts are calculated, or the parameter CW may be used. When multiple edges are in use, different core lifetimes are required for different edges. Although a fix for problems occurring when
two edges (e.g. K and L3) of the same element are in use is described below, it is better to calculate sets of both scattering and
excited atom phaseshifts for each edge. In this case, instead of using the global parameter CW, the spectrum specific parameters
CWi should be used. The index i is incremented for every atom number specifying an excited atom, and thus relates to a block
of potentials or their associated parameters. It does not necessarily bear any relation to the spectrum index as used in READ EXP,
although it normally ought to do so. In the table below (given by LIST PHASE), CW1 would apply to atoms 1 to 3, and CW2 to
atoms 4 to 6. If parameters CWi are to be used, CW must be set to -1. 1 ATOM 47 (AG*) ION 0.00 MTR 1.437 ALF 0.6667 CMAG 0.000 XE 0.000 2 ATOM 35 (BR ) ION 0.00 MTR 1.396 ALF 0.6667 CMAG 0.000 XE 0.000 3 ATOM 47 (AG ) ION 0.00 MTR 1.442 ALF 0.6667 CMAG 0.000 XE 0.000 4 ATOM 35 (BR*) ION 0.00 MTR 1.396 ALF 0.6667 CMAG 0.000 XE 0.000 5 ATOM 47 (AG ) ION 0.00 MTR 1.442 ALF 0.6667 CMAG 0.000 XE 0.000 6 ATOM 35 (BR ) ION 0.00 MTR 1.396 ALF 0.6667 CMAG 0.000 XE 0.000 If the HL excited state option is not being used, then no inelastic losses are included. A term due to the electron inelastic losses
should then also be included at this stage (or in CW). This will normally add 1 to 6 eV to the effective core width. When the command CALC PHASE is executed, a prompt is issued: Do you require the atomic absorption [no] ? The default option will depend on the setting of ATOMABS, the response will normally differ from the default only if XANES
calculations are to be performed. The next prompt is for the core hole width, as discussed above. If CW (or CWi) is defined, that is the default. Otherwise, a
tabulated value is the default. If tabulated values are being used, a value rather higher than the default is normally appropriate.
Experience suggests that in the current version of the program, twice the default value will normally give the best fit. It is sensible
to check the tabulated value against published data, as the tables use approximate interpolated values only. Once potentials and phaseshifts have been calculated, they can be written to files using the PRINT command (PRINT PHASE).
They can be read in to later sessions using READ. The PLOT command can be used to examine phaseshifts graphically. P PHASE The EXAFS theory is updated in response to a number of commands, such as FIT, PLOT, PRINT. FIT also gives fitting statistics. An example, for Cu-foil, is given below. Fit Index: with k**3 Weighting 3.0023 R-Factor : with k**3 Weighting 27.8242 Amplitude of Experiment 65.0893 R(exafs) = 27.8242 Weight: 1.00 R(distances)= 0.0000 Weight: 0.00 R(angles) = 0.0000 Weight: 0.00 N(ind) = 61.2 Np = 7. Chi^2 = 73.7025E-6 If you wish to update the FT as well (for example, so you can use CP 1 after the next update), type FIT FT. Single scattering (SS) is always calculated using curved wave theory (CW). Multiple scattering (MS) may be calculated using
curved wave theory, or in the small atom (SA) or Rehr and Albers (RA) approximations. These are selected using SET THEORY
CURVED_WAVE (for CW) or SET THEORY SMALL_ATOM (for both SA and RA). There are angular momentum restrictions on the MS CW theory (DLMAX for double scattering, TLMAX for higher orders). Use LIST
DIMENSION to display the maximum allowed values. DLMAX also controls SA/RA theory, although it is only restricted to the
maximum of LMAX in this case. If the theory is changed to CW, DLMAX will change. It will not be changed if the theory is switched
back to SA/RA. This must be done manually to recover the previous result: CHANGE DLMAX LMAX By default, the 'small-atom' option, is equivalent to the Rehr and Albers full-matrix approximation (NUMAX=2). C NUMAX 1 Reduces the accuracy C NUMAX 0 Gives the original small-atom approximation (Gurman, 1988). Refinement of parameters so as to achieve optimum agreement with experimental data is the primary purpose of the program. The only command required is REFINE which, given a list of parameters to refine, will determine their optimum values and calculate
statistical errors at the minimum it locates. For example, to refine the variables EF, R1 and R2: REFINE 150 (no keyword, so LSR assumed, step parameter set to 150) Enter parameter name or "=" to skip EF Enter parameter name or "=" to skip R1 Enter parameter name or "=" to skip Least squares refinement using k**3 weighting Initial parameters 1 EF 17.240 0.00272 17.51203 2 R1 1.500 0.00002 1.50159 Enter : CONTINUE to refine interactively a number to edit, add to, or delete from the list 3 We forgot R2, 3 means that we want to add parameter number 3 to the list. Note that there is no default for the above prompt, its
just too easy to start the refinement accidently. Enter parameter name or "=" to skip R2 Least squares refinement using k**3 weighting Initial parameters 1 R1 1.500 0.00002 1.50159 2 EF 17.240 0.00272 17.51203 3 R2 2.922 0.00003 2.92553 Enter : CONTINUE to refine interactively a number to edit, add to, or delete from the list C (for continue) R1 = 1.50000 EF = 17.23991 R2 = 2.92244 Call 1 F 351.5847 R 177.353 F- 351.585 Rex 177.353 RD 0.000 Rx 0.00 636 27 Select Next,Exit,Complete,Stats or Plot N 33 (or just 33) Carry out 33 refinement cycles. The program will carry on altering the values of R1, R2 and EF until the fit-index reaches a
minimum or the specified number of steps has been carried out. In this case the refinement should end with: R1 = 1.95281 EF = 21.12186 R2 = 2.87818 Call 50 F= 19.9091 R= 44.305 F(min)= 19.909 R(ex)= 44.305 R(dis)= 0.000 Minimum Predicted, Accuracy = 0.2817E-02 Cond = 2 At Call 50 Filename for statistics: [excora1.dat] ? The program prints 2σ errors and correlations along with the best values of the parameters at the predicted minimum, after
requesting a filename (default excora1.dat etc.) to provide a permanent record of the information (it also goes into the logfile). Type
'=' if you do not require a statistics file. The weighting used by the refinement is that same as that used by FIT, and can be altered using the SET option WEIGHTING. The command uses numerical estimates of the derivatives whose values can depend on the step parameter. Convergence is often
poor and refinements using a range of values of the step parameter are often required. A useful method of improving convergence
is to refine different sub-sets of the variables required separately. Refinement can often lead to physically unrealistic parameters, either as a result of underdeterminacy, strong correlations
between parameters (as revealed in the correlation matrix produced as above), or because other parameters are wrong. For
example, if EF is wrong, but only A parameters are being refined, they will refine to high values, as the errors due to the phase
being wrong are reduced if the amplitude is reduced. The program uses soft constraints to reduce this effect. If a parameter is
beyond a certain limiting value, there is a contribution to the fit-index proportional to the square of the excess. The most useful
constraint is for A parameters. The limits can be set using DWMIN and DWMAX. C DWMIN .001;C DWMAX .03 Will in general keep A parameters between .001 and .03 during a refinement. Multiple parameters may be refined using the list form for parameters (e.g. A[1-3,5]). Parameters may also be linked to refined
variables using rules. RULE R2;R1*2 REF;R1;C;C; Will refine R1 and R2 (R2=2*R1) A full list of options is presented in the command reference section on the REFINE command. If you wish to compare the results of several refinements, SAVE will store both the parameters and their errors. TABLE may then be used to display all the sets of parameters, with or without their errors. The wide range of plotting commands are described more fully in the command reference sections for PLOT and COPLOT. Amongst the more useful are: PLOT EXAFS To plot the EXAFS spectrum To plot EXAFS and FT together Other options include those to plot previous generations of theory with either EXAFS alone or in conjunction with the COPLOT
command. P EX 1 (or just P 1) will plot experiment, theory and the previous theory. CP 1 is similar. Note however, that for this to work, the previous theory must have been updated via a CP, P FT, PRINT or FIT FT, not
just a FIT. A full list of options is given in the command reference section on the PLOT command. It is often necessary to save output as disc files. Parameters should be saved at regular intervals in case of program failure, or
in case an unintentional change results in loss of the correct structure (but note that there is an UNDO and a RECOVER facility
to cope with many situations). Phaseshifts should be saved to avoid having to calculate them each time and to ensure consistency
in calculations. It is also useful to keep records of spectra, in case it is necessary to reproduce the results at a later stage, or to
compare them with later calculations. Most of all, spectra are required as input to graphics packages and word-processors. Files
for input to other programs may also be required. All these forms of output may be produced using the command PRINT. Many other commands can produce files. One obvious example is TABLE whose purpose is to produce formatted output for display
or for inclusion in papers. REFINE produces files containing best-fit parameters, correlations and errors. EXTRACT produces files
containing details of multiple scattering paths, and a filtered sum of paths. All spectra may be written in column format for use with other plotting packages. Parameters may be written in Brookhaven or
EXCURVE format, again using PRINT. Disorder is introduced by means of shell parameters A, B, C and D. A, B and C are the second, third and fourth cumulants (see the theory section on disorder. A is actually defined as 2xC2 (or 2σ2), B as 10xC3 (Å3) and C as 100xC4 (Å4). Two schemes are in use to calculate theory. In one the integral in equation (26) is calculated numerically (SET DIS EXACT). This
will take into account spherical wave effects. The other (SET DISORDER APPROX) uses a conventional Debye-Waller term as
given by equations (26-27). When only the second cumulant term is used, the difference between the two methods appears to be
very small in the majority of cases. The approximate method is recommended therefore on the grounds of efficiency. A is always the dominant term, but in some situations other terms are significant. Solids near their melting points are an obvious
example. The effect in k-space of a distribution with a given set of cumulants can, in principal, be calculated accurately, provided
spherical wave effects are ignored. In order to generate a pair distribution function for a given set of cumulants, however, a specific
model is assumed - the anharmonic oscillator. In this model, the allowed value of higher cumulants is determined by the values
of the lower ones. Thus in general, most values for the higher cumulants will not generate sensible values. For this reason the
EXACT option is certainly not recommended when higher cumulants are in use. One situation where the two methods should give
similar results, is if the values are commensurate with the anharmonic oscillator model. That is if: (42) Where α is the coefficient of linear expansion. It is possible to make use of this in determining the best values of the third cumulant.
The model works very well for the first and fourth shells of Cu at reasonable temperatures (Edwards et al.,1997). The program will
calculate B values automatically using this relationship provided both the coefficient of linear expansion (CLE) and the temperature
(TEMP) is defined. CLE is actually 106 x the coefficient, TEMP is in K. Use of this method avoids a big problem with using B parameters - they largely effect the phase, and are strongly correlated with
R and EF, hence may lead to a good fit being obtained with erroneous distances. A second scheme for treatment of disorder involves using A and D parameters to describe the convolution of a Gaussian with a
truncated exponential. This model is appropriate for cases where a cumulant expansion does not rapidly converge, as in ionic
conductors. D parameters should not be used in conjunction with B and C parameters. Where an atom occupies several sites, it is not unusual for it to have two different oxidation states. An obvious example is staurolite
(Fe2,Mg)2(Al,Fe3)9O6(SiO4)4(O,OH)2. When this occurs, the chemical shift of the absorption edge (caused by differences in the
screening of the nucleus by the different valence shells) will result in the associated components of the spectrum being out of phase
at low energy. This can be accommodated by the atom-type dependent terms XEn, where n is the atom number associated with
Tn. This is a correction to EF for each atom-type. It will be necessary to calculate a second set of phaseshifts for the central atom
and each scattering atom, and to specify two clusters, about Fe2 and Fe3. The central atom of the second cluster should have
CLUSn parameter of -2. There should be only two values for the XE parameters, i.e. all values associated with the same cluster
should be the same. (The combined XAFS/PD program P more sensibly uses a cluster-dependent XE parameter, preventing the
use of XE's as general fudge factors). There are other, and possibly better ways of treating multiple oxidation states, for example
calculating two sets of phaseshifts with different V0 values. When two or more EXAFS spectra are in use, variables such as EF, AFAC, EMIN, EMAX etc. are unlikely to be the same for both.
For these variables, as well as a common variable, which applies to all spectra, there are individual variables which apply to only
one. These are indexed by spectrum number, as used in READ EXP. In some instances the common variable may be more
appropriate - for example EF for different polarisation directions. It may also be useful to combine common and individual variables
- AFAC may be the same for all but one of the spectra. The rules for combining variables, in the case of spectrum i is as follows: AFAC The program uses AFAC*AFACi EMIN The program uses MAX(EMIN,EMINi) EMAX The program uses MIN(EMAX,EMAXi) CW The program uses CW unless CW=-1, when it will use CWi unless that is -1, but note that i is not a spectrum
index here, it marks a 'block' of atom numbers which commences with an excited atom (see calculating phaseshifts in this chapter). BETH There is no BETH - it uses BETHi (similarly with BEPHI, BEW) The common variables may also be written, for example, EF0. This can be useful in resetting variables. e.g. C AFAC[0-3] 1 If only one spectrum has been used, the indexed variables can be ignored. If you are working with one spectrum, but a parameter
file has been read which has been used with multiple spectra, check that the indexed parameters do not affect the result. To
facilitate this, commands such as LIST will display the first spectrum indexed variables even when they are not required. Differences in core-hole lifetime normally mean duplicating the scattering phaseshifts, using different values for the effective core-hole widths. It is also possible to use a single set of scattering phaseshifts, and to compensate for differences using spectrum
dependent values of VPI. Shell parameters for multiple spectra will normally require two clusters (see Reading the spectra in this chapter, and READ in
Command Documentation for other possibilities). An example is shown below for AgBr (NaCl structure, Oh symmetry at each
atom). Here cluster 1 is centred on Ag and cluster 2 on Br. Only one parameter can vary in this case, the cell parameter, so only
ACELL is refined (it just scales all the distances). Note also that Ag-Br and Br-Ag Debye-Waller terms must be the same, so these
are refined as A[1,6], A[2,7] etc. The atom types are defined in the table above - see Calculating the phaseshifts in this chapter. Shell 0 N0 = 1.000 T0 = 1(AG) R0 = 0.000 A0 = 0.010 B0 = 0.000-1 0 Shell 1 N1 = 6.000 T1 = 2(BR) R1 = 2.867 A1 = 0.008 B1 = 0.000 1 0 Shell 2 N2 = 12.000 T2 = 3(AG) R2 = 4.054 A2 = 0.011 B2 = 0.000 1 0 Shell 3 N3 = 8.000 T3 = 2(BR) R3 = 4.965 A3 = 0.012 B3 = 0.000 1 0 Shell 4 N4 = 6.000 T4 = 3(AG) R4 = 5.734 A4 = 0.019 B4 = 0.000 1 0 Shell 5 N5 = 1.000 T5 = 4(BR) R5 = 0.000 A5 = 0.010 B5 = 0.000-2 0 Shell 6 N6 = 6.000 T6 = 5(AG) R6 = 2.867 A6 = 0.008 B6 = 0.000 2 0 Shell 7 N7 = 12.000 T7 = 6(BR) R7 = 4.054 A7 = 0.011 B7 = 0.000 2 0 Shell 8 N8 = 8.000 T8 = 5(AG) R8 = 4.965 A8 = 0.012 B8 = 0.000 2 0 Shell 9 N9 = 6.000 T9 = 6(BR) R9 = 5.734 A9 = 0.019 B9 = 0.000 2 0 When refining multiple spectra, it may be necessary to weight each one differently, to prevent the spectrum with the highest
amplitude or the best signal to noise from dominating the fit. This is done by the weighting terms WEXi. There default value is 1.
They may be adjusted to alter the relative weightings of the spectra. Ideally the sum of the terms in use should equal the number
of spectra being used. e.g., for three spectra the scheme might : C WEX1 .5;.C WEX2 2;C WEX3 .5 Multiple scattering is best implemented using the option SET MS ALL This calculates all the paths within all the clusters, up to a maximum pathlength given by PLMAX. Paths may also be limited to
those involving certain angles by MINANG and by magnitude using MINMAG. These and other related parameters are described
in the parameter documentation, chapter 4. Their effect can be seen using EXTRACT (q.v.). PLMAX should be set to at least twice
the distance of any strongly 'shadowed' atom - for example twice the fourth shell distance in Cu - this is higher than the default
distance of 10Å. By default, only 2nd and 3rd order paths are calculated (those involving 2 or 3 scattering events). It is possible to increase the order
of scattering to 4 or 5 however by setting OMAX to 4 or 5. This still leaves a restriction that there can be no more than 2 different
scattering atoms. This is relaxed by setting ATMAX to 3, 4 or 5 rather than 2. There are two options for the choice of theory in multiple scattering calculations: The default option is the approximation due to Rehr and Albers (1990). The alternative is the curved-wave theory (Gurman, Binsted
and Ross, 1986) as used for single scattering. The former is implemented in three levels, using the parameter NUMAX to designate
different matrix sizes. The minimum value, NUMAX=0, corresponds exactly to the small-atom theory (Gurman, 1988). They are
selected using: SET THEORY Curved_wave or SET THEORY Small_atom The parameters which control the number of angular momentum terms in expansions about scattering centres are very important.
Strictly, the same value of LMAX should be used as for single scattering (typically around 25 for Cu foil). In practice, it is often
possible to use much lower values if only light element scatterers are present and only low energies are required. The values
required must be determined by inspection. The use of too low a value will result in MS amplitudes at high energy which are too
high - sometimes an order of magnitude too high. The energy range is very critical, and whereas an LMAX of 12 may be
acceptable at, say 800 eV, it may be far too low at 850 eV. For the curved_wave theory, there are absolute limits on angular
momentum values of 12 for double scattering paths, and 9 for higher order paths. The values used are controlled by the parameters
DLMAX and TLMAX respectively. For the small atom theory, DLMAX is used to control all orders of scattering and it has the same
limit as LMAX (usually 25). With NUMAX=2, the small atom theory is almost as accurate as the curved wave theory. Although it
is not significantly faster for low values of DLMAX, it is much faster for DLMAX>=12. The errors due to restricting DLMAX to lower
values are in most instances much greater than using the approximate theory. The small atom theory, with NUMAX=2 is therefore
recommended for all but near-edge calculations. There are angular momentum restrictions on the MS CW theory (DLMAX for double scattering, TLMAX for higher orders). Use LIST
DIMENSION to display the maximum allowed values. DLMAX also controls SA/RA theory, although it is only restricted to the
maximum of LMAX in this case. If the theory is changed to CW, DLMAX will change. It will not be changed if the theory is switched
back to SA/RA. This must be done manually to recover the previous result: If you wish to compare the two theories, using a similar angular momentum basis, you should first restrict DLMAX using: CHANGE DLMAX TLMAX To revert to maximum accuracy, DLMAX should be reset, whichever theory is used. In order to inspect the current values of multiple scattering parameters, it is best to use the command Typical output is: OMIN 1.000 OMAX 3.000 PLMIN 0.000 PLMAX 10.000 MINANG -1.000 MINMAG 0.000 DLMAX 6.000 TLMAX 5.000 ATMAX 3.000 NUMAX 2.000 OUTPUT 0.000 It is important to try to ensure that important paths do not move in and out of the range allowed by PLMAX during refinement, as
this will prevent convergence. There is a mechanism (EXTRACT FILTER) to 'fix' the set of paths to be calculated, but it is unreliable
at present. The mechanism for calculating only the most important paths using MINMAG relies on the path indices (entries such
as 7.02 in the table generated by extract) being constant. The set of path indices is always slightly greater than the number of
entries calculated. The indices will remain constant, provided that the number of whole shells does not differ between calculations.
If distances are altered to the extent that they differ by more than 5%, if the radii of two shells near to the path-length limit differ
by less than the maximum change in bond length, or if the number of shells, order of scattering etc. are changed, then using
MINMAG will give erroneous results. MINMAG is best used when only EF, AFAC and A parameters are varied, at least by novice
users. If distances are to be included, they should be refined using a high value of the step parameter in REFINE. path overflow: Although the number of paths which may be included in a calculation is not limited (the record is about 2 million),
the number that may be stored is limited (the number is displayed at the start of the program). If the path dimension is exceeded,
the entries will not appear in EXTRACT tables, and MINMAG and FILTER must not be used. The mechanism for rapid refinements
when only AFAC and A parameters are varied will also fail (results will still be correct, but the calculations may be hundreds of
times slower). If the path-number dimension, is exceeded, then EXTRACT will be valid but MINMAG and FILTER may not be
used, and fast refinement of A and AFAC will not work. site vacancies: Special difficulties arise when using site vacancies (see mixed sites) to model disordered sites. In this case the
structure may contain two adjacent partially occupied sites which are very close to each other. In this case multiple scattering paths
are required between the partially occupied sites except for mutually complementary sites which together represent a single atom.
The interatomic vectors between such sites are usually small and may be restricted by use of the variable MINDIST. A path,
whatever its length, is excluded if any single leg is less than MINDIST. Sometimes it is useful to subdivide the structure to facilitate either model-building or refinement. Usually this situation occurs with
molecular compounds, where specific ligands are treated as separate entities. When building a model it is useful to be able to add
or subtract an entire ligand, and it may also be useful to treat it as a more or less rigid entity during refinement. In addition, although
it may be quite easy to determine which ligands are present and their distance from the central atom, it may be more difficult to
determine the relationship between them in three-dimensions. As multiple scattering paths tend to be strongest when they include
bonded distances, including intra-ligand paths but excluding inter-ligand paths may allow a reasonable fit to the spectrum, without
having to develop a complete structural model. In EXCURVE parts of the structure treated in this way are called units. They are
identified by a common unit number shared by their constituent atoms. The unit number of atoms which are not part of a unit is
0. A unit may be defined using, for example: This defines unit 1, which may be, say, a CO3 group. Units themselves have a number of parameters associated with them. If the position of a ligand is to be moved without altering
its bonding distance, then it must be rotated about one of its atoms. The atom about which a rotation takes place is called the
pivotal atom. It can be defined by a unit parameter (that is, a parameter indexed by unit number) called PIVn (n is the unit
number). If PIVn is undefined, then whenever a rotation is performed, the atom nearest to the central atom at the time is used.
Leaving PIV parameters undefined is dangerous, as refinement may at least temporarily alter the nearest atom, resulting in loss
of coherency of the unit. In most circumstances the pivotal atom is part of the unit. In a few instances it may be outside the unit.
For example, in P(Ph)3- compounds, it may be a good idea to define each phenyl group as a unit, but the bonded P atom as the
pivotal atom. If a unit is to be moved nearer or farther from the central atom, one atom must be selected for translation. If the pivotal
atom is both defined and within the unit, this atom will be used. Otherwise the nearest atom to the centre at the time will be
selected, giving rise to the problems seen above. Each atom in the unit, other than the pivotal atom, has an angle associated with
it which is expressed in the parameter ANGn (n is of course a shell index, not a unit index). For some purposes, the plane of a
unit is important. As units of more than three atoms cannot in general be exactly planar, three atoms are selected to define the
unit plane - PIVn, plus two more atoms defined by PLAn and PLBn (the program will select atoms if PLA/B are undefined). A unit
must contain equal occupation numbers for all shells. The occupation numbers may however be more than one - the unit may
be duplicated by the point symmetry. Constrained refinement uses rigid body constraints. That is, it involves movement of a complete unit, maintaining all of the
interatomic distances and angles with the exception of those associated with the central atom. The only occasion on which the
geometry might change, is when the central atom is itself part of a unit, as constrained refinement never results in movement of
the central atom. Constrained refinement is turned on by setting the option constraints to on. Once this is done, various other
parameters can be used to manipulate the unit. In addition to the parameters describing the coordinates of individual atoms within
the unit, there are three rotational parameters -ROTn, TWSTn and TILTn. Formal definitions are given in the parameters section
(chapter 4) - note however that they are shell parameters not unit parameters. Two types of motion are possible: A. Movement of the pivotal atom (see the definition above), either along a vector passing through the central atom (if X,
Y, Z or R for the pivotal atom are changed), or by means of a rotation about an axis passing through the central atom (if TH, PHI,
ROT, TWST and TILT for the pivotal atom are changed). B. Movement of one of the atoms other than the central atom or pivotal atom. In every case this involves a rotation about
an axis passing through the pivotal atom while the position of the pivotal atom itself is unchanged. Type a motion: Movement of type A is achieved by changing or refining the shell variable associated with the shell for the pivotal atom. e.g., if
the pivotal atom for unit 1 is in shell 7, then a command might be C R7 2.2 or (if the pivotal atom is the pivotal atom) C R[PIV1]
2.2. The latter form removes the need to remember the index of the pivotal atom, which might change, after a SORT or after new
atoms have been added. The rules governing motion in each case are (c is the central atom, p is the pivotal atom, u is the vector
c-p - see the parameter section, chapter 4 for more precise definitions): R motion is along the vector c-p so that the pivotal atom has the new R value requested. TH the unit is rotated about an axis through c and normal both to u and the z-axis, so that the new value of theta for the
pivotal atom is achieved. PHI the unit is rotated about and axis though c and parallel to the z-axis so that the pivotal atom has the new coordinate
requested. X the unit is translated along a vector through c parallel to the x-axis. Y translation parallel to the y-axis. Z translation parallel to the z-axis. ANG as the c-p-p angle is undefined (0) this is not a valid option. ROT the unit is rotated about the vector defined by VECA and VECB. The user must ensure that VECA refers to the central
atom and VECB is neither 0 nor the pivotal atom. TILT the unit is rotated about the tilt axis nXu by the change in value. TWST the unit is rotated about n the normal to the unit plane. All these type A motions are straightforward if the central atom is at the centre of the coordinate system (0,0,0). If this is not the
case, motion is more complex as the parameter values to be satisfied are the absolute values not the values relative to the central
atom. Under these circumstances, some values or PHI may be unobtainable, and the program does not allow TH to be changed
at all. Note that none of these type A motions will effect the single scattering or intra-unit multiple scattering EXAFS except for R. Type B motion: Motions of type B are achieved by changing any shell variables associated with the unit other than those of the pivotal or central
atoms. The atom given by PLA may often be appropriate.e.g. C R7 2.2 or C R[PLA1] 2.2. Note that C PANG1 235 is exactly equivalent to C ANG[PIV1] 235. The rules governing motion in each case are: R The unit is rotated about the tilt axis uXn TH the unit is rotated about the tilt axis uXn PHI the unit is rotated about n the normal to th unit plane X,Y the unit is rotated about n Z the unit is rotated about the tilt axis uXn ANG the unit is rotated about uX(s-p), where s is the vector defining the shell in question ROT the unit is rotated the axis defined by VECA and VECB. TWST the unit is rotated about n TILT the unit is rotated about the tilt axis uXn Sometimes, constrained refinement is required only for a single operation. It becomes tedious to switch it on, change a parameter
and switch it off again. It is also normally disastrous to perform an operation when the constrinaed refinement status is not what
you thought it was. For these cases, a single command CCHANGE (constrained change) is provided (see the command
documentation, chapter 5). A much simpler scheme for fixing one parameter in terms of another is to use the command RULE (see chapter 5). This allows,
for example, the 4th shell to be maintained at twice the first shell distance. It can be used for any parameters, not just coordinates. The parameters ACELL, BCELL, CCELL can be used to scale all the coordinates, as is appropriate for multi-shell fits to high
symmetry solids, where shell ratios must be preserved to obtain a meaningful result. See the section on cell parameters in chapter
5. Restrained refinement is, like constrained refinement, a system for ensuring the coherence of well defined elements in the
structure. It differs in that some distortion of the ideal distances is allowed, and there is no need to define units, or ensure that shells
to be coupled have similar occupation numbers. It simply defines a set of preferred distances, which given suitable weightings will
remain more or less fixed during refinement. The procedure is formally defined in the theory section, chapter 1, and examples are
given in the paper by Binsted, Strange and Hasnain (1990). To get started: Select a relative weighting for EXAFS and restraints: C WEX .5;C WD .5 Set up some ideal distances: C D2:1 1.2;C D3:2 1.1;C D3:1 1.3 and some bond-weightings C W2:1 1;C W3:2 1;C W3:1 1 FIT will display actual and ideal distances, and statistics. CHANGE (in chapter 5) and the parameter section (chapter 4) describe
the variables and easy ways of changing them for complicated structures. Metallo-proteins - amino acids, Brookhaven files and torsion angles A number of options are provided primarily for use with proteins. These include additional unit parameters such as a unit name
UNAMEi, torsion angles, to allow residues to be moved while the protein backbone remains intact. There are also easy ways to
set up structures, either by reading from files in Brookhaven database format, reading .car files from the Cerius Explorer modelling
program, or using a ligand database based on PROTIN restraint files. Additional unit parameters, as well as PIV, PLA, PLB etc.
are defined if units are set up in this way. These parameters are most easily edited using the command PLOT UNIT n which has
an edit facility. This is not usually the best way of displaying individual, complex units however, DRAW UNIT n is better, but does
not have an edit facility. Torsion angle refinement Torsion angle refinement, like constrained refinement, involves simultaneous movement of atoms within a unit. In this case
however, three of the four atoms defining the dihedral angle remain stationary while the others rotate until the desired angle is
obtained. Three torsion angles may be defined. By default one (TORC) is defined by atoms c-p-a-b, while the others (TORA and
TORB) are undefined except when a Brookhaven ( or .pdb ) file is read, when the atoms N-CA-CB-CG and CA-CB-CG-CD are
used. The atoms defining the torsion angle for a unit may be changed at any time using PLOT UNIT which allows interactive
changes to unit parameters. It is not relevant whether CONSTRAINTS is on or off when changing or refining torsion angles. Brookhaven files Brookhaven format files (often .pdb files on Unix systems) may be read using READ PARAMETERS BROOK (R PAR BR). The
atom number of the central atom is requested, which allows atoms within MAXRAD angstroms of the central atom to be read. If
these include atoms within many-atom residues, the whole residue is read. The variable MAXRAD is set to 5 Å by default. It may
be changed if required. Brookhaven format files may be written using PRINT PARAMETERS BROOK. It is also possible to update
a file, for which the EXCURVE parameters are a part, using PRINT PARAMETERS REPLACE. This is only possible if the same
file has previously been read, as the atom numbers must correspond. Cerius Explorer files Cerius (.car) files, contain similar information to .pdb files, they may be also be read using: R PAR CAR. The ligand database resides in the file ideals and is based on the database used by the crystallographic refinement program
prolsq. It has been extended to include non-amino acid ligands and information for defining the default relationship to the central
atom for use in EXAFS analysis. In order to use the database the shell for the pivotal atom and the coordinates of the pivotal atom
should be selected. E.g.: C X3 1.3;C Y3 1.3;C Z3 0. The ligand can then be selected using: C T3 (ligand name) The ligand name may be an amino acid symbol (TYR, HIS etc.) or a chemical formula (CO3, SO4). A list of current names may
be obtained using: C T3 ? The ligand will be assigned to the next vacant unit number. The pivotal atom will be assigned to the specified shell, and shell
numbers above NS will be used for the other atoms. The ligand name will be used for the unit name. The unit parameters PIV, PLA,
PLB, will be set up, as will any relevant torsion angles. Bond angles and torsion angles will be defined from database information.
The only undefined aspect of the ligand is the rotation about the vector u. This may be changed using the ROT parameter with
VECA=VECB=0. As ROT has no absolute significance it is not possible to use the current setting to define the orientation. This
coordinate has no effect on the theory unless inter-unit multiple scattering or XANES calculations are performed. Idealisation and the Spin command Because protein crystal refinements have been subject to generally weaker constraints on the interatomic distances and angles
within amino-acid side-chains than is usual with EXAFS, it may be desirable to idealise the side chains. This is done by replacing
them with amino-acids from the ligand database. The command used to do this is IDEALISE. The syntax is: idealise ([spin/nospin]) unit-number The optional keyword determines whether the spin command (described below) is also executed. If no unit number is specified,
all units are idealised. Idealisation assumes that all four main-chain atoms (N, O, C, CA) remain unchanged and that the torsion angles TORA and TORB
remain unchanged. These two assumptions mean that the cumulative effect of small changes in interatomic distances, angles and
torsion angles may result in very large movements of the atoms coordinated to the metal atom. The metal-ligand distance, the
principal angle, and the torsion angle TORC can be optimised, without changing other distances or angles, using either the
command spin or map. In each case TORA and TORB are varied. It is also possible to perform a general rotation about any bond,
although unlike changing a torsion angle this will also result in changes to the main-chain atoms. A rotation can be achieved using
a ROT parameter, after setting VECA and VECB to the correct shells numbers. The command spin has the syntax: spin [A or B or R]([long or short]) unit-number e.g., spin along 2, spin blong 3 A signifies TORA, B TORB and R a general rotation ROT. long or short determine the amount of output. The unit number must be specified, and the unit parameters piv, pla and plb must be defined. The command requests an optimum bond angle and central atom torsion angle, and determines the optimum value of TORA, etc.
to achieve both these angles and the shortest metal-ligand distance. One problem with spin is that the torsion angles cannot in general be optimised sequentially. An alternative is to use map to find
the optimum value. To do this restrained refinement is used with the optimum distance (and/or angle) specified and with WEX
a little higher than 0 and WD close to 1. TORA and TORB for the unit in question are used as map parameters. The analysis of both single crystal and surface spectra makes use of the polarisation dependence of the technique. As the program can analyze both polarised and unpolarised spectra from the same sample simultaneously, it is necessary to
specify how each spectrum is to be treated when it is read. Theory calculation can be considerably speeded up if several spectra
can be calculated together. This is only possible if they are of the same edge, have the same energy range, and share a common
value of EF. If this is the case, they may be treated as members of a polarisation set. The sequence number in each set is
requested by READ EXPERIMENT. Sequence number in polarisation set [0] Entering 0 indicates no polarisation dependence 1 is the first or only member of a set 2 is a spectrum equivalent in every way except in terms of beam parameters Once polarised spectra have been read, polarisation dependence is initiated using SET POLARISATION ON or SET
POLARISATION EXACT and setting the appropriate beam parameters. ON simply takes the ratio of the amplitudes for the
polarised and unpolarised cases in the small atom approximation. EXACT calculates the directional effects arising from angular
part of the dipole matrix element for the full Z-matrix. It is much slower, and for multiple scattering calculations will only work at
present for shells with single atoms (use EXPAND). For both single and multiple scattering, one effect of using ON is to ignore
spherical-wave effects which are important very near the edge. It is exact, but totally useless, for cubic point groups ! In addition,
for multiple scattering, the polarisation dependence is approximated by a scattering factor depending only on the angle between
the first and last leg of the scattering path (γ). It is quite accurate for γ ~ 0, but very inaccurate for γ ~ 90. At γ = 90, it is zero,
however important the path is. The approximation can therefore be extremely poor, for some paths. The EXACT polarisation
dependence makes use of the Curved Wave theory option, and so is restricted in terms of angular momentum values in the same
way. It is most useful in fitting the edge region, where curved wave effects are largest. It is however extremely slow. When performing polarisation dependent calculations, two or three spectra will normally have been read, each referring to a
different beam or polarisation direction. In this case, there are a set of beam parameters associated with each spectrum. The beam
parameters are defined in terms of the cluster coordinates by BETHn and BEPHIn (beam θ and φ parameters) and an azimuthal
parameter BEWn, the projection of the direction of the photon e-vector onto the xy plane. This system of reference is illustrated
in Gurman, (1988). Selection of the correct beam parameters is facilitated by the command: which displays in addition to the above parameters, the actual direction of the e-vector in both cartesian and polar coordinates. The
parameters have indices n given by the spectrum number, so if only a single spectrum has been read, the index is 1 (BETH1 etc).
Typical output from the command is given below. Spectrum 1 Spectrum 2 Spectrum 3 BETH1 = 0.000 BETH2 = 0.000 BETH3 = 90.000 BEPHI1= 0.000 BEPHI2= 0.000 BEPHI3= 0.000 BEW1 = 90.000 BEW2 = 0.000 BEW3 = 90.000 EX1 = 1.000 EX2 = 0.000 EX3 = 0.000 EY1 = 0.000 EY2 = 1.000 EY3 = 0.000 EZ1 = 0.000 EZ2 = 0.000 EZ3 = -1.000 ETH1 = 90.000 ETH2 = 90.000 ETH3 = 180.000 EPHI1 = 0.000 EPHI2 = 270.000 EPHI3 = 0.000 Domain averaging. For some surfaces, although the normal incidence e//x and e//y are in principal different, because of random
orientation of surface domains, no difference is seen in practice. In this instance it is necessary to average the theory over the two
orthogonal directions. This is implemented by the command SET AVERAGE ON. When this is done the first two spectra are
averaged - it is thus usually necessary to read one of the spectra twice - the theory will then be calculated twice, and the average
used for each of the first two spectra. The program does not ensure that the beam direction is normal to the surface or that the
two azimuthal angles are orthogonal as required so this must be done by the user. This procedure may also be used with twinned
crystals. Surface analysis may use a number of parameters not required elsewhere. The adsorbate layer is often defined by knowledge
of the coverage, the LEED pattern, and the lattice spacing of the substrate. In this instance, the atoms forming this layer need not
be refined. Below a few atomic layers, the substrate is also unlikely to differ greatly from the bulk phase, and may also be defined
by knowledge of the bulk cell parameters. Between these to regions it is possible to define a surface layer. In many instances,
refining the adsorbate-substrate distance, the relaxation of the surface layer, and reconstructions within it allows a large number
of shells to be fitted, using a small number of parameters, providing a rigorous test of a particular model of the surface. The adsorbate-substrate distance can be refined using DELTAZ which changes the Z-coordinates of all atoms below the XY plane.
It is not necessary to specify constrained refinement in order to use this. The surface layer is defined using ZMIN and ZMAX. The adsorbate is usually assigned coordinates with Z >= 0, with surface atoms
lying below the XY plane, with Z -ve. In this case, ZMIN and ZMAX will be of the order -1.5 and -.5 respectively (for a monatomic
surface layer). It is important that ZMIN and ZMAX are defined so that random changes in the coordinates during refinement do
not cause atoms to move in and out of the layer. Once the surface layer is defined, a simple model of surface relaxation may be tested by refining SURREL. This moves the
substrate relative to the surface layer. The initial value of SURREL is 1, so refined values of the order of 1.1 are likely. More complex relaxations may be achieved by refining the Z-coordinates of atoms in groups - e.g. Z[3,5,9,11], Z[4,6,8,12]. In order to treat reconstructions, it is also useful to define a surface cell. This is done using ACELL and BCELL. These parameters
should be defined before any of the atom coordinates, the reason being that changing them will alter the x- and y- values of all
the atoms in the cluster, even if constrained refinement is not selected. BCELL ( and for bulk crystals, CCELL) will reflect the point
symmetry of the cluster. Thus if the symmetry is defined as C4v, changing ACELL will also change BCELL. Once the cell is
defined, The parameters SDXn, and SDYn can be used to effect reconstructions. Although this approach may have limited
applications, it can be applied to some quite complex reconstructions. See for example Binsted and Norman [n]. Another type of transformation may be achieved using SURROT which rotates atoms about the origin of their surface cell. All these transformations are restricted by the requirement that they are compatible with the point symmetry of the cluster, and
therefore are of greatest use with clusters described by low symmetry. A simple demonstration of some of these options is given in the examples section of the manual (example 3). Further discussion
and examples appear in the Parameters and Related Topics section under surface parameters. Some typical steps surface analysis 1. Set up the initial parameters ( e.g. using PAXAS ) so the adsorbate atoms are at Z=0 or above. Ensure all the shells are
compatible with the point group. 2. Define the point group. 3. Change ZMIN and ZMAX to refine the surface layer: -1.0 and -0.1 will often suffice. 4. Set the beam polarisation parameters for each experiment to be analyzed. 5. Set SURFACE to ON. 6. Adjust the adsorbate-substrate distance using DELTAZ. 7. Adjust the surface relaxation using SURREL. 8. Adjust the bulk spacing using ACELL, BCELL, CCELL as required by symmetry (use ACELL[1,3] or ACELL[1-3] for isotropic
refinement in non-cubic point groups. 9. Adjust the displacements of the adsorbate and reconstruction of the surface layer using SDX and SDY. Whole-spectrum fitting is the analysis of the total absorbance spectrum, including both the atomic and scattering contributions,
over the whole energy range - from below the edge to the limit of measurable EXAFS. The procedure is selected using It requires atomic transition rates, calculated for real (rather than complex) potentials, so potentials and phaseshifts should be re-calculated after the option is selected. It also requires that EMIN is given a negative value, so the pre-edge region can be seen.
The data to be fitted should consist of absorbance vs. either eV above the edge, or absolute photon energy. If the latter, or if the
edge position is not accurately defined, the variable E0 can be used to generate the correct energy scale. The edge position so
defined should accurately reflect the position of the Fermi level when the spectrum is read. There is no suitable 'fudge factor' to
alter the edge position after it is read in. Other parameters required are TEMP - this defines the effective temperature used to define a T-dependent Fermi function, which
is convoluted with the edge. As this is also used to approximate lifetime and resolution, a value very much higher than the actual
temperature may be needed to match the experimental resolution. The parameters PRE0i, PRE1i, PRE2i correct for residual pre-edge background (the index i is only required for multiple spectra). The parameters POST0i ... POST8i and OFFSET (POSTn is a polynomial in (eV-OFFSET) correct the theoretical atomic transition
rate - this must be accurate to a few % in order for the EXAFS to be refinable - an impossible task theoretically. The procedure is discussed in more detail in Binsted and Hasnain, 1996. Results obtainable now that most of the improvements
recommended in the program have been introduced are much better however. The first step is usually to set up a startup file. This contains commands to read in spectra, set options that are invariably used,
read parameter and phaseshift files etc. It is not absolutely essential to do this as the commands can be entered at the terminal
in exactly the same format, but it is difficult to remember as well as tedious to enter a dozen or more lines every time the program
is used. A startup file may look like this ( you can include comments starting with ! ) !Set up some aliases for plotting hard copies !make sure alias names are otherwise incomprehensible to the program !T0 is 'plot the exafs as hard copy' alias T0 gs dev 2;p;;gs dev 1 !T1 is a hard-copy coplot alias T1 gs dev 2;cp;;gs dev 1 !T2 plots 3 spectra in separate frames alias T2 gs fr sel;p;1;2;3;;gs fr 1 !T3 plots 3 FT's in 3 frames alias T3 gs fr sel;p ft;1;2;3;;gs fr 1 !read in two EXAFS spectra r ex 1;ba2ino3f.ex1;1;12;BA K;; r ex 2;ba2ino3f.ex2;1;12;IN K;; !read in excurve parameters r par;expara1 !set the weighting for the refinement s w k3 !set a few phaseshift related options ft nd 2;s rel of;s ground x;s tort v !set the limits for DW factors c dwmin 0;c dwmax .03 !set the value of CW c CW 3 !calculate some phaseshifts ca pot;;;;;5;;;; ca ph;;;;;;;;;;; All this could be put in a file called, for example stba2 ( traditionally excurve command files start with st). To run it, start the program: excurve %stba2( run the startup file) You can then enter further input from the terminal. The bond valence sum, given by Brown as: (43)
is a useful check of the validity of cluster coordinates. It may be check using FIT BOND_VALENCE, which uses values of ION (or
default values if ION parameters are zero), and BOND to establish the cuttoff for bonded atoms. Note that ION, which is index by
atom-type also effects phaseshift calculations, and should be reset after use for bond valence calculations. The examples below are essentially the same as the examples used to demonstrate the program. Comment lines start with a ! as in EXCURVE command files Program output other than prompts is either indented by two spaces or in fixed-space font (tables) Prompts issued by the program are not indented User responses are indented by eight spaces and in bold. [return] signifies a carriage return. Some of the output below is, by default, written only to the log file exout.lis. Options are available to send more output !to he
terminal if required. C OUTPUT 1, 2 ,3 etc. gives more terminal output throughout The CALC command prompts for the amount of output produced during potential and phaseshift calculations Example 1. Refinement of Copper Foil !Fitting Cu foil (100 K) - typical run time ~ 6 hours !Read in the data ENTER COMMAND: r ex Filename for Experiment 1 ? culn.exn Point frequency [1] ? [return] Column combination [12] ? [return] Edge ? [CU K] [return] Sequence-number in polarisation set [0] [return] Number of clusters for this experiment [1] [return] !The next lines are the headers on the data file EXAFS : CULN.EX3 EV EXAFS Number of points read: 290 !Define the atom-types ENTER COMMAND: c atom[1-2] cu !Calculate the potentials using all default options ENTER COMMAND: ca pot;;;;;;;;;;; Z= 29 (CU) 1s1/2 1 2s1/2 2 2p1/2 2 2p3/2 4 3s1/2 2 3p1/2 2 3p3/2 4 3d3/2 4 3d5/2 6 4s1/2 2 - total 29 Z= 29 (CU) 1s1/2 2 2s1/2 2 2p1/2 2 2p3/2 4 3s1/2 2 3p1/2 2 3p3/2 4 3d3/2 4 3d5/2 6 4s1/2 1 - total 29 Excited atom code : 1 Log grid increment : 0.02784 Lattice constant : 2.56000 Muffin-tin radius : 1.28000 349 Atomic volume : 11.86329 Wigner-seitz radius: 1.41483 352 Exchange constant : 0.66670 Madelung constant : 0.00000 I R RHO(AT) RHO(SUP) RV(AT) RV(SUP) RV(MT) V(MT)) 50 0.0006 0.0478 0.0478 -57.8672 -57.8672 -57.8932 -0.987E+05 100 0.0024 0.6551 0.6551 -57.4673 -57.4673 -57.5661 -0.244E+05 150 0.0095 6.5809 6.5809 -55.8846 -55.8856 -56.2250 -0.592E+04 200 0.0382 20.6102 20.6103 -50.2474 -50.2514 -51.0456 -0.134E+04 250 0.1537 41.2852 41.2869 -34.3744 -34.3911 -36.0022 -0.234E+03 300 0.6183 24.1379 24.1672 -10.3973 -10.4697 -12.6942 -0.205E+02 348 2.3524 1.2220 2.2412 -0.7806 -1.4629 -3.2688 -0.139E+01 349 2.4188 1.1539 2.2887 -0.7235 -1.4591 -3.2973 -0.136E+01 350 2.4871 1.0892 2.3569 -0.6691 -1.4640 -3.3396 -0.134E+01 CU (CU) Rho0: 0.3775 Efermi: -5.566 V0: -18.149 Electrons: 28.903 (Z= 29) Z= 29 (CU) 1s1/2 2 2s1/2 2 2p1/2 2 2p3/2 4 3s1/2 2 3p1/2 2 3p3/2 4 3d3/2 4 3d5/2 6 4s1/2 1 - total 29 Z= 29 (CU) 1s1/2 2 2s1/2 2 2p1/2 2 2p3/2 4 3s1/2 2 3p1/2 2 3p3/2 4 3d3/2 4 3d5/2 6 4s1/2 1 - total 29 Log grid increment : 0.02784 Lattice constant : 2.56000 Muffin-tin radius : 1.28000 349 Atomic volume : 11.86329 Wigner-seitz radius: 1.41483 352 Exchange constant : 0.66670 Madelung constant : 0.00000 I R RHO(AT) RHO(SUP) RV(AT) RV(SUP) RV(MT) V(MT)) 50 0.0006 0.0845 0.0845 -57.8385 -57.8385 -57.8698 -0.986E+05 100 0.0024 1.1583 1.1583 -57.3521 -57.3520 -57.4714 -0.243E+05 150 0.0095 11.6431 11.6431 -55.4412 -55.4422 -55.8521 -0.588E+04 200 0.0382 35.3221 35.3222 -49.0428 -49.0468 -49.9952 -0.131E+04 250 0.1537 39.7602 39.7619 -33.3862 -33.4030 -34.9944 -0.228E+03 300 0.6183 22.4335 22.4629 -10.3392 -10.4121 -12.5856 -0.204E+02 348 2.3524 1.1058 2.1223 -0.6822 -1.3663 -3.1437 -0.134E+01 349 2.4188 1.0323 2.1638 -0.6330 -1.3705 -3.1788 -0.131E+01 350 2.4871 0.9641 2.2280 -0.5863 -1.3831 -3.2281 -0.130E+01 CU (CU) Rho0: 0.3574 Efermi: -5.456 V0: -17.590 Electrons: 29.055 (Z= 29) !Calculate the phaseshifts - increment the core width to allow for experimental resolution ENTER COMMAND: ca ph;;2.761 !Define the structure ENTER COMMAND: c acell 2.5*1.414213562 ENTER COMMAND: c r1 2.5 ENTER COMMAND: c t[1-20] 2 !Use a 7-shell cluster initially ENTER COMMAND: c ns 7;str fcc !Set the maximum length for MS paths ENTER COMMAND: c plmax 14 !Guess some initial values for the Debye-Waller factors ENTER COMMAND: c a1 .01;c a[2-20] .02 !Initial value of EF (the refinement will not work well if you start from 0) ENTER COMMAND: c ef 1 !Inspect the parameters ENTER COMMAND: l LMAX = 25.000 DLMAX= 25.000 TLMAX= 9.000 WP = 0.100 NS = 7.000 EF = 1.000 VPI = 0.000 AFAC = 1.000 EMIN = 3.000 EMAX =3000.000 EF1 = 0.000 VPI1 = 0.000 AFAC1= 1.000 EMIN1=-311.500 EMAX1=1525.540 Shell 0 N0 = 1.000 T0 = 1(CU) R0 = 0.000 A0 = 0.010 B0 = 0.000-1 0 Shell 1 N1 = 12.000 T1 = 2(CU) R1 = 2.500 A1 = 0.010 B1 = 0.000 1 0 Shell 2 N2 = 6.000 T2 = 2(CU) R2 = 3.536 A2 = 0.020 B2 = 0.000 1 0 Shell 3 N3 = 24.000 T3 = 2(CU) R3 = 4.330 A3 = 0.020 B3 = 0.000 1 0 Shell 4 N4 = 12.000 T4 = 2(CU) R4 = 5.000 A4 = 0.020 B4 = 0.000 1 0 Shell 5 N5 = 24.000 T5 = 2(CU) R5 = 5.590 A5 = 0.020 B5 = 0.000 1 0 Shell 6 N6 = 8.000 T6 = 2(CU) R6 = 6.124 A6 = 0.020 B6 = 0.000 1 0 Shell 7 N7 = 48.000 T7 = 2(CU) R7 = 6.614 A7 = 0.020 B7 = 0.000 1 0 !Switch on MS - only third order, two different scatterers so far ENTER COMMAND: s ms al !Refine EF, ACELL and A1 ENTER COMMAND: ref ENTER COMMAND: Parameter refinement with constraints off: ef;acell;a1;= Least squares refinement using k**2 weighting Initial parameters 1 EF 1.000 0.27212 0.00000 27.21164 0.03675 2 ACELL 3.536 0.00212 0.00000 0.21167 16.70298 3 A1 0.010 0.00028 0.00000 0.02800 0.35711 Enter: Write to save the list of variables Continue to refine interactively a number to edit, add to, or delete from the list c EF = 1.00000 ACELL = 3.53553 A1 = 0.01000 Call 1 F 66.0630 R 76.554 F- 66.063 Rex 76.554 RD 0.00 Rx 0.000 192 7 Select Next,Exit,Complete,Stats or Plot !Plot toggles plotting during refinement p EF = 1.27212 ACELL = 3.53553 A1 = 0.01000 Call 2 F 54.5289 R 68.497 F- 52.937 Rex 68.497 RD 0.00 Rx 0.000 192 7 Select Next,Exit,Complete,Stats or Plot !Next n (or just n) disables prompts for the next n cycles 333 EF = 1.00000 ACELL = 3.53765 A1 = 0.01000 Call 3 F 53.6014 R 67.680 F- 52.937 Rex 67.680 RD 0.00 Rx 0.000 192 7 EF = 1.00000 ACELL = 3.53553 A1 = 0.01028 Call 4 F 52.6384 R 67.541 F- 52.638 Rex 67.541 RD 0.00 Rx 0.000 192 7 EF = -0.76881 ACELL = 3.53020 A1 = 0.01060 Call 5 F 43.8959 R 64.220 F- 43.896 Rex 64.220 RD 0.00 Rx 0.000 192 7 EF = -2.21235 ACELL = 3.53958 A1 = 0.01100 Call 6 F 36.2281 R 58.990 F- 36.228 Rex 58.990 RD 0.00 Rx 0.000 192 7 EF = -3.76135 ACELL = 3.55790 A1 = 0.01134 Call 7 F 27.6812 R 51.503 F- 27.681 Rex 51.503 RD 0.00 Rx 0.000 192 7 EF = -6.25711 ACELL = 3.56873 A1 = 0.01136 Call 8 F 18.7336 R 43.870 F- 18.734 Rex 43.870 RD 0.00 Rx 0.000 192 7 EF = -7.88148 ACELL = 3.58281 A1 = 0.01070 Call 9 F 13.7414 R 37.167 F- 13.741 Rex 37.167 RD 0.00 Rx 0.000 192 7 EF = -7.92222 ACELL = 3.58326 A1 = 0.01097 Call 10 F 13.7654 R 37.431 F- 13.741 Rex 37.431 RD 0.00 Rx 0.000 192 7 EF = -8.08531 ACELL = 3.58141 A1 = 0.01070 Call 11 F 13.8338 R 36.869 F- 13.741 Rex 36.869 RD 0.00 Rx 0.000 192 7 EF = -9.58525 ACELL = 3.59704 A1 = 0.01040 Call 12 F 11.4548 R 33.323 F- 11.455 Rex 33.323 RD 0.00 Rx 0.000 192 7 EF = -9.92608 ACELL = 3.60063 A1 = 0.01028 Call 13 F 11.3707 R 33.062 F- 11.371 Rex 33.062 RD 0.00 Rx 0.000 192 7 Minimum Predicted, Accuracy = 0.3743E-01 Cond = 2 At Call 13 Filename for statistics ? Filename: [excora1.dat] ? <return> Call 13, R(min)= 11.3707 Best Values Are : EF = -9.92608 ACELL = 3.60063 A1 = 0.01028 !Now include all third order paths. Refine the DW factors for the !first 6 shells (4th order terms are required at the distance !of the 7th shell) ENTER COMMAND: c atmax 3 ENTER COMMAND: ref Enter parameter name or "=" to skip ef;acell;a1;a2;a3;a4;a5;a6;= Least squares refinement using k**2 weighting Initial parameters 1 EF -9.926 0.27212 0.00000 27.21164 -0.36477 2 ACELL 3.601 0.00212 0.00000 0.21167 17.01051 3 A1 0.010 0.00029 0.00000 0.02878 0.35711 4 A2 0.020 0.00056 0.00000 0.05601 0.35711 5 A3 0.020 0.00056 0.00000 0.05601 0.35711 6 A4 0.020 0.00056 0.00000 0.05601 0.35711 7 A5 0.020 0.00056 0.00000 0.05601 0.35711 8 A6 0.020 0.00056 0.00000 0.05601 0.35711 Enter: Write to save the list of variables Continue to refine interactively a number to edit, add to, or delete from the list c;p;c;; EF = -9.92608 ACELL = 3.60063 A1 = 0.01028 A2 = 0.02000 A3 = 0.02000 A4 = 0.02000 A5 = 0.02000 A6 = 0.02000 Call 1 F 9.0099 R 29.860 F- 9.010 Rex 29.860 RD 0.00 Rx 0.000 192 7 EF = -9.65396 ACELL = 3.60063 A1 = 0.01028 A2 = 0.02000 A3 = 0.02000 A4 = 0.02000 A5 = 0.02000 A6 = 0.02000 Call 2 F 9.6026 R 30.736 F- 9.010 Rex 30.736 RD 0.00 Rx 0.000 192 7 etc. EF = -10.37898 ACELL = 3.59946 A1 = 0.01022 A2 = 0.01639 A3 = 0.01365 A4 = 0.01434 A5 = 0.01799 A6 = 0.01818 Call 22 F 6.1436 R 24.564 F- 6.144 Rex 24.564 RD 0.00 Rx 0.000 192 7 Minimum Predicted, Accuracy = 0.6371E-02 Cond = 2 At Call 22 EF = -10.37898 ACELL = 3.59946 A1 = 0.01022 A2 = 0.01639 A3 = 0.01365 A4 = 0.01434 A5 = 0.01799 A6 = 0.01818 Call 23 F 6.1436 R 24.564 F- 6.144 Rex 24.564 RD 0.00 Rx 0.000 192 7 etc. EF = -10.37898 ACELL = 3.59946 A1 = 0.01022 A2 = 0.01639 A3 = 0.01365 A4 = 0.01434 A5 = 0.01799 A6 = 0.01874 Call 31 F 6.1454 R 24.571 F- 6.144 Rex 24.571 RD 0.00 Rx 0.000 192 7 Fit index * k**2 : 0.6145E-03 R-factor : 24.5712 Chisqu : 0.3329E-05 Rexafs 24.5712 Rdist 0.0000 !Now extend the model to 10 shells, with all 5th order paths !This will take ages ENTER COMMAND c ns 10 ENTER COMMAND: str fcc q c plmax 16;c omax 5;c atmax 5 !Refine the DW factors only ENTER COMMAND ref;a1;a2;a3;a4;a5;a6;a7;a8;a[9-10];= Least squares refinement using k**2 weighting Initial parameters 1 A1 0.010 0.00029 0.46548 0.02861 0.35711 2 A2 0.016 0.00046 0.00391 0.04589 0.35711 3 A3 0.014 0.00038 0.00052 0.03824 0.35711 4 A4 0.014 0.00040 0.00426 0.04017 0.35711 5 A5 0.018 0.00050 0.00154 0.05038 0.35711 6 A6 0.018 0.00051 0.00186 0.05092 0.35711 7 A7 0.020 0.00056 0.00602 0.05601 0.35711 8 A8 0.020 0.00056 0.01806 0.05601 0.35711 9 A[9-10] 0.020 0.00056 0.00000 0.05601 0.35711 Enter: Write to save the list of variables Continue to refine interactively a number to edit, add to, or delete from the list c;p;c; A1 = 0.01022 A2 = 0.01639 A3 = 0.01365 A4 = 0.01434 A5 = 0.01799 A6 = 0.01818 A7 = 0.02000 A8 = 0.02000 A[9-10] = 0.02000 Call 1 F 4.5154 R 20.310 F- 4.515 Rex 20.310 RD 0.00 Rx 0.000 192 10 A1 = 0.01050 A2 = 0.01639 A3 = 0.01365 A4 = 0.01434 A5 = 0.01799 A6 = 0.01818 A7 = 0.02000 A8 = 0.02000 A[9-10] = 0.02000 Call 2 F 4.5573 R 20.519 F- 4.515 Rex 20.519 RD 0.00 Rx 0.000 192 10 etc. A1 = 0.01020 A2 = 0.01596 A3 = 0.01355 A4 = 0.01395 A5 = 0.01708 A6 = 0.01176 A7 = 0.01450 A8 = 0.01388 A[9-10] = 0.02227 Call 15 F 4.3090 R 19.786 F- 4.309 Rex 19.786 RD 0.00 Rx 0.000 192 10 Minimum Predicted, Accuracy = 0.3193E-02 Cond = 2 At Call 15 Call 15, R(min)= 4.3090 Best Values Are : A1 = 0.01020 A2 = 0.01596 A3 = 0.01355 A4 = 0.01395 A5 = 0.01708 A6 = 0.01176 A7 = 0.01450 A8 = 0.01388 A[9-10] = 0.02227 !Save the parameters, and create a table of MS paths (using the default filename in each case ENTER COMMAND: pr par; Fit Index: with k^2 Weighting 0.4309 R-Factor : with k^2 Weighting 19.7860 Amplitude of Experiment 53.8497 R(exafs) = 19.7860 Weight: 1.00 R(distances)= 0.0000 Weight: 0.00 R(angles) = 0.0000 Weight: 0.00 N(ind) = 31.5 Np = 1. Chi^2 = 2.3179E-6 Parameters printed in expara1.dat ENTER COMMAND extr q; Paths printed in extaba1.dat !Plot the final result ENTER COMMAND cp !Obtain a hard copy ENTER COMMAND: gs dev p;cp !Finish ENTER COMMAND: END A much better fit can be obtained using 20 shells at considerable extra computational expense. Figure 2 - 10 shell fit to Cu foil Example 1a. Whole Spectrum Refinement of Whole Sectrum Refinement ofCopper Foil !Select the plotter amd set up some aliases gs plot ps_file alias pcp;gs dev p;cp;gs dev t alias pp;gs dev p;p;gs dev t !Set the e0 value in order to read the experiment c e0 8979.66 r ex;culn.bsa;;;;;;;;;;;;;;;;;; !Read the paramaters - from a seven shell EXAFS refinement r par;cu.par !!Define the symmetry sym oh !Select the whole-spectrum option s atom on !Allow calculation below the edge c emin -3 !Select the core wdith (1 ev for experimental resolution) c cw 2.761 !Include multiple scattering s ms al !Calculate potentials and phaseshifts !!These are defaults - using different options can give better results ca pot;;;;;;;;;;;;;;; ca ph;;;;;;;;;;;;;;;; !Select 5th order correction c npost 5 !Refine the post-edge polynomial correction parameters, ! plus effective temperature and the E0 correction ! First step will be slow ref post 6;offset;7;temp;8;de0 c;p;c;= cp !Turn off the k^2 weighting to see the edge s we n p !Display the atomic-background parameters l back Figure 3 - Unweighted whole spectrum fit to Cu
foil Figure 4 - Whole spectrum fit to k^2 weighted Cu
foil Find the cell parameter and Debye-Waller factors of GaSb by simultaneously fitting the Ga K-edge and Sb K-edge The compound has the cubic sphalerite structure, with Sb and Ga each in 4-fold coordination. The author wishes to thank
Dr.A.Sapelkin for permission to use this data. !Read the data for the gallium edge ENTER COMMAND: r ex 1 Filename for Experiment 1 ? cgasb.exb Point frequency [1] ? 1 Column combination [12] ? 32 Edge ? [CU K] ga k Sequence-number in polarisation set [0] ? [return] Number of clusters for this experiment [1] ? [return] HARTREES ABSORPTION EV Number of points read: 359 SPECTRUM : 1 359 0.670 1511.020 !Read the data for the antimony edge ENTER COMMAND: r ex 2;csbga.exb;;32;sb k;;;; HARTREES ABSORPTION EV Number of points read: 437 SPECTRUM : 2 437 0.270 0.081 1493.870 0.000 !Read some pre-existing phaseshift files ENTER COMMAND: r ph Enter number of phaseshift files 6 Filename for central atom ? exphsa1.gac EXCURVE >>> 23/ 3/98 20:52:05 GA phaseshifts (Z= 31). mtr 1.4100 alpha 0.667 madelung const. 0.00 hole-code 1 V0 -17.3443 Ef -5.6451 RHO0 0.3384 Rlife 7.0000 Number Of Energy Points: 90 Filename for second atom ? .sb;.ga;.sbc;.ga2;.sb2 SB phaseshifts (Z= 51). mtr 1.5900 alpha 0.667 madelung const. 0.00 hole-code 0 V0 -16.6932 Ef -6.2505 RHO0 0.2854 Rlife 7.0000 Number Of Energy Points: 90 GA phaseshifts (Z= 31). mtr 1.4100 alpha 0.667 madelung const. 0.00 hole-code 0 V0 -17.0346 Ef -5.9492 RHO0 0.3121 Rlife 7.0000 Number Of Energy Points: 90 SB phaseshifts (Z= 51). mtr 1.5900 alpha 0.667 madelung const. 0.00 hole-code 1 V0 -16.3918 Ef -5.7126 RHO0 0.2951 Rlife 7.0000 Number Of Energy Points: 90 GA phaseshifts (Z= 31). mtr 1.4100 alpha 0.667 madelung const. 0.00 hole-code 0 V0 -17.0346 Ef -5.9492 RHO0 0.3121 Rlife 7.0000 Number Of Energy Points: 90 SB phaseshifts (Z= 51). mtr 1.5900 alpha 0.667 madelung const. 0.00 hole-code 0 V0 -16.6932 Ef -6.2505 RHO0 0.2854 Rlife 7.0000 Number Of Energy Points: 90 !Generate the structure - this is made slightly more !complicated by the need to generate two clusters, one for each edge !First set the atom parameters c atom[1,3,5] ga;c atom[2,4,6] sb !Set the first shell distance to 6.095/sqrt(16/3) - the approximate !crystallographic value at RT c r1 2.63921 !Set the atom types for the first two shells and the cluster size c t1 sb;c t2 ga;c ns 13 !define the structure ENTER COMMAND: str sph q ENTER COMMAND: l LMAX = 25.000 DLMAX= 25.000 TLMAX= 9.000 WP = 0.100 NS = 13.000 EF = 0.000 VPI = 0.000 AFAC = 1.000 EMIN = 3.000 EMAX =3000.000 EF1 = 0.000 VPI1 = 0.000 AFAC1= 1.000 EMIN1= 0.670 EMAX1=1511.020 EF2 = 0.000 VPI2 = 0.000 AFAC2= 1.000 EMIN2= 0.670 EMAX2=1511.020 Shell 0 N0 = 1.000 T0 = 1(GA) R0 = 0.000 A0 = 0.010 B0 = 0.000-1 0 Shell 1 N1 = 4.000 T1 = 6(SB) R1 = 2.639 A1 = 0.010 B1 = 0.000 1 0 Shell 2 N2 = 12.000 T2 = 5(GA) R2 = 4.310 A2 = 0.010 B2 = 0.000 1 0 Shell 3 N3 = 12.000 T3 = 6(SB) R3 = 5.054 A3 = 0.010 B3 = 0.000 1 0 Shell 4 N4 = 6.000 T4 = 5(GA) R4 = 6.095 A4 = 0.010 B4 = 0.000 1 0 Shell 5 N5 = 12.000 T5 = 6(SB) R5 = 6.642 A5 = 0.010 B5 = 0.000 1 0 Shell 6 N6 = 12.000 T6 = 5(GA) R6 = 7.465 A6 = 0.010 B6 = 0.000 1 0 Shell 7 N7 = 12.000 T7 = 5(GA) R7 = 7.465 A7 = 0.010 B7 = 0.000 1 0 Shell 8 N8 = 4.000 T8 = 6(SB) R8 = 7.918 A8 = 0.010 B8 = 0.000 1 0 Shell 9 N9 = 12.000 T9 = 6(SB) R9 = 7.918 A9 = 0.010 B9 = 0.000 1 0 Shell 10 N10= 12.000 T10= 5(GA) R10 = 8.620 A10 = 0.010 B10 = 0.000 1 0 Shell 11 N11= 24.000 T11= 6(SB) R11 = 9.015 A11 = 0.010 B11 = 0.000 1 0 Shell 12 N12= 24.000 T12= 5(GA) R12 = 9.637 A12 = 0.010 B12 = 0.000 1 0 Shell 13 N13= 12.000 T13= 6(SB) R13 = 9.992 A13 = 0.010 B13 = 0.000 1 0 !Remove any existing file PAR1 (O/S dependent) ENTER COMMAND: ^del par1 !store the result for the Ga edge ENTER COMMAND: pr par Fit Index: with k^2 Weighting 3.4382 R-Factor : with k^2 Weighting 116.5080 Amplitude of Experiment 101.5911 R(exafs) = 116.5080 Weight: 1.00 R(distances)= 0.0000 Weight: 0.00 R(angles) = 0.0000 Weight: 0.00 N(ind) = 88.4 Np = 1. Chi^2 = 4.8704E-6 Filename: [expara4.dat ] ? par1 Parameters printed in par1 !Repeat for the Sb edge - remember to redefine the central atom type ENTER COMMAND: c t0 sb;c t1 ga;c t2 sb;str sph q ENTER COMMAND: l LMAX = 25.000 DLMAX= 25.000 TLMAX= 9.000 WP = 0.100 NS = 13.000 EF = 0.000 VPI = 0.000 AFAC = 1.000 EMIN = 3.000 EMAX =3000.000 EF1 = 0.000 VPI1 = 0.000 AFAC1= 1.000 EMIN1= 0.670 EMAX1=1511.020 EF2 = 0.000 VPI2 = 0.000 AFAC2= 1.000 EMIN2= 0.670 EMAX2=1511.020 Shell 0 N0 = 1.000 T0 = 6(SB) R0 = 0.000 A0 = 0.010 B0 = 0.000-1 0 Shell 1 N1 = 4.000 T1 = 5(GA) R1 = 2.639 A1 = 0.010 B1 = 0.000 1 0 Shell 2 N2 = 12.000 T2 = 6(SB) R2 = 4.310 A2 = 0.010 B2 = 0.000 1 0 Shell 3 N3 = 12.000 T3 = 5(GA) R3 = 5.054 A3 = 0.010 B3 = 0.000 1 0 Shell 4 N4 = 6.000 T4 = 6(SB) R4 = 6.095 A4 = 0.010 B4 = 0.000 1 0 Shell 5 N5 = 12.000 T5 = 5(GA) R5 = 6.642 A5 = 0.010 B5 = 0.000 1 0 Shell 6 N6 = 12.000 T6 = 6(SB) R6 = 7.465 A6 = 0.010 B6 = 0.000 1 0 Shell 7 N7 = 12.000 T7 = 6(SB) R7 = 7.465 A7 = 0.010 B7 = 0.000 1 0 Shell 8 N8 = 4.000 T8 = 5(GA) R8 = 7.918 A8 = 0.010 B8 = 0.000 1 0 Shell 9 N9 = 12.000 T9 = 5(GA) R9 = 7.918 A9 = 0.010 B9 = 0.000 1 0 Shell 10 N10= 12.000 T10= 6(SB) R10 = 8.620 A10 = 0.010 B10 = 0.000 1 0 Shell 11 N11= 24.000 T11= 5(GA) R11 = 9.015 A11 = 0.010 B11 = 0.000 1 0 Shell 12 N12= 24.000 T12= 6(SB) R12 = 9.637 A12 = 0.010 B12 = 0.000 1 0 Shell 13 N13= 12.000 T13= 5(GA) R13 = 9.992 A13 = 0.010 B13 = 0.000 1 0 ENTER COMMAND: ^del par2 pr par;par2 Fit Index: with k^2 Weighting 2.0194 R-Factor : with k^2 Weighting 94.2218 Amplitude of Experiment 101.4107 R(exafs) = 94.2218 Weight: 1.00 R(distances)= 0.0000 Weight: 0.00 R(angles) = 0.0000 Weight: 0.00 N(ind) = 88.5 Np = 1. Chi^2 = 2.8606E-6 Parameters printed in par2 !Now re-read the parameters for each cluster ENTER COMMAND: r par;par1 Experiment in : cgasb.exb Fit index with K**2 weight: 0.3438E-02 R-factor :116.5080 Chisqu : 4.87043 ENTER COMMAND: r par add;par2 Experiment in : cgasb.exb Fit index with K**2 weight: 0.2019E-02 R-factor : 94.2218 Chisqu : 2.86061 !Fix the cluster parameters for the second cluster ENTER COMMAND: c clus14 -2;c clus[15-ns] 2 !Select the useful energy range ENTER COMMAND: c emin 5;c emax 1250 !Make sure the point group (Td) is defined for both clusters ENTER COMMAND: sym td;td !Select the MS parameters - 5th order but paths !involve only 3 DIFFERENT scatterers - initially only include paths to 12 angstroms ENTER COMMAND: c atmax 3;c omax 5;c plmax 12 !Turn the MS on (the ALL option selects all paths in the cluster within filter limits) ENTER COMMAND: s ms al !Select some initial DW factors ENTER COMMAND: c a[1-ns] .02 ENTER COMMAND: c a[1,15] .005;c a[2-4,16-18] .015 !Two energy zero terms are also needed - set these to some value other than 0 at the start ENTER COMMAND: c ef1 20;c ef2 1 ENTER COMMAND: l LMAX = 25.000 DLMAX= 25.000 TLMAX= 9.000 WP = 0.100 NS = 27.000 EF = 0.000 VPI = 0.000 AFAC = 1.000 EMIN = 5.000 EMAX =1250.000 EF1 = 5.000 VPI1 = 0.000 AFAC1= 1.000 EMIN1= 0.670 EMAX1=1511.020 EF2 = 5.000 VPI2 = 0.000 AFAC2= 1.000 EMIN2= 0.670 EMAX2=1511.020 Shell 0 N0 = 1.000 T0 = 1(GA) R0 = 0.000 A0 = 0.010 B0 = 0.000-1 0 Shell 1 N1 = 4.000 T1 = 6(SB) R1 = 2.639 A1 = 0.005 B1 = 0.000 1 0 Shell 2 N2 = 12.000 T2 = 5(GA) R2 = 4.310 A2 = 0.015 B2 = 0.000 1 0 Shell 3 N3 = 12.000 T3 = 6(SB) R3 = 5.054 A3 = 0.015 B3 = 0.000 1 0 Shell 4 N4 = 6.000 T4 = 5(GA) R4 = 6.095 A4 = 0.015 B4 = 0.000 1 0 Shell 5 N5 = 12.000 T5 = 6(SB) R5 = 6.642 A5 = 0.020 B5 = 0.000 1 0 Shell 6 N6 = 12.000 T6 = 5(GA) R6 = 7.465 A6 = 0.020 B6 = 0.000 1 0 Shell 7 N7 = 12.000 T7 = 5(GA) R7 = 7.465 A7 = 0.020 B7 = 0.000 1 0 Shell 8 N8 = 4.000 T8 = 6(SB) R8 = 7.918 A8 = 0.020 B8 = 0.000 1 0 Shell 9 N9 = 12.000 T9 = 6(SB) R9 = 7.918 A9 = 0.020 B9 = 0.000 1 0 Shell 10 N10= 12.000 T10= 5(GA) R10 = 8.620 A10 = 0.020 B10 = 0.000 1 0 Shell 11 N11= 24.000 T11= 6(SB) R11 = 9.015 A11 = 0.020 B11 = 0.000 1 0 Shell 12 N12= 24.000 T12= 5(GA) R12 = 9.637 A12 = 0.020 B12 = 0.000 1 0 Shell 13 N13= 12.000 T13= 6(SB) R13 = 9.992 A13 = 0.020 B13 = 0.000 1 0 Shell 14 N14= 1.000 T14= 6(SB) R14 = 0.000 A14 = 0.020 B14 = 0.000-2 0 Shell 15 N15= 4.000 T15= 5(GA) R15 = 2.639 A15 = 0.005 B15 = 0.000 2 0 Shell 16 N16= 12.000 T16= 6(SB) R16 = 4.310 A16 = 0.015 B16 = 0.000 2 0 Shell 17 N17= 12.000 T17= 5(GA) R17 = 5.054 A17 = 0.015 B17 = 0.000 2 0 Shell 18 N18= 6.000 T18= 6(SB) R18 = 6.095 A18 = 0.015 B18 = 0.000 2 0 Shell 19 N19= 12.000 T19= 5(GA) R19 = 6.642 A19 = 0.020 B19 = 0.000 2 0 Shell 20 N20= 12.000 T20= 6(SB) R20 = 7.465 A20 = 0.020 B20 = 0.000 2 0 Shell 21 N21= 12.000 T21= 6(SB) R21 = 7.465 A21 = 0.020 B21 = 0.000 2 0 Shell 22 N22= 4.000 T22= 5(GA) R22 = 7.918 A22 = 0.020 B22 = 0.000 2 0 Shell 23 N23= 12.000 T23= 5(GA) R23 = 7.918 A23 = 0.020 B23 = 0.000 2 0 Shell 24 N24= 12.000 T24= 6(SB) R24 = 8.620 A24 = 0.020 B24 = 0.000 2 0 Shell 25 N25= 24.000 T25= 5(GA) R25 = 9.015 A25 = 0.020 B25 = 0.000 2 0 Shell 26 N26= 24.000 T26= 6(SB) R26 = 9.637 A26 = 0.020 B26 = 0.000 2 0 Shell 27 N27= 12.000 T27= 5(GA) R27 = 9.992 A27 = 0.020 B27 = 0.000 2 0 !Plot the result (this will be slow) ENTER COMMAND: cp !Refine the structure - as it is cubic, only ACELL can sensibly be refined - it will just scale all the distances ENTER COMMAND: ref 80 Parameter refinement with constraints off: Enter parameter name or = : acell ef1 ef2 = Least squares refinement using k**2 weighting Initial parameters 1 ACELL 1.000 0.00532 0.00000 0.21167 4.72432 2 EF1 20.000 0.68353 0.00000 27.21164 0.73498 3 EF2 1.000 0.68353 0.00000 27.21164 0.03675 Enter : Write to save the list of variables Continue to refine interactively a number to edit, add to, or delete from the list !This is correct so no editing needed c ACELL = 1.00000 EF1 = 20.00000 EF2 = 1.00000 Call 1 F 2.7502 R 32.476 F- 2.750 Rex 32.48 RD 0.00 Rx 0.00 636 27 !The continue option will complete the refinement without further prompts Select Next,Exit,Complete,Stats or Plot p;c ACELL = 1.00532 EF1 = 20.00000 EF2 = 1.00000 Call 2 F 5.3635 R 46.184 F- 2.750 Rex 46.18 RD 0.00 Rx 0.00 636 27 ACELL = 1.00000 EF1 = 20.68353 EF2 = 1.00000 Call 3 F 2.6433 R 32.044 F- 2.643 Rex 32.04 RD 0.00 Rx 0.00 636 27 ACELL = 1.00000 EF1 = 20.68353 EF2 = 1.68353 Call 4 F 2.8511 R 33.532 F- 2.643 Rex 33.53 RD 0.00 Rx 0.00 636 27 ACELL = 0.99875 EF1 = 21.21262 EF2 = 0.59809 Call 5 F 2.3736 R 30.243 F- 2.374 Rex 30.24 RD 0.00 Rx 0.00 636 27 ACELL = 0.99843 EF1 = 21.84350 EF2 = 0.42212 Call 6 F 2.3325 R 29.980 F- 2.333 Rex 29.98 RD 0.00 Rx 0.00 636 27 ACELL = 1.00352 EF1 = 22.04184 EF2 = 0.42212 Call 7 F 4.5556 R 42.192 F- 2.333 Rex 42.19 RD 0.00 Rx 0.00 636 27 ACELL = 0.99841 EF1 = 21.83211 EF2 = 0.20828 Call 8 F 2.3242 R 29.906 F- 2.324 Rex 29.91 RD 0.00 Rx 0.00 636 27 ACELL = 0.99750 EF1 = 22.21673 EF2 = 0.76116 Call 9 F 2.4023 R 30.615 F- 2.324 Rex 30.62 RD 0.00 Rx 0.00 636 27 ACELL = 0.99891 EF1 = 21.42129 EF2 = -0.33415 Call 10 F 2.3064 R 29.660 F- 2.306 Rex 29.66 RD 0.00 Rx 0.00 636 27 Minimum Predicted, Accuracy = 0.1541E-02 Cond = 2 At Call 10 Fit index * k^2: 0.2306 Rdistances: 0.0000 Chisqu: 0.3662E-06 Rexafs: 29.6604 Rxray: 0.0000 Rd: 0.0000 Call 10, R(min)= 2.3064 Best Values Are : 1 ACELL = 0.99891 +/- 0.00067 (2 sigma) 2 EF1 = 21.42129 +/- 0.48710 (2 sigma) 3 EF2 = -0.33415 +/- 0.51268 (2 sigma) !Replot ENTER COMMAND: cp ENTER COMMAND: !Add some more MS paths ENTER COMMAND: c plmax 16 !The DW factors for Ga-Sb and Sb-Ga at the same distance must be the same !Here it is also assumed that Ga-Ga and Sb-Sb DW factors are the same !(this turns out to be a reasonable approximation) ENTER COMMAND: ref;a[1,15];a[2,16];a[3,17];a[4,18];a[5,19];a[6,7,20,21];a[8,9,22,23];= Parameter refinement with constraints off: Least squares refinement using k**2 weighting Initial parameters 1 A[1,15] 0.005 0.00014 0.00000 0.01400 0.35711 2 A[2,16] 0.015 0.00042 0.00000 0.04200 0.35711 3 A[3,17] 0.015 0.00042 0.00000 0.04200 0.35711 4 A[4,18] 0.015 0.00042 0.00000 0.04200 0.35711 5 A[5,19] 0.020 0.00056 0.00000 0.05601 0.35711 6 A[6,7,20,2 0.020 0.00056 0.00000 0.05601 0.35711 7 A[8,9,22,2 0.020 0.00056 0.00000 0.05601 0.35711 Continue to refine interactively a number to edit, add to, or delete from the list c;p !First cycle will be slow Call 1 F 2.1960 R 28.800 F- 2.196 Rex 28.80 RD 0.00 Rx 0.00 636 27 A[1,15] = 0.00514 A[2,16] = 0.01500 A[3,17] = 0.01500 A[4,18] = 0.01500 A[5,19] = 0.02000 A[6,7,20= 0.02000 A[8,9,22= 0.02000 Call 2 F 2.1843 R 28.745 F- 2.184 Rex 28.74 RD 0.00 Rx 0.00 636 27 A[1,15] = 0.00514 A[2,16] = 0.01542 A[3,17] = 0.01500 A[4,18] = 0.01500 A[5,19] = 0.02000 A[6,7,20= 0.02000 A[8,9,22= 0.02000 !Selecting complete avoids all further prompts but the last - which is for the filename for the statistics Select Next,Exit,Complete,Stats or Plot c A[1,15] = 0.00514 A[2,16] = 0.01542 A[3,17] = 0.01500 A[4,18] = 0.01500 A[5,19] = 0.02000 A[6,7,20= 0.02000 A[8,9,22= 0.02000 Call 3 F 2.2438 R 29.107 F- 2.184 Rex 29.11 RD 0.00 Rx 0.00 636 27 A[1,15] = 0.00514 A[2,16] = 0.01500 A[3,17] = 0.01542 A[4,18] = 0.01500 A[5,19] = 0.02000 A[6,7,20= 0.02000 A[8,9,22= 0.02000 Call 4 F 2.1939 R 28.784 F- 2.184 Rex 28.78 RD 0.00 Rx 0.00 636 27 A[1,15] = 0.00514 A[2,16] = 0.01500 A[3,17] = 0.01500 A[4,18] = 0.01542 A[5,19] = 0.02000 A[6,7,20= 0.02000 A[8,9,22= 0.02000 Call 5 F 2.1849 R 28.736 F- 2.184 Rex 28.74 RD 0.00 Rx 0.00 636 27 A[1,15] = 0.00514 A[2,16] = 0.01500 A[3,17] = 0.01500 A[4,18] = 0.01500 A[5,19] = 0.02056 A[6,7,20= 0.02000 A[8,9,22= 0.02000 Call 6 F 2.1818 R 28.704 F- 2.182 Rex 28.70 RD 0.00 Rx 0.00 636 27 A[1,15] = 0.00514 A[2,16] = 0.01500 A[3,17] = 0.01500 A[4,18] = 0.01500 A[5,19] = 0.02056 A[6,7,20= 0.02056 A[8,9,22= 0.02000 Call 7 F 2.1869 R 28.721 F- 2.182 Rex 28.72 RD 0.00 Rx 0.00 636 27 A[1,15] = 0.00514 A[2,16] = 0.01500 A[3,17] = 0.01500 A[4,18] = 0.01500 A[5,19] = 0.02056 A[6,7,20= 0.02000 A[8,9,22= 0.02056 Call 8 F 2.1835 R 28.717 F- 2.182 Rex 28.72 RD 0.00 Rx 0.00 636 27 A[1,15] = 0.00527 A[2,16] = 0.01088 A[3,17] = 0.01440 A[4,18] = 0.01496 A[5,19] = 0.02071 A[6,7,20= 0.01953 A[8,9,22= 0.01984 A[1,15] = 0.00514 A[2,16] = 0.01542 A[3,17] = 0.01500 A[4,18] = 0.01500 A[5,19] = 0.02000 A[6,7,20= 0.02000 A[8,9,22= 0.02000 Call 9 F 1.7333 R 25.821 F- 1.733 Rex 25.82 RD 0.00 Rx 0.00 636 27 A[1,15] = 0.00513 A[2,16] = 0.01048 A[3,17] = 0.01355 A[4,18] = 0.01438 A[5,19] = 0.02213 A[6,7,20= 0.01508 A[8,9,22= 0.01719 Call 10 F 1.6575 R 25.037 F- 1.658 Rex 25.04 RD 0.00 Rx 0.00 636 27 A[1,15] = 0.00518 A[2,16] = 0.01055 A[3,17] = 0.01344 A[4,18] = 0.01372 A[5,19] = 0.02242 A[6,7,20= 0.01335 A[8,9,22= 0.01463 Call 11 F 1.6566 R 24.974 F- 1.657 Rex 24.97 RD 0.00 Rx 0.00 636 27 A[1,15] = 0.00518 A[2,16] = 0.01050 A[3,17] = 0.01339 A[4,18] = 0.01417 A[5,19] = 0.02257 A[6,7,20= 0.01347 A[8,9,22= 0.01708 Call 12 F 1.6538 R 24.985 F- 1.654 Rex 24.98 RD 0.00 Rx 0.00 636 27 Minimum Predicted, Accuracy = 0.7764E-03 Cond = 2 At Call 12 Filename: [excora6.dat ] ? [return] Value of fit index at predicted minimum 1.6538 Fit index * k^2: 0.1654 Rdistances: 0.0000 Chisqu: 0.2626E-06 Rexafs: 24.9848 Rxray: 0.0000 Rd: 0.0000 Call 12, R(min)= 1.6538 Best Values Are : 1 A[1,15] = 0.00518 +/- 0.00018 (2 sigma) 2 A[2,16] = 0.01050 +/- 0.00071 (2 sigma) 3 A[3,17] = 0.01339 +/- 0.00139 (2 sigma) 4 A[4,18] = 0.01417 +/- 0.00488 (2 sigma) 5 A[5,19] = 0.02257 +/- 0.00195 (2 sigma) 6 A[6,7,20,21] = 0.01347 +/- 0.00356 (2 sigma) 7 A[8,9,22,23] = 0.01708 +/- 0.00519 (2 sigma) Correlation matrix printed in excora6.dat !Display the final R-factor and plot the results Figure 5 - Fit to Ga and Sb K-edges of
GaSb ENTER COMMAND: fit Fit Index: with k^2 Weighting 0.1654 R-Factor : with k^2 Weighting 24.9848 Amplitude of Experiment 102.4680 R(exafs) = 24.9848 Weight: 1.00 R(distances)= 0.0000 Weight: 0.00 R(angles) = 0.0000 Weight: 0.00 ENTER COMMAND: cp ENTER COMMAND: end Total cpu time used = 1289.76 Elapsed time = 1290.00 Example 3. Cu-Zn Superoxide dismutase Start the program - this example requires a version with large dimension limits excurve big !Start the example in file stsod %stsod !Henceforth the i/o is as for direct input from the terminal - some of this is supressed when the command file is run unless GSET
PROMPT MAX is selected !Read the Cu K-edge data ENTER COMMAND: r ex 1 Filename for Experiment 1 ? r51991a.SODCU.exback Point frequency [1] ? [return] Column combination [12] ? 32 Sequence-number in polarisation set [0] [return] Edge ? [CU K] cu k Number of clusters for this experiment [1] [return] !These are the title records for this experiment 0.00000 0.08302 25.23699 Background subtracted spectrum 304 Hartrees Absorption Ev Edgenor pre Number of points read: 304 !Read the Zn K-edge data (avoid prompts this time) ENTER COMMAND: r ex 2;r52042a.SODZN.exback;1;32;zn k;;;; 0.00000 0.08706 27.33692 Background subtracted spectrum 314 Hartrees Absorption Ev Edgenor pre Number of points read: 314 !Set up the atom types - the * indicates a central atom. Here we are using the same !scattering phaseshifts for both Cu and Zn edges (the imaginary parts should really ! be slightly different). ENTER COMMAND: c atom1 cu*;c atom2 c;c atom3 n;c atom4 o;c atom5 zn;c atom6 cu;c atom7 zn* !Set the search radius for residues and bond distance for molecular graphics etc. ENTER COMMAND: c maxrad 8.5 ENTER COMMAND: c bond 2.1 !Read the .pdb files ENTER COMMAND: r par br Reading data with format: BROOKHAVEN (.pdb) Filename for parameters ? sod Enter Cluster Number: [1] [return] Central atom number [1 ] 2168 !This is how to set up an indepenent cluster for Zn !r par br;sod.pdb;2;2169;;;; !Here the clusters are to be linked however !Calculate potentials - the order depends on how the atom variables are defined ca pot;c;n;1;c;c;n;n;n;n; Calculating potentials for all atoms: Z= 29 (CU) 1s1/2 1 2s1/2 2 2p1/2 2 2p3/2 4 3s1/2 2 3p1/2 2 3p3/2 4 3d3/2 4 3d5/2 6 4s1/2 2 - total 29 Z= 7 (N) 1s1/2 2 2s1/2 2 2p1/2 1 2p3/2 2 - total 7 CU (N ) Rho0: 0.5100 Efermi: -3.014 V0: -18.393 Electrons: 29.145 (Z= 29) ! ..... output for additional atoms follows .... Z= 16 (S) 1s1/2 2 2s1/2 2 2p1/2 2 2p3/2 4 3s1/2 2 3p1/2 2 3p3/2 2 - total 16 Z= 30 (ZN) 1s1/2 2 2s1/2 2 2p1/2 2 2p3/2 4 3s1/2 2 3p1/2 2 3p3/2 4 3d3/2 4 3d5/2 6 4s1/2 2 - total 30 S (ZN) Rho0: 0.4364 Efermi: -3.665 V0: -17.525 Electrons: 15.657 (Z= 16) !Calculate phaseshifts ca ph Calculating phaseshifs for all atoms: Enter core width (FWHM eV) [ 1.761] [return] Enter core width (FWHM eV) [ 1.936] [return] !Sort in order of radius sort Shells Have Been Put in Order of Ascending R !Generate a second cluster centred on the Zn at 6 angstroms !The cluster is to be a LINKED cluster ENTER COMMAND: gen link 2 Enter atom number for cluster 2 53 !The Zn was in shell 53 of the Cu cluster (cluster 1) !Remember to change the atom-type for the excited atoms in cluster 2 !Sort again sort Shells Have Been Put in Order of Ascending R !Remove the unwanted part of the Zn cluster ENTER COMMAND: c un[284-ns] 0 ENTER COMMAND: c ns 283 !Set some initial values of EF ENTER COMMAND: c ef[1-2] -2 !Set up DW factors based on model compounds ENTER COMMAND: c a[1-4] .005;c a[5-14] .009;c a[15-26] .016;c a[26-250] .025 !Zn cluster model compounds ENTER COMMAND: c a[252-255] .005;c a[256-265] .009;c a[266-275] .016;c a[276-ns] .025 !Define the symmetry ENTER COMMAND: sym c1;c1 !Draw the 2 clusters ENTER COMMAND: dr cl 1 dr cl 2 ENTER COMMAND: !Plot the result ENTER COMMAND: cp !Refine the EFs and central atom positions ENTER COMMAND: ref 50;ef1;ef2;x0;y0;z0;x53;y53;z53;= Parameter refinement with constraints off: Least squares refinement using k**2 weighting Initial parameters 1 EF1 -2.000 2.72116 0.00000 27.21164 -0.07350 2 EF2 -2.000 2.72116 0.00000 27.21164 -0.07350 3 X0 0.001 0.02117 0.00000 0.21167 0.00472 4 Y0 0.001 0.02117 0.00000 0.21167 0.00472 5 Z0 0.001 0.02117 0.00000 0.21167 0.00472 6 X53 -3.014 0.02117 0.00000 0.21167 -14.23909 7 Y53 3.907 0.02117 0.00000 0.21167 18.45790 8 Z53 -3.572 0.02117 0.00000 0.21167 -16.87525 Enter : Write to save the list of variables Continue to refine interactively a number to edit, add to, or delete from the list c EF1 = -2.00000 EF2 = -2.00000 X0 = 0.00100 Y0 = 0.00100 Z0 = 0.00100 X53 = -3.01400 Y53 = 3.90700 Z53 = -3.57200 Call 1 F 20.9503 R 79.159 F- 20.950 Rex 79.16 RD 0.00 Rx 0.00 594 283 EF1 = 0.72116 EF2 = -2.00000 X0 = 0.00100 Y0 = 0.00100 Z0 = 0.00100 X53 = -3.01400 Y53 = 3.90700 Z53 = -3.57200 Call 2 F 23.8008 R 83.411 F- 20.950 Rex 83.41 RD 0.00 Rx 0.00 594 283 !Additional output follows ............... EF1 = 0.83020 EF2 = 1.20080 X0 = 0.31601 Y0 = -0.03229 Z0 = 0.17469 X53 = -2.97089 Y53 = 3.82933 Z53 = -3.66482 Call 30 F 9.4960 R 51.599 F- 9.496 Rex 51.60 RD 0.00 Rx 0.00 594 283 PRED+2.PPAR.AP.(DMAX-AP) le ACCURACY Minimum Predicted, Accuracy = 0.4685E-01 Cond = 2 At Call 30 Filename for statistics ? Filename: [excora1.dat] ? [return] *** Correlation matrix and statistical errors *** EF1 EF2 X0 Y0 Z0 X53 Y53 Z53 EF1 1.00 -0.14 0.47 -0.27 0.42 0.16 0.00 0.17 EF2 -0.14 1.00 -0.23 0.00 -0.19 0.09 0.19 -0.13 X0 0.47 -0.23 1.00 0.38 0.58 0.04 -0.17 0.24 Y0 -0.27 0.00 0.38 1.00 -0.22 -0.33 -0.37 -0.23 Z0 0.42 -0.19 0.58 -0.22 1.00 0.38 0.33 0.65 X53 0.16 0.09 0.04 -0.33 0.38 1.00 0.52 0.39 Y53 0.00 0.19 -0.17 -0.37 0.33 0.52 1.00 0.69 Z53 0.17 -0.13 0.24 -0.23 0.65 0.39 0.69 1.00 DSTEP 2.721 2.721 0.021 0.021 0.021 0.021 0.021 0.021 Value of fit index at predicted minimum 9.4960 Fit index * k^2 : 0.9496 Rdistances : 0.0000 Chisqu : 0.1613E-05 Rexafs 51.5987 Rxray 0.0000 Rd 0.0000 !Repeat the refinement, avoiding all further prompts ENTER COMMAND: ref res 100;c;p;c;= 1 EF1 = 0.83020 +/- 0.58330 (2 sigma) 2 EF2 = 1.20080 +/- 0.43868 (2 sigma) 3 X0 = 0.31601 +/- 0.04502 (2 sigma) 4 Y0 = -0.03229 +/- 0.04876 (2 sigma) 5 Z0 = 0.17469 +/- 0.03559 (2 sigma) 6 X53 = -2.97089 +/- 0.02298 (2 sigma) 7 Y53 = 3.82933 +/- 0.04581 (2 sigma) 8 Z53 = -3.66482 +/- 0.03874 (2 sigma) Parameter refinement with correlation off: EF1 = 0.83020 EF2 = 1.20080 X0 = 0.31601 Y0 = -0.03229 Z0 = 0.17469 X53 = -2.97089 Y53 = 3.82933 Z53 = -3.66482 Call 1 F 9.4960 R 51.599 F- 9.496 Rex 51.60 RD 0.00 Rx 0.00 594 283 EF1 = 1.10232 EF2 = 1.20080 X0 = 0.31601 Y0 = -0.03229 Z0 = 0.17469 X53 = -2.97089 Y53 = 3.82933 Z53 = -3.66482 Call 2 F 9.5108 R 51.911 F- 9.496 Rex 51.91 RD 0.00 Rx 0.00 594 283 !Additional output follows ............... EF1 = 1.08041 EF2 = 0.84231 X0 = 0.35989 Y0 = -0.01875 Z0 = 0.20371 X53 = -2.97934 Y53 = 3.79551 Z53 = -3.66970 Call 32 F 9.2000 R 49.948 F- 9.198 Rex 49.95 RD 0.00 Rx 0.00 594 283 PRED+2.PPAR.AP.(DMAX-AP) le ACCURACY Minimum Predicted, Accuracy = 0.6715E-02 Cond = 2 At Call 32 *** Correlation matrix and statistical errors *** EF1 EF2 X0 Y0 Z0 X53 Y53 Z53 EF1 1.00 0.01 0.51 -0.31 0.42 -0.01 0.00 -0.01 EF2 0.01 1.00 0.01 0.00 0.00 0.11 0.21 -0.14 X0 0.51 0.01 1.00 0.20 0.55 -0.03 -0.01 0.01 Y0 -0.31 0.00 0.20 1.00 -0.38 -0.01 0.00 0.01 Z0 0.42 0.00 0.55 -0.38 1.00 -0.02 -0.01 0.01 X53 -0.01 0.11 -0.03 -0.01 -0.02 1.00 0.69 0.55 Y53 0.00 0.21 -0.01 0.00 -0.01 0.69 1.00 0.65 Z53 -0.01 -0.14 0.01 0.01 0.01 0.55 0.65 1.00 DSTEP 0.272 0.272 0.002 0.002 0.002 0.002 0.002 0.002 Value of fit index at predicted minimum 9.1981 Fit index * k^2 : 0.9200 Rdistances : 0.0000 Chisqu : 0.1563E-05 Rexafs 49.9477 Rxray 0.0000 Rd 0.0000 1 EF1 = 1.08393 +/- 0.52541 (2 sigma) 2 EF2 = 0.83260 +/- 0.38846 (2 sigma) 3 X0 = 0.35998 +/- 0.04687 (2 sigma) 4 Y0 = -0.01880 +/- 0.04269 (2 sigma) 5 Z0 = 0.20375 +/- 0.04408 (2 sigma) 6 X53 = -2.97803 +/- 0.02579 (2 sigma) 7 Y53 = 3.79946 +/- 0.06112 (2 sigma) 8 Z53 = -3.66646 +/- 0.05609 (2 sigma) ENTER COMMAND: !And again ...... ref res 150;c;p;c;= Parameter refinement with correlation off: EF1 = 1.08393 EF2 = 0.83260 X0 = 0.35998 Y0 = -0.01880 Z0 = 0.20375 X53 = -2.97803 Y53 = 3.79946 Z53 = -3.66646 Call 1 F 9.1981 R 49.963 F- 9.198 Rex 49.96 RD 0.00 Rx 0.00 594 283 !Additional output follows ............... EF1 = 1.18360 EF2 = 0.71829 X0 = 0.40260 Y0 = 0.01504 Z0 = 0.22739 X53 = -2.97873 Y53 = 3.79873 Z53 = -3.66719 Call 30 F 9.0944 R 49.019 F- 9.094 Rex 49.02 RD 0.00 Rx 0.00 594 283 PRED+2.PPAR.AP.(DMAX-AP) le ACCURACY Minimum Predicted, Accuracy = 0.2057E-02 Cond = 2 At Call 30 *** Correlation matrix and statistical errors *** EF1 EF2 X0 Y0 Z0 X53 Y53 Z53 EF1 1.00 0.01 0.29 -0.38 0.31 0.00 0.01 -0.01 EF2 0.01 1.00 0.00 -0.01 0.00 0.11 0.18 -0.17 X0 0.29 0.00 1.00 0.34 0.55 0.00 0.00 -0.01 Y0 -0.38 -0.01 0.34 1.00 -0.28 0.00 -0.01 0.01 Z0 0.31 0.00 0.55 -0.28 1.00 -0.01 0.00 -0.01 X53 0.00 0.11 0.00 0.00 -0.01 1.00 0.71 0.53 Y53 0.01 0.18 0.00 -0.01 0.00 0.71 1.00 0.65 Z53 -0.01 -0.17 -0.01 0.01 -0.01 0.53 0.65 1.00 DSTEP 0.027 0.027 0.000 0.000 0.000 0.000 0.000 0.000 Value of fit index at predicted minimum 9.0944 Fit index * k^2 : 0.9094 Rdistances : 0.0000 Chisqu : 0.1545E-05 Rexafs 49.0189 Rxray 0.0000 Rd 0.0000 1 EF1 = 1.18360 +/- 0.47498 (2 sigma) 2 EF2 = 0.71829 +/- 0.37500 (2 sigma) 3 X0 = 0.40260 +/- 0.04543 (2 sigma) 4 Y0 = 0.01504 +/- 0.05078 (2 sigma) 5 Z0 = 0.22739 +/- 0.04305 (2 sigma) 6 X53 = -2.97873 +/- 0.02577 (2 sigma) 7 Y53 = 3.79873 +/- 0.05970 (2 sigma) 8 Z53 = -3.66719 +/- 0.05730 (2 sigma)0 ENTER COMMAND: !Switch on MS ENTER COMMAND: s ms un ENTER COMMAND: ref res;c;p;c;= Parameter refinement with constraints off: EF1 = 1.18360 EF2 = 0.71829 X0 = 0.40260 Y0 = 0.01504 Z0 = 0.22739 X53 = -2.97873 Y53 = 3.79873 Z53 = -3.66719 Call 1 F 9.1004 R 47.329 F- 9.100 Rex 47.33 RD 0.00 Rx 0.00 594 283 EF1 = 1.45572 EF2 = 0.71829 X0 = 0.40260 Y0 = 0.01504 Z0 = 0.22739 X53 = -2.97873 Y53 = 3.79873 Z53 = -3.66719 Call 2 F 9.7217 R 48.225 F- 9.100 Rex 48.23 RD 0.00 Rx 0.00 594 283 !Additional output follows ............... EF1 = -1.87525 EF2 = -0.66724 X0 = 0.41083 Y0 = 0.02129 Z0 = 0.27967 X53 = -2.96679 Y53 = 3.80051 Z53 = -3.62263 Call 38 F 3.3382 R 32.309 F- 3.335 Rex 32.31 RD 0.00 Rx 0.00 594 283 Fit index is not decreasing *** Correlation matrix and statistical errors *** EF1 EF2 X0 Y0 Z0 X53 Y53 Z53 EF1 1.00 0.11 0.42 -0.39 0.35 0.14 0.31 -0.32 EF2 0.11 1.00 0.19 0.00 0.29 0.16 -0.02 0.03 X0 0.42 0.19 1.00 0.11 0.75 0.28 0.49 -0.12 Y0 -0.39 0.00 0.11 1.00 0.16 0.08 0.03 0.68 Z0 0.35 0.29 0.75 0.16 1.00 0.40 0.66 -0.03 X53 0.14 0.16 0.28 0.08 0.40 1.00 0.27 -0.14 Y53 0.31 -0.02 0.49 0.03 0.66 0.27 1.00 -0.36 Z53 -0.32 0.03 -0.12 0.68 -0.03 -0.14 -0.36 1.00 DSTEP 0.272 0.272 0.002 0.002 0.002 0.002 0.002 0.002 Value of fit index at predicted minimum 3.3348 Fit index * k^2 : 0.3338 Rdistances : 0.0000 Chisqu : 0.5671E-06 Rexafs 32.3089 Rxray 0.0000 Rd 0.0000 1 EF1 = -1.91691 +/- 0.24213 (2 sigma) 2 EF2 = -0.63885 +/- 0.27333 (2 sigma) 3 X0 = 0.41076 +/- 0.02465 (2 sigma) 4 Y0 = 0.02186 +/- 0.01918 (2 sigma) 5 Z0 = 0.27986 +/- 0.02877 (2 sigma) 6 X53 = -2.96714 +/- 0.01137 (2 sigma) 7 Y53 = 3.79991 +/- 0.01202 (2 sigma) 8 Z53 = -3.62449 +/- 0.01431 (2 sigma) !Draw the clusters again ENTER COMMAND: dr cl 1 dr cl 2 !Plot the result ENTER COMMAND: cp ENTER COMMAND: !Create a modified .pdb file ENTER COMMAND: pr par rep; Fit Index: with k^2 Weighting 0.3335 R-Factor : with k^2 Weighting 32.3093 Amplitude of Experiment 101.2935 R(exafs) = 32.3093 Weight: 1.00 R(distances)= 0.0000 Weight: 0.00 R(angles) = 0.0000 Weight: 0.00 N(ind) = 110.5 Np = 1. Chi^2 = 0.5665E-6 Filename: [exbrka5.pdb] ? File to be modified filename ? sod.pdb !sod.pdb is modified and written to exbrka5.pdb - sod.pdb is not itself changed Parameters printed in exbrka5.pdb Figure 6 - Cu site of Superoxide Dismutase Figure 7 - Result of refining metal atom positions
of Superoxide Dismutase Example 4. Tetrakis imidazole Copper(II) Nitrate Example 5. Simple demonstration of Surface parameters ! Define the symmetry of a four-fold hollow site sym c4v ! Define the surface cell c acell 2.4 ! Define the atom types c atom1 s;c atom2 ni;c atom3 s ! Define the shell parameters c ns 3;c t[1-2] ni;c t3 s c x1 1.2;c y1 1.2;c z1 -1;c n1 4 c x2 0;c y2 0;c z2 -2.2;c n2 1 c x3 2.4;c y3 2.4;c z3 0;c n3 4 ! Define the surface layer c zmin -1.5;c zmax .5 ! List the cartesian coordinates l car log EF = 0.000 VPI = 0.000 AFAC= 0.800 EMIN= 3.000 EMAX=1650.000 RMIN= 0.100 RMAX= 10.000 WIND= 2.000 WP = 0.100 NS = 3.000 Shell 1 N1 = 4.000 T1 = 2(NI) X1 = 1.200 Y1 = 1.200 Z1 = -1.000 Shell 2 N2 = 1.000 T2 = 2(NI) X2 = 0.000 Y2 = 0.000 Z2 = -2.200 Shell 3 N3 = 4.000 T3 = 3(S ) X3 = 2.400 Y3 = 2.400 Z3 = 0.000 ! List the radial coordinates l log EF = 0.000 VPI = 0.000 AFAC= 0.800 EMIN= 3.000 EMAX=1650.000 RMIN= 0.100 RMAX= 10.000 WIND= 2.000 WP = 0.100 NS = 3.000 Shell 1 N1 = 4.000 T1 = 2(NI) R1 = 1.970 A1 = 0.010 B1 = 0.000 Shell 2 N2 = 1.000 T2 = 2(NI) R2 = 2.200 A2 = 0.010 B2 = 0.000 Shell 3 N3 = 4.000 T3 = 3(S ) R3 = 3.394 A3 = 0.010 B3 = 0.000 ! Draw the structure dr ! Test the effect of DELTAZ c deltaz .1 l car log EF = 0.000 VPI = 0.000 AFAC= 0.800 EMIN= 3.000 EMAX=1650.000 RMIN= 0.100 RMAX= 10.000 WIND= 2.000 WP = 0.100 NS = 3.000 Shell 1 N1 = 4.000 T1 = 2(NI) X1 = 1.200 Y1 = 1.200 Z1 = -0.900 Shell 2 N2 = 1.000 T2 = 2(NI) X2 = 0.000 Y2 = 0.000 Z2 = -2.100 Shell 3 N3 = 4.000 T3 = 3(S ) X3 = 2.400 Y3 = 2.400 Z3 = 0.000 ! Reset DELTAZ c deltaz 0 ! Test the effect of SDX2 - the 2 refers to Nickel scattering atoms - not shell 2 c sdx2 .1 l car log EF = 0.000 VPI = 0.000 AFAC= 0.800 EMIN= 3.000 EMAX=1650.000 RMIN= 0.100 RMAX= 10.000 WIND= 2.000 WP = 0.100 NS = 3.000 Shell 1 N1 = 4.000 T1 = 2(NI) X1 = 1.440 Y1 = 1.200 Z1 = -1.000 Shell 2 N2 = 1.000 T2 = 2(NI) X2 = 0.000 Y2 = 0.000 Z2 = -2.200 Shell 3 N3 = 4.000 T3 = 3(S ) X3 = 2.400 Y3 = 2.400 Z3 = 0.000 c sdx2 0 ! Test the effect of SURREL c surrel 1.1 l car log EF = 0.000 VPI = 0.000 AFAC= 0.800 EMIN= 3.000 EMAX=1650.000 RMIN= 0.100 RMAX= 10.000 WIND= 2.000 WP = 0.100 NS = 3.000 Shell 1 N1 = 4.000 T1 = 2(NI) X1 = 1.200 Y1 = 1.200 Z1 = -1.000 Shell 2 N2 = 1.000 T2 = 2(NI) X2 = 0.000 Y2 = 0.000 Z2 = -2.080 Shell 3 N3 = 4.000 T3 = 3(S ) X3 = 2.400 Y3 = 2.400 Z3 = 0.000 c surrel 1 (Data kindly donated by Dr. Walkiria Schlindwein) The command file to perform these calculations is listed below, followed by a detailed description !Set up an empty site - as the ligand is bidentate, this will be used for the pivotal atom site e;c;0;= !Read the Fe K-edge data r ex;r41799.exb;1;32;fe k;;; !Set emergy limits to the range where background subtraction is adequate c emin 19.8;c emax 590 !Read in crystallographic parameters r par br;fedmpp1;1;1 !Remove the hydrogen atoms and sort in order of distance sort del 5 !Draw the original structure dr !Remove furthest atoms c ns 31 !Check the orientation of 'rotation axis' using the 3rd shell N atoms !(the molecule has approximate C3 symmetry) plane;23;24;25;0;0;1 !The axis is almost exactly parallel to z - no need to rotate it !Divide the cluster into 3 units, + one odd atom !(necessary because the .pdb file defined everything as one residue) c un[1-ns] 0 c un[1,4,7,9,13,16,20,23,26,29] 1 c un[2,6,8,11,14,17,21,24,27,30] 2 c un[3,5,10,12,15,18,22,25,28,31] 3 !Put the single O atom on the z-axis c th19 0 !Remove units 1 and 3, increase the occupation number for unit 2 c uoc[1,3] 0 c uoc2 3 !Remove empty shells sort del !Create the pivotal atom between atoms 1 and 2 c ns ns+1 c x[ns] .5*x1+.5*x2 c y[ns] .5*y1+.5*y2 c z[ns] .5*z1+.5*z2 !Ensure the occupation number and unit are the same as the other atoms !The occupation number should be changed first c n[ns] 3;c un[ns] 2 !Rename unit 2 as unit 1 c un[1-6,8-ns] 1 !Define the extra shell as the pivotal atom for unit 1 c piv1 ns !Define the symmetry sym c3 !Draw the symmetrised structure on top of the original dr noc !Define an atom parameter for the empty site c atom5 e !Define the atom type for the extra atom c t[ns] e !Calculate the potentials ca pot;C;O;1;FE;C;;; !Use the constant interstitial potential option s con v ca pot;C;O;1;FE;C;;; !Calculate the phaseshifts !The core-hole lifetime is adjusted, so the amplitude is correct !For AFAC=1 c cw .5 ca ph;;;;;;;;;;;;;;;;;;; !Set up rules for DW factors - makes refinements easier rule a2;a1 rule a4;a3 rule a6;a5 rule a9;a8 rule a10;a8 rule a11;a8 !Set DW factor limits c dwmin .003;c dwmax .025 !Set starting value for EF c ef -2 !Refine EF and A values without MS ref ef+a c;c;= !Set up MS parameters c atmax 5;c omax 5;c plmax 12 s ms units !Re-refine As ref a;c;c;= !Constrained refinement s cor on !Reduce MS using filter c minmag .005 !Refine the pivotal atom distance ref;r12;=;c;c; !Refine the DW factors ref a;c;c;= !Turn off constrained refinement s cor of !Use restrained refinement c wd .5;c wex .5 c d: c w: -5 1;1.5 c w: -2 1.5;2.3 fit ref x+y+z c;c; ref a;c;c;= !Remove filters and re-refine DW factors c minmag 0 ref a;c;c;= EXCURVE 9.26 >>> 29/09/02 at 06:12:21 ENTER COMMAND: Start the command file %stm Set up an empty site - as the ligand is bidentate, this will be used for the pivotal atom site e;c;0;= Read the Fe K-edge data r ex;r41799.exb;1;32;fe k;;; Set energy limits to the range where background subtraction is adequate c emin 19.8;c emax 590 Read in crystallographic parameters r par br;fedmpp1;1;1 Reading data with format: BROOKHAVEN (.pdb) Remove the hydrogen atoms and sort in order of distance sort del H Draw the original structure dr Remove furthest atoms c ns 31 Check the orientation of 'rotation axis' using the 3rd shell N atoms (the molecule has approximate C3 symmetry) plane;23;24;25;0;0;1 The axis is almost exactly parallel to z - no need to rotate it Divide the cluster into 3 units, + one odd atom (necessary because the .pdb file defined everything as one residue) c un[1-ns] 0 c un[1,4,7,9,13,16,20,23,26,29] 1 c un[2,6,8,11,14,17,21,24,27,30] 2 c un[3,5,10,12,15,18,22,25,28,31] 3 Put the single O atom on the z-axis c th19 0 Remove units 1 and 3, increase the occupation number for unit 2 c uoc[1,3] 0 c uoc2 3 Remove empty shells sort del Create the pivotal atom between atoms 1 and 2 c ns ns+1 c x[ns] .5*x1+.5*x2 c y[ns] .5*y1+.5*y2 c z[ns] .5*z1+.5*z2 Ensure the occupation number and unit are the same as the other atoms The occupation number should be changed first c n[ns] 3;c un[ns] 2 Rename unit 2 as unit 1 c un[1-6,8-ns] 1 Define the extra shell as the pivotal atom for unit 1 c piv1 ns Define the symmetry sym c3 Draw the symmetrised structure on top of the original dr noc Define the atom type for the extra atom c t[ns] e Calculate the potentials ca pot;C;O;1;FE;C;;; Z= 26 (FE) 1s1/2 1 2s1/2 2 2p1/2 2 2p3/2 4 3s1/2 2 3p1/2 2 3p3/2 4 3d3/2 4 3d5/2 3 4s1/2 2 - total 26 Z= 8 (O) 1s1/2 2 2s1/2 2 2p1/2 2 2p3/2 2 - total 8 NPTS = 349 JWS = 353 NEND = 451 NEND(2) = 451 Excited atom code : 1 Log grid increment : 0.02779 Lattice constant : 2.26000 Muffin-tin radius : 1.26000 349 Atomic volume : 10.88370 Wigner-seitz radius: 1.37477 352 Exchange constant : 0.66670 Madelung constant : 0.00000 I R RHO(AT) RHO(SUP) RV(AT) RV(SUP) RV(MT) V(MT)) 50 0.0006 0.0330 0.0330 -51.8887 -51.8887 -51.9117 -0.887E+05 100 0.0024 0.4607 0.4607 -51.5524 -51.5524 -51.6402 -0.220E+05 150 0.0094 4.8826 4.8826 -50.2211 -50.2211 -50.5279 -0.536E+04 200 0.0379 17.9767 17.9767 -45.4394 -45.4402 -46.1972 -0.122E+04 250 0.1520 34.1266 34.1275 -31.8463 -31.8519 -33.3602 -0.220E+03 300 0.6100 18.1612 18.1782 -11.1538 -11.1795 -13.2020 -0.216E+02 348 2.3158 1.3239 2.6169 -1.0125 -1.4249 -3.3026 -0.143E+01 349 2.3811 1.2571 2.7524 -0.9452 -1.4034 -3.3311 -0.140E+01 350 2.4482 1.1941 2.9309 -0.8808 -1.3916 -3.3779 -0.138E+01 FE (O ) Rho0: 0.4987 Efermi: -3.578 V0: -18.728 Electrons: 25.667 (Z= 26) etc. Calculate the phaseshifts The core-hole lifetime is adjusted, so the amplitude is correct For AFAC=1 c cw .5 ca ph;;;;;;;;;;;;;;;;;;; Set up rules for DW factors - makes refinements easier rule a2;a1 rule a4;a3 rule a6;a5 rule a9;a8 rule a10;a8 rule a11;a8 Set DW factor limits c dwmin .003;c dwmax .025 Set starting value for EF c ef -2 Refine EF and A values without MS ref ef+a Least squares refinement using k**2 weighting Initial parameters 1 EF -2.000 0.27212 0.00000 27.21164 -0.07350 2 A1 0.022 0.00063 0.00000 0.06301 0.35711 3 A3 0.022 0.00063 0.00000 0.06301 0.35711 4 A5 0.022 0.00063 0.00000 0.06301 0.35711 5 A7 0.022 0.00063 0.00000 0.06301 0.35711 6 A8 0.022 0.00063 0.00000 0.06301 0.35711 c;c;= EF = -2.00000 A1 = 0.02250 A3 = 0.02250 A5 = 0.02250 A7 = 0.02250 A8 = 0.02250 Call 1 F 15.3191 R 45.164 F- 15.319 Rex 45.16 RD 0.00 Rx 0.00 220 12 EF = -1.72788 A1 = 0.02250 A3 = 0.02250 A5 = 0.02250 A7 = 0.02250 A8 = 0.02250 Call 2 F 16.2460 R 46.154 F- 15.319 Rex 46.15 RD 0.00 Rx 0.00 220 12 EF = -2.00000 A1 = 0.02313 A3 = 0.02250 A5 = 0.02250 A7 = 0.02250 A8 = 0.02250 Call 3 F 15.7096 R 45.872 F- 15.319 Rex 45.87 RD 0.00 Rx 0.00 220 12 etc. EF = -3.62000 A1 = 0.00709 A3 = 0.02313 A5 = 0.02464 A7 = 0.01724 A8 = 0.00770 Call 74 F 3.8024 R 21.746 F- 3.802 Rex 21.75 RD 0.00 Rx 0.00 220 12 Minimum Predicted, Accuracy = 0.1083E-01 Cond = 2 At Call 74 *** Correlation matrix and statistical errors *** EF A1 A3 A5 A7 A8 EF 1.00 0.18 -0.35 -0.28 -0.12 0.19 A1 0.18 1.00 -0.13 -0.11 -0.08 0.05 A3 -0.35 -0.13 1.00 0.71 0.18 -0.55 A5 -0.28 -0.11 0.71 1.00 0.48 -0.66 A7 -0.12 -0.08 0.18 0.48 1.00 0.34 A8 0.19 0.05 -0.55 -0.66 0.34 1.00 DSTEP 0.272 0.001 0.001 0.001 0.001 0.001 Value of fit index at predicted minimum 3.8024 Fit index * k^2 : 0.3802 Rdistances : 0.0000 Chisqu : 0.1845E-05 Rexafs : 21.7463 Rxray : 0.0000 Rd : 0.0000 Best Values Are: 1 EF = -3.62719 +/- 0.23554 (2 sigma) 2 A1 = 0.00709 +/- 0.00108 (2 sigma) 3 A3 = 0.00770 +/- 0.00462 (2 sigma) 4 A5 = 0.02464 +/- 0.00564 (2 sigma) 5 A7 = 0.01724 +/- 0.07940 (2 sigma) 6 A8 = 0.02209 +/- 0.03739 (2 sigma) Set up MS parameters
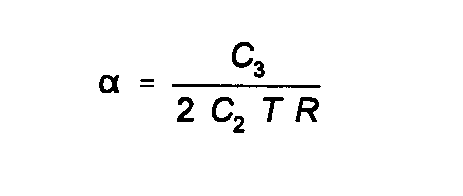
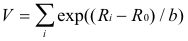
![]() (43)
(43)
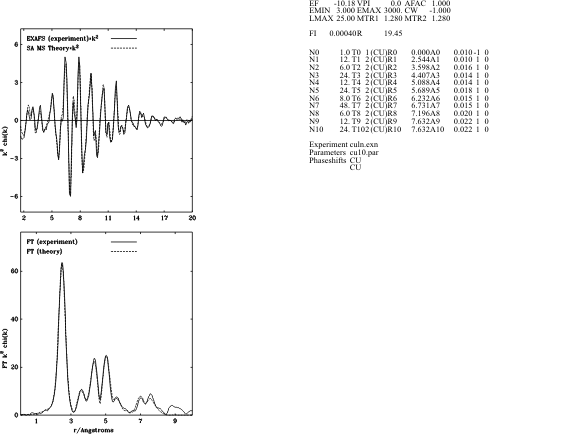
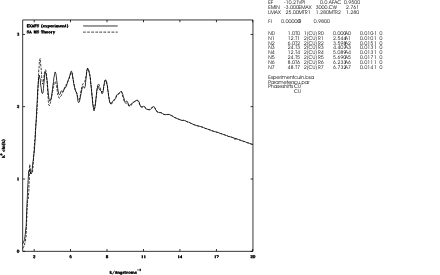
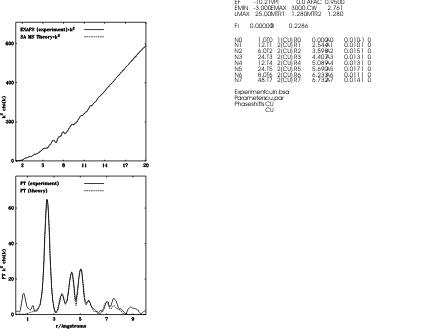
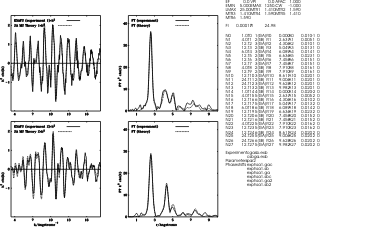
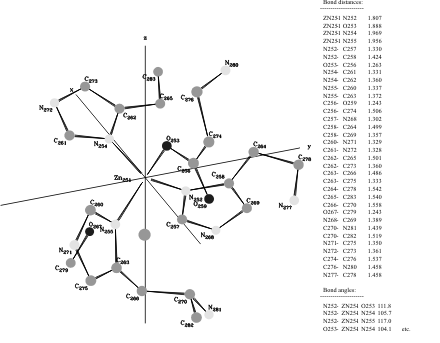
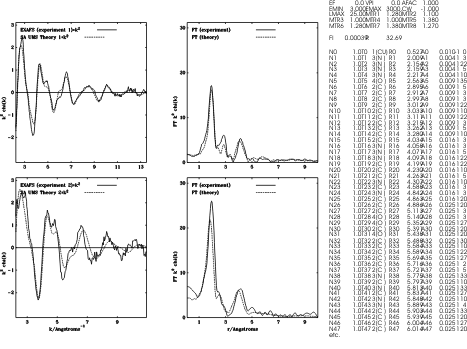
c atmax 5;c omax 5;c plmax 12
s ms units
Re-refine “A”s
ref a;c;c;=
A1 = 0.00709 A3 = 0.00770 A5 = 0.02464
A7 = 0.01724 A8 = 0.02209
Call 1 F 1.8720 R 14.555 F- 1.872 Rex 14.56 RD 0.00 Rx 0.00 220 12
A1 = 0.00729 A3 = 0.00770 A5 = 0.02464
A7 = 0.01724 A8 = 0.02209
Call 2 F 1.8830 R 14.511 F- 1.872 Rex 14.51 RD 0.00 Rx 0.00 220 12
A1 = 0.00709 A3 = 0.00792 A5 = 0.02464
A7 = 0.01724 A8 = 0.02209
Call 3 F 1.8790 R 14.609 F- 1.872 Rex 14.61 RD 0.00 Rx 0.00 220 12
etc.
A1 = 0.00676 A3 = 0.00621 A5 = 0.00638
A7 = 0.02450 A8 = 0.01274
Call 38 F 1.3343 R 12.559 F- 1.334 Rex 12.56 RD 0.00 Rx 0.00 220 12
Minimum Predicted, Accuracy = 0.1324E-02 Cond = 2 At Call 38
*** Correlation matrix and statistical errors ***
A1 A3 A5 A7 A8
A1 1.00 -0.03 0.00 0.01 0.03
A3 -0.03 1.00 -0.37 0.28 0.04
A5 0.00 -0.37 1.00 0.77 -0.12
A7 0.01 0.28 0.77 1.00 -0.31
A8 0.03 0.04 -0.12 -0.31 1.00
DSTEP 0.000 0.000 0.001 0.000 0.001
Value of fit index at predicted minimum 1.3343
Fit index * k^2 : 0.1334 Rdistances : 0.0000 Chisqu : 0.6473E-06
Rexafs : 12.5593 Rxray : 0.0000 Rd : 0.0000
Best values are:
1 A1 = 0.00676 +/- 0.00061 (2 sigma)
2 A3 = 0.00621 +/- 0.00198 (2 sigma)
3 A5 = 0.00638 +/- 0.00319 (2 sigma)
4 A7 = 0.02450 +/- 0.00058 (2 sigma)
5 A8 = 0.01274 +/- 0.00982 (2 sigma)
Constrained refinement
s cor on
Reduce MS using filter
c minmag .005
Refine the pivotal atom distance
ref;r12;=;c;c;
R12 = 1.54021
Call 1 F 1.2404 R 11.867 F- 1.240 Rex 11.87 RD 0.00 Rx 0.00 220 12
R12 = 1.54184
Call 2 F 1.2613 R 11.864 F- 1.240 Rex 11.86 RD 0.00 Rx 0.00 220 12
Minimum Predicted, Accuracy = 0.8771E-03 Cond = 2 At Call 2
Correlation matrix printed in excorb9.dat
Refine the DW factors
ref a;c;c;=
A1 = 0.00676 A3 = 0.00621 A5 = 0.00638
A7 = 0.02450 A8 = 0.01274
Call 1 F 1.2403 R 11.867 F- 1.240 Rex 11.87 RD 0.00 Rx 0.00 220 12
A1 = 0.00695 A3 = 0.00621 A5 = 0.00638
A7 = 0.02450 A8 = 0.01274
Call 2 F 1.2454 R 11.904 F- 1.240 Rex 11.90 RD 0.00 Rx 0.00 220 12
etc.
A1 = 0.00667 A3 = 0.00602 A5 = 0.00777
A7 = 0.02451 A8 = 0.01292
Call 10 F 1.2332 R 11.773 F- 1.233 Rex 11.77 RD 0.00 Rx 0.00 220 12
Minimum Predicted, Accuracy = 0.8771E-03 Cond = 2 At Call 10
*** Correlation matrix and statistical errors ***
A1 A3 A5 A7 A8
A1 1.00 -0.01 0.01 0.00 0.02
A3 -0.01 1.00 -0.10 -0.08 0.04
A5 0.01 -0.10 1.00 0.39 -0.17
A7 0.00 -0.08 0.39 1.00 -0.11
A8 0.02 0.04 -0.17 -0.11 1.00
DSTEP 0.000 0.000 0.000 0.001 0.000
Value of fit index at predicted minimum 1.2332
Fit index * k^2 : 0.1233 Rdistances : 0.0000 Chisqu : 0.5983E-06
Rexafs : 11.7731 Rxray : 0.0000 Rd : 0.0000
Best Values Are:
1 A1 = 0.00667 +/- 0.00058 (2 sigma)
2 A3 = 0.00602 +/- 0.00180 (2 sigma)
3 A5 = 0.00777 +/- 0.00309 (2 sigma)
4 A7 = 0.02451 +/- 0.00006 (2 sigma)
5 A8 = 0.01292 +/- 0.00787 (2 sigma)
Turn off constrained refinement
s con of
c wd .5;c wex .5
c d:
c w: -5
1;1.5
c w: -2
1.5;2.3
fit
Atom(I) Atom(J) Ideal Theory Difference Error % Weighting Error*Weight*1
1(O ) 0(FE) 1.998 1.998 0.0000 0.00 2.000 0.0000
2(O ) 0(FE) 2.046 2.046 0.0000 0.00 2.000 0.0000
3(C ) 1(O ) 1.346 1.346 0.0000 0.00 5.000 0.0000
4(C ) 2(O ) 1.288 1.288 0.0000 0.00 5.000 0.0000
4(C ) 3(C ) 1.410 1.410 0.0000 0.00 5.000 0.0000
5(C ) 3(C ) 1.400 1.400 0.0000 0.00 5.000 0.0000
6(C ) 4(C ) 1.424 1.424 0.0000 0.00 5.000 0.0000
8(C ) 5(C ) 1.469 1.469 0.0000 0.00 5.000 0.0000
9(N ) 5(C ) 1.362 1.362 0.0000 0.00 5.000 0.0000
10(C ) 6(C ) 1.352 1.352 0.0000 0.00 5.000 0.0000
10(C ) 9(N ) 1.368 1.368 0.0000 0.00 5.000 0.0000
11(C ) 9(N ) 1.476 1.476 0.0000 0.00 5.000 0.0000
12(E ) 0(FE) 1.540 1.540 0.0000 0.00 2.000 0.0000
12(E ) 1(O ) 1.310 1.310 0.0000 0.00 5.000 0.0000
12(E ) 2(O ) 1.310 1.310 0.0000 0.00 5.000 0.0000
12(E ) 3(C ) 1.365 1.365 0.0000 0.00 5.000 0.0000
12(E ) 4(C ) 1.368 1.368 0.0000 0.00 5.000 0.0000
Fit Index: with k^2 Weighting 0.0308
R-Factor : with k^2 Weighting 11.7731
Amplitude of Experiment 52.4801
R(exafs) = 11.7731 Weight: 0.50
R(distances)= 0.0000 Weight: 0.50
R(angles) = 0.0000 Weight: 0.00
N(ind) = 24.4 Np = 1. Chi^2 = 0.1357E-6
ref x+y+z
Least squares refinement using k**2 weighting
Initial parameters
1 X1 -1.411 0.00212 0.00000 0.21167 -6.66601
2 X2 -0.389 0.00212 0.00000 0.21167 -1.83776
3 X3 -1.759 0.00212 0.00000 0.21167 -8.31009
4 X4 -1.202 0.00212 0.00000 0.21167 -5.67863
etc.
30 Z11 -1.432 0.00212 0.00000 0.21167 -6.76522
c;c;
X1 = -1.41100 X2 = -0.38900 X3 = -1.75900
X4 = -1.20200 X5 = -2.63800 X6 = -1.56100
X8 = -3.19600 X9 = -2.95800 X10 = -2.42501
X11 = -3.85700 Y1 = -0.87100 Y2 = -1.62800
Y3 = -2.07800 Y4 = -2.45100 Y5 = -2.92700
Y6 = -3.72800 Y8 = -2.57400 Y9 = -4.12800
Y10 = -4.51100 Y11 = -5.08400 Z1 = -1.11400
Z2 = 1.17700 Z3 = -0.63000 Z4 = 0.61100
Z5 = -1.31300 Z6 = 1.12800 Z8 = -2.62500
Z9 = -0.75700 Z10 = 0.44300 Z11 = -1.43200
Call 1 F 0.3083 R 11.773 F- 0.308 Rex 11.77 RD 0.00 Rx 0.00 237 12
etc.
X1 = -1.40884 X2 = -0.38901 X3 = -1.75689
X4 = -1.20198 X5 = -2.63797 X6 = -1.55889
X8 = -3.19389 X9 = -2.95588 X10 = -2.42497
X11 = -3.85700 Y1 = -0.86891 Y2 = -1.62802
Y3 = -2.07794 Y4 = -2.44889 Y5 = -2.92696
Y6 = -3.72796 Y8 = -2.57191 Y9 = -4.12796
Y10 = -4.51099 Y11 = -5.08399 Z1 = -1.11391
Z2 = 1.17912 Z3 = -0.62796 Z4 = 0.61102
Z5 = -1.31297 Z6 = 1.13011 Z8 = -2.62498
Z9 = -0.75493 Z10 = 0.44305 Z11 = -1.43200
Call 35 F 0.3423 R 12.319 F- 0.303 Rex 12.20 RD 0.12 Rx 0.00 237 12
Correlation matrix printed in excorc1.dat
ref a;c;c;=
A1 = 0.00667 A3 = 0.00602 A5 = 0.00777
A7 = 0.02451 A8 = 0.01292
Call 1 F 0.3030 R 11.920 F- 0.303 Rex 11.80 RD 0.12 Rx 0.00 237 12
A1 = 0.00686 A3 = 0.00602 A5 = 0.00777
A7 = 0.02451 A8 = 0.01292
Call 2 F 0.3052 R 11.977 F- 0.303 Rex 11.86 RD 0.12 Rx 0.00 237 12
etc.
A1 = 0.00649 A3 = 0.00602 A5 = 0.00774
A7 = 0.02451 A8 = 0.01285
Call 7 F 0.3020 R 11.890 F- 0.302 Rex 11.77 RD 0.12 Rx 0.00 237 12
Minimum Predicted, Accuracy = 0.2143E-03 Cond = 2 At Call 7
*** Correlation matrix and statistical errors ***
A1 A3 A5 A7 A8
A1 1.00 -0.01 0.00 -0.02 0.01
A3 -0.01 1.00 -0.09 0.00 0.04
A5 0.00 -0.09 1.00 -0.01 -0.15
A7 -0.02 0.00 -0.01 1.00 -0.03
A8 0.01 0.04 -0.15 -0.03 1.00
DSTEP 0.000 0.000 0.000 0.001 0.000
Value of fit index at predicted minimum 0.3020
Fit index * k^2 : 0.3020E-01 Rdistances : 0.1158 Chisqu : 0.1329E-06
Rexafs : 11.7739 Rxray : 0.0000 Rd : 0.1158
1 A1 = 0.00649 +/- 0.00055 (2 sigma)
2 A3 = 0.00602 +/- 0.00169 (2 sigma)
3 A5 = 0.00774 +/- 0.00303 (2 sigma)
4 A7 = 0.02451 +/- 0.00003 (2 sigma)
5 A8 = 0.01285 +/- 0.00758 (2 sigma)
Remove filtersand re-refine DW factors
c minmag 0
ref a;c;c;=
A1 = 0.00649 A3 = 0.00602 A5 = 0.00774
A7 = 0.02451 A8 = 0.01285
Call 1 F 0.3302 R 12.683 F- 0.330 Rex 12.57 RD 0.12 Rx 0.00 237 12
A1 = 0.00667 A3 = 0.00602 A5 = 0.00774
A7 = 0.02451 A8 = 0.01285
Call 2 F 0.3306 R 12.690 F- 0.330 Rex 12.57 RD 0.12 Rx 0.00 237 12
etc.
A1 = 0.00651 A3 = 0.00589 A5 = 0.00632
A7 = 0.02451 A8 = 0.01444
Call 10 F 0.3275 R 12.747 F- 0.327 Rex 12.63 RD 0.12 Rx 0.00 237 12
Minimum Predicted, Accuracy = 0.2335E-03 Cond = 2 At Call 10
*** Correlation matrix and statistical errors ***
A1 A3 A5 A7 A8
A1 1.00 -0.01 0.01 0.00 0.03
A3 -0.01 1.00 -0.09 0.01 0.07
A5 0.01 -0.09 1.00 -0.14 -0.14
A7 0.00 0.01 -0.14 1.00 0.12
A8 0.03 0.07 -0.14 0.12 1.00
DSTEP 0.000 0.000 0.000 0.001 0.000
Value of fit index at predicted minimum 0.3275
Fit index * k^2 : 0.3275E-01 Rdistances : 0.1158 Chisqu : 0.1441E-06
Rexafs : 12.6309 Rxray : 0.0000 Rd : 0.1158
Best Values Are:
1 A1 = 0.00651 +/- 0.00057 (2 sigma)
2 A3 = 0.00589 +/- 0.00176 (2 sigma)
3 A5 = 0.00632 +/- 0.00293 (2 sigma)
4 A7 = 0.02451 +/- 0.00003 (2 sigma)
5 A8 = 0.01444 +/- 0.00787 (2 sigma)
fit
Atom(I) Atom(J) Ideal Theory Difference Error % Weighting Error*Weight*1
1(O ) 0(FE) 1.998 1.995 0.0024 0.12 2.000 0.0000
2(O ) 0(FE) 2.046 2.047 -0.0012 -0.06 2.000 0.0000
3(C ) 1(O ) 1.346 1.349 -0.0027 -0.20 5.000 -0.0001
4(C ) 2(O ) 1.288 1.287 0.0004 0.03 5.000 0.0000
4(C ) 3(C ) 1.410 1.407 0.0033 0.23 5.000 0.0002
5(C ) 3(C ) 1.400 1.402 -0.0024 -0.17 5.000 -0.0001
6(C ) 4(C ) 1.424 1.425 -0.0014 -0.10 5.000 -0.0001
8(C ) 5(C ) 1.469 1.468 0.0003 0.02 5.000 0.0000
9(N ) 5(C ) 1.362 1.362 -0.0004 -0.03 5.000 0.0000
10(C ) 6(C ) 1.352 1.354 -0.0014 -0.10 5.000 -0.0001
10(C ) 9(N ) 1.368 1.365 0.0027 0.20 5.000 0.0001
11(C ) 9(N ) 1.476 1.478 -0.0023 -0.15 5.000 -0.0001
Fit Index: with k^2 Weighting 0.0327
R-Factor : with k^2 Weighting 12.7468
Amplitude of Experiment 52.4801
R(exafs) = 12.6309 Weight: 0.50
R(distances)= 0.1158 Weight: 0.50
R(angles) = 0.0000 Weight: 0.00
N(ind) = 24.4 Np = 1. Chi^2 = 0.1441E-6
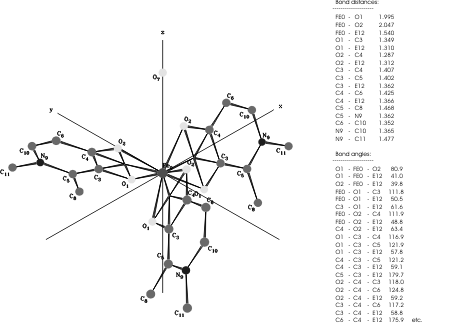
Figure 8 - Structure of Fe(DMPP)3, including dummy atoms for positioning the ligands.
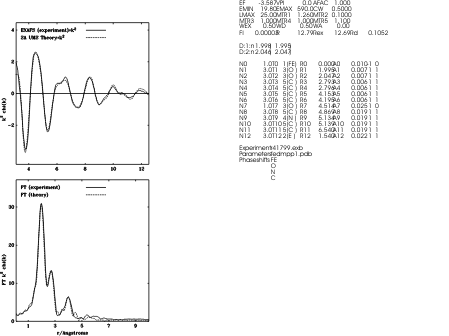
Figure 9 - Fit to Fe-edge of Fe(DMPP)3
4. Parameters and Related Topics
Parameters are of thirteen types:
1. Cell parameters ACELL,BCELL,ALPHA
2. Shell parameters are indexed by shell numbers - N1,T2,X3
3. Unit parameters are indexed by unit number - PANG1,PIV1,UOC2
4. Phaseshift parameters are indexed by atom type - MTR1,ALF2
5. Site parameters are indexed by site number - PERCA1, PERCB2
6. Spectrum parameters are either unindexed (common parameters) or indexed by spectrum number - WEX,EF1,AFAC2
There are five groups of unindexed parameters:
7. Theory parameters, including those concerned with phaseshift and multiple scattering calculations - LMAX, MINANG
8. Control parameters, influencing refinement, output etc. - BOND, SIZE, MAXRAD
9. Fourier transform parameters - WP, RMIN, RMAX
10. Surface parameters - ZMIN, DELTAX, SURREL
11. XANES parameters are used only by XANES calculations - XEMIN, XLOUT
12. Restrained refinement parameters are indexed by two shell numbers - D3:1,V5:0
13. Bond parameters are index by atom numbers - B253:72
Central atom - each of the clusters represented in the parameter table has one central atom. Cluster 1 has the central atom in shell 0, i.e. has coordinates X0, Y0, Z0 or R0, θ0, φ0. It is often, but not necessarily, at the centre of the coordinate system. The central atom of other clusters may occupy any shell, and is distinguished by a negative cluster number. The central atom is abbreviated by c below.
Absolute and Relative coordinates - the spatial coordinates X, Y, Z, R, θ and φ are all absolute coordinates. If the central atom is not at the origin in these coordinates the theory is determined by the relative coordinates, with the central atom at (0,0,0). It is possible to display, but not to directly change the relative coordinates using LIST RADIAL. Compare the cordinates given by this command with those given by L SPHERICAL and L CART. To avoid complications it is usually preferable to leave the central atom at absolute position (0,0,0) and to move the ligands. One occasion when this may not be the case is in protein refinements where the main-chain atomic positions are to be retained, but the central atom and ligands such as bicarbonate or hydroxyl are to be moved.
Clusters - the parameter table represents one or more clusters which consists of atoms surrounding crystallographically distinct excited atom sites. A cluster may also be an average over several such sites.
Shell - a shell consists of a set of atoms equidistant from the origin (usually the central atom but see above). Neither the θ nor φ values of the other atoms in the same shell need be the same. If the point symmetry of each central atom is defined, then the coordinates of all the atoms in a shell are defined. If no point symmetry is specified, then for the purpose of theory calculation the coordinates of all te atoms in a shell are assumed to be the same. Many program commands and options are unavailable if the symmetry is undefined.
Unit - a unit is a set of atoms, for example those comprising one ligand, identified by a common, non-zero, unit number. The unit is to allow constrained/restrained refinement, restricted (intra-unit) multiple scattering, and easy addition to, or removal of, ligands from a cluster.
Pivotal atom - the pivotal atom of a unit is the atom about which rotations of the unit are defined. If no pivotal atom is defined, the nearest atom to the centre is assumed. As the nearest atom may change giving unexpected results, it is advisable to set the parameter PIVn to one of the atoms in the unit (i.e. to a shell number) to avoid ambiguity. PIVn then defines the pivotal atom for unit n. In some circumstances, PIVn may be outside the unit (see torsion angles below), it does not then define the pivotal atom. The pivotal atom is abbreviated by p below. The vector c-p is abbreviated by u.
Unit plane - the unit plane is defined by p-a-b, where p is the pivotal atom, a is an atom defined by the unit parameter PLA and b is an atom defined by the unit parameter PLB. The unit plane is defined by the vector n which is a normal to the plane through p. If either p, a or b are undefined ( or equivalent ) then n is taken as a normal to u in the vertical plane. If p is undefined or u is parallel to the z-axis, n is taken as the z-axis.
Principal angle - the principal angle is the angle c-p-a. Its sign is determined by the sign of the z-component of the vector cross-product uX(a-p). If this is zero, the y- or if necessary the -x component is used.
1. Cell parameters (neither essential, nor used rigorously in EXCURVE, and mainly used for scaling of atomic distances)
ACELL crystallographic cell dimension a
BCELL crystallographic cell dimension b
CCELL crystallographic cell dimension c
ALPHA crystallographic cell parameter α
BETA crystallographic cell parameter β
GAMMA crystallographic cell parameter γ
R radial distance of one of the atoms in the shell (Å)
A Debye-Waller term 2σ2 (Å2) (or 2 C2 where C2 is the second cumulant of the pair distribution function)
C fourth cumulant C4 x100 ( Å4)
D exponent for truncated exponential disorder
UN number of the unit of which the shell is a part, or 0
ANG angle c-p-i for shell i. c = central, p = pivotal atom (0)
TH polar coordinate θ of one of the atoms in the shell (0)
PHI polar coordinate φ of one of the atoms in the shell (0)
X cartesian coordinate x of one of the atoms (Å)
Y cartesian coordinate y (Å)
Z cartesian coordinate z (Å)
ROT rotation about u (unit vector c-p) or VECA-VECB (0)
TWST rotation about n (normal to unit plane)(0)
TILT rotation about nXu (tilt axis)(0)
CLUS cluster to which shell belongs, -ve for an excited atom
LINK shell to which the shell is linked, in linked clusters
PLA atom a defining the unit plane, and the principal angle
PLB atom b defining the unit plane
TORA torsion angle ψ (N-CA-CB-CG)
TORB torsion angle ϖ (CA-CB-CG-CD)
TORC torsion angle at centre (c-p-a-b)
UOC unit occupancy - changes N for all atoms in the unit
All the unit parameters above can be changed using the command CHANGE. The torsion angle parameters can be refined using REFINE as well as being read from files. A number of other unit variables can only be modified by reading from the ligand database, or from a Brookhaven (.pdb) file or by changing them interactively using PLOT UNIT. These are:
NTORA[1-4] the atoms defining torsion angle A
NTORB[1-4] the atoms defining torsion angle B
RESNAME the name of the ligand ( residue in .PDB files)
4. Phaseshift parameters (indexed by ATOM TYPE)
ATOM atomic number associated with an atom type
ALF exchange term for X-α ground state exchange
CMAG Correction to Madelung potential
XE Atom-type specific correction to EF
The following parameters are, like the parameters above, indexed by atomi type, but are involved in changing the structure, not the phaseshifts
SDXi Displacement in x-direction for atoms of type i with Z between ZMIN and ZMAX
SDYi Displacement in y-direction for atoms of type i with Z between ZMIN and ZMAX
SDZi Displacement in z-direction for atoms of type i with Z between ZMIN and ZMAX
Other parameters which effect phaseshifts include EF0, RHO0 and V0 (see theory parameters).
Parameters used for adjusting the occupancy of mixed sites
PERCAi Fraction of component A in site i
PERCBi Fraction of component B in site i
PERCCi Fraction of component C in site i
Most of these variables have both a common term which is unindexed, and a term indexed by its spectrum number. The common variable applies to all spectra, the individual variables apply to only one. These are indexed by spectrum number, as used in READ EXP. Variables of this type are indicated by an optional index below. e.g. EF(i). A few variables must be indexed: these are indicated by BEWi etc.
EF(i) edge position (fermi energy), relative to calculated vacuum zero
VPI(i) energy-independent correction imaginary potential
AFAC(i) amplitude reduction due to many-electron processes etc.
EMIN(i) minimum energy for EXAFS theory calculations
EMAX(i) maximum energy for EXAFS theory calculations
WEX(i) weighting for EXAFS component of the fit-index
CW(i) the effective core-hole width, in eV. Theoretical values used if -1
BETH(i) beam direction - theta value, for spectrum i
BEPHI(i) beam direction - phi value, for spectrum i
BEW(i) polarisation azimuth, for experimental spectrum i
coefficients of pre-edge correction factor for whole-spectrum fitting
coefficients of polynomial correction factor for whole-spectrum fitting
In some instances it may be useful to combine common and individual variables - e.g., AFAC may be the same for all but one of the spectra. The rules for combining variables, in the case of spectrum i is as follows:
EF The program uses EF+EFi
AFAC The program uses AFAC*AFACi
EMIN The program uses MAX(EMIN,EMINi)
EMAX The program uses MIN(EMAN,EMANi)
VPI The program uses VPI+VPIi
WEX The program uses WEX*WEXi
BETH There is no BETH - it uses BETHi (similarly with BEPHI, BEW)
The common variables may also be written, for example, EF0. This can be useful in resetting variables. e.g.
C AFAC[0-3] 1
ATMAX maximum number of different atoms in a MS path
DE0 refinable correction to E0, for whole spectrum fitting
NS the number of shells used for calculations
LMAX maximum angular momentum for single scattering theory
KMIN minimum wave vector for EXAFS theory
KMAX maximum wave vector for EXAFS theory
OMIN minimum order of scattering for theory calculations
OMAX maximum order of scattering for theory calculations
DLMAX maximum angular momentum for double-scattering theory
TLMAX maximum angular momentum for triple-scattering theory
DWMIN minimum reasonable value of A
DWMAX maximum reasonable value of A
TEMP temperature (K) - used in calculating edge-width (whole-spectrum fitting) and C3 (if CLE ne 0)
CLE coefficient of linear expansion - if ne 0, used to calculate T-dependent C3 in anharmonic oscillator model
E0 absolute edge position - for whole-spectrum fitting only
NIND number of independent points in χ2 calculation
V0 global interstitial potential value for potential calculation (eV)
FE0 global Fermi energy value for potential calculation (eV)
RHO0 global charge density value for potential calculation (4/3πr2 x e/(bohr rad)3)
OFFSET fitting parameter defining the independent variables origin in the whole-spectrum polynomial correction
MINANG minimum angle for multiple scattering paths
MINDIST minimum distance for inter-atomic paths in multiple scattering paths
MINMAG minimum magnitude in principal path selection
PLMIN minimum pathlength for multiple scattering paths
PLMAX maximum pathlength for multiple scattering paths
D1A parameter A for use with CHANGE D1
D1B parameter B for use with CHANGE D1
D2A parameter A for use with CHANGE D2
D2B parameter B for use with CHANGE D2
OUTPUT determines the output level
BOND minimum bonding distance for plots etc.
SIZE atom size in plots (% of MTR)
VX x-coordinate of viewing position for 3D plots
VY y-coordinate of viewing position for 3D plots
VZ z-coordinate of viewing position for 3D plots
WD weighting for structural component of the fit-index
WA weighting for angle component of the fit-index
NGRID number of points in grid used for potential calculations
DFAC Relates maximum to initial step size in REFINE
MAXRAD maximum radius for inclusion of ligands from Brookhaven format file
NPARS number of refined parameters used in χ2 calculation
WNOISE k-weighting of noise component of theory
SPARE8 select specific scattering angular momentum components - must be -1 for curve-fitting
SPAR11 suppress pi-shift if 99 and controls the way matrix elements are calculated from complex potentials- should normally be 97
VECA shell A used in defining a rotation vector
VECB shell B used in defining a rotation vector
YMIN y-axis minimum for plots (for GSET RANGE YMIN/YMAX)
YMAX y-axis maximum
9. Fourier Transform parameters
RMIN minimum distance for Fourier transforms
RMAX maximum distance for Fourier transforms
WP Fourier transform window parameter
ZMIN minimum depth of surface layer
ZMAX maximum depth of surface layer
DELTAX substrate displacement in x-direction
DELTAY substrate displacement in y-direction
DELTAZ substrate displacement in z-direction
SURROT surface rotation - rotation of top layer of substrate
SURREL surface relaxation - increase in top layer spacing relative to bulk
XMIN minimum energy for XANES theory calculations
XMAX maximum energy for XANES theory calculations
XNP number of points in XANES calculations
XLMAX maximum angular momentum for XANES calculations
XLOUT maximum angular momentum for single centre expansion
XOPT determines treatment of central atom phase.
12. Restrained refinement parameters
These are indexed by two shell numbers, n and m
Dn:m ideal value of distance n-m, e.g. D3:2 is the distance between atoms in shells 3 and 2
Wn:m weighting for distance n-m
An:m ideal value of angle 0-n-m
Vn:m weighting for angle 0-n-m
These are indexed by two atom numbers, n and m. They are used only by DRAW
Bn:m if non-zero, indicates a bond between atom numbers n and m. The atom numbers are given by the ID column in SYMMETRY output.
1. The crystallographic cell parameters
The cell parameters ACELL, BCELL, CCELL, ALPHA, BETA, and GAMMA have a number of uses. They are used to add the cell outline to pictures created with DRAW, they are used to calculate and display fractional coordinates, and they are used to allow simultaneous changes to all shells in accordance with changes in the cell parameters. The cell parameters are subject to symmetry conditions, e.g. for cubic point groups (T, Td, O, Oh ) A=B=C, α=β=γ=90. For this reason the point symmetry of the cluster should be defined before they are changed, using the command SYMMETRY. As changing the cell parameters also moves atoms, it is also important that the cell parameters are entered before the atomic coordinates. If this procedure has not been followed, the best solution is to write the parameters to a file using PRINT PARAMS, and then edit the file so that the cell parameters are correct before reading it again. The fractional coordinates of atoms may be listed using the command LIST FRAC.
Normally, it is not necessary to enter actual values of the cell parameters. For example, to determine the lattice constant of a cubic metal, it is simply to necessary to refine ACELL. The extent it differs from the starting value of 1 will indicate the fractional change in the cell parameter compared to the stating value. This provides an easy way of refining the coordinates of all the atoms, without having to use constrained refinement (constraints on) or the RULE command.
One use of CCELL, is to simulate the effect of surface relaxation. If only adsorbate atoms and two layers of substrate atoms are included, then the inter-layer spacing will vary. The adsorbate-substrate spacing will also change as a result, but this can be corrected by adjusting DELTAZ as described below (surface parameters).
N If the point symmetry of the cluster is defined, the occupation numbers must be an integer number consistent with the symmetry position occupied. This may sometimes mean that a shell of atoms must be split into two - as for HCP structures where c/a is exactly (8/3)1/2.
T The atom type is stored as a number, which is the same as that used in reading phaseshift files and in indexing phaseshift variables, including the ATOMn variables, where the index n corresponds to the value of T. If the ATOM variables are defined, then T may be assigned an element symbol instead of a number.
R When the central atom of the cluster is at the centre of the coordinate system, the R values of all the atoms in a shell are there same. This is not the case, however, when the central atom has other coordinates.
ROT ROT defines a rotation about an axis, but there is no absolute reference for the rotation, so only the change in its value is significant. For this reason it is often simpler to enter commands such as C ROT13 ROT13+90. The ROT parameters provide a convenient means of changing the fundamental coordinates x, y, z, r, θ, and φ, they play no prat in the theory. If a new set of parameters is read while ROT parameters are non-zero, then the starting point for any changes to the new data will be non-zero. As this is potentially confusing, these parameters are set to zero whenever a new parameter file is read. The axis used is determined by the parameters VECA and VECB which determine two points defining the rotation vector. VECA is always a shell number ( 0 for central atom of cluster 1). VECB may be a positive shell number, negative, or zero:
VECB > 0 the second point is shell VECA ( not shell 0)
VECB = 0 the second point is the pivotal atom for the unit to which the shell belongs ( the unit must be defined).
VECB < 0 VECB is a code defining a direction relative to VECA. These codes are -1 for <100>, -2 for <010>, -3 for <001>, and -4 for <111>. What -5 does is a complete mystery.
Note that if, as is often the case, the central atom in shell 0 is required, this must be specified as VECA not VECB.
CLUS This parameter determines which cluster the shell belongs to and whether the shell is an excited atom or not. An excited atom is signified by a negative value.
LINK If two excited atoms are in close proximity, such as for Fe atoms in cubane, it would be normal to link one cluster with the other. This is achieved by setting the LINK parameter for a shell in one cluster to a shell in another, and defining an offset vector for the second cluster. This is normally done by means of the command GENERATE, but LINKS may be inspected or changed manually.
Many shell parameters are effected when constrained refinement is used. Details are discussed in the section Constrained Refinement in the program Guide (chapter 2).
Torsion angles - The program allows torsion angles to be defined and then changed for certain atoms within units. Changing a torsion angle, like constrained refinement, involves simultaneous movement of atoms within a unit. In this case however, three of the four atoms defining the dihedral angle remain stationary while the others rotate until the desired angle is obtained. Three torsion angles may be defined. By default one (TORC) is defined by atoms c-p-a-b, while the others (TORA and TORB) are undefined except when a Brookhaven ( or .pdb ) file is read, when the atoms N-CA-CB-CG and CA-CB-CG-CD are used (if there is no CG, NG or CG1 etc. may be used). The atoms defining the torsion angle for a unit may be changed at any time using PLOT UNIT which allows interactive changes to unit parameters. It is not relevant whether CONSTRAINTS is on or off when changing or refining torsion angles. Note that the program has no sense of which part of the unit is to remain stationary during rotation. The atoms defining the rotation axis (e.g. -CA-CB- ) will of course be unchanged. Otherwise those with residue names C, N, O, CA and the central atom, will remain unchanged, all others will be rotated. For this reason torsion angles are not generally used except in conjunction with parameters read from .pdb or .car files, or with structures created using the ligand database.
In order to make TORCi more useful. p is taken as PIVi, even if it is not within a unit. This has a special application in moving the side chains of complex polydentate ligands.
The ligand database - The ligand database resides in the file ideals and is based on the database used by the crystallographic refinement program prolsq. It has been extended to include non-amino acid ligands and information for defining the default relationship to the central atom for use in EXAFS analysis. In order to use the database the shell for the pivotal atom and the coordinates of the pivotal atom should be selected. E.g.:
C X3 1.3;C Y3 1.3;C Z3 0.
The ligand can then be selected using:
C T3 (ligand name)
The ligand name may be an amino acid symbol (TYR, HIS etc.) or a chemical formula (CO3, SO4). A list of current names may be obtained using:
C T3 ?
The ligand will be assigned to the next vacant unit number. The pivotal atom will be assigned to the specified shell, and shell numbers above NS will be used for the other atoms. The ligand name will be used for the unit name. The unit parameters PIV, PLA, PLB, will be set up, as will any relevant torsion angles. Bond angles and torsion angles will be defined from database information. The only undefined aspect of the ligand is the rotation about the vector u. This may be changed using the ROT parameter with VECA=VECB=0. As ROT has no absolute significance it is not possible to use the current setting to define the orientation. This coordinate has no effect on the theory unless inter-unit multiple scattering or XANES calculations are performed.
MTR values are set to default values (dependent on ION values), whenever ATOM values are changed. They may thereafter be changed or refined. Potentials and phaseshifts are automatically updated during refinements, but not when MTR values are changed manually If the COMMON V, EF or RHO options are in use, the Muffin-tin radius is initially set to the corresponding MTR value. The value of the Muffin-tin radius is then refined. The value used is written to the terminal and logfile during potential calculations, and to the phaseshift files genereated by PRINT PHASE, but the values of MTR variables are not changed..
At present, the atoms associated with a site definition can only be defined by the SITE command, whereas the occupancy factors, PERCA1 etc. are parameters. The PERC parameters are only meaningful within the context of the site definition. It is therefore necessary to define the site before reading a parameter file, which contains no record of either the site name or its constituent atoms.
The ATOM parameters which allow T parameters to refer to the sites, are stored as numbers, not simbols. The order in which the sites are defined must therefore be the same as that in which they are used in any parameter files that are read .For further discussion of site parameters see the section Mixed Sites in the Program Guide (Chapter 2).
6a. Spectrum parameters - beam polarisation parameters
Beam parameters are used in polarised (single crystal or surface) EXAFS to describe the orientation of the beam in terms of the axial system used to describe the crystal. There are sets of beam parameters for each experimental spectrum.
(e.g., BETH1, BETH2, BETH3). The beam orientation is given by BETH1 and BEPHI1 etc., the polarisation direction by BEW1. The default settings are for the polarisation e vector parallel to x, y and z respectively. The parameter values and the corresponding e vector directions are shown below.
l spars
Spectrum 1 Spectrum 2 Spectrum 3
BETH1 = 0.000 BETH2 = 0.000 BETH3 = 90.000
BEPHI1= 0.000 BEPHI2= 0.000 BEPHI3= 0.000
BEW1 = 90.000 BEW2 = 0.000 BEW3 = 90.000
EX1 = 1.000 EX2 = 0.000 EX3 = 0.000
EY1 = 0.000 EY2 = 1.000 EY3 = 0.000
EZ1 = 0.000 EZ2 = 0.000 EZ3 = -1.000
ETH1 = 90.000 ETH2 = 90.000 ETH3 = 180.000
EPHI1 = 0.000 EPHI2 = 270.000 EPHI3 = 0.000
7a. Theory parameters - multiple scattering parameters
A number of parameters apply only to MS. DLMAX and TLMAX determine the number of angular momentum terms for double and triple scattering respectively. Although the number of terms required is usually less than for single scattering, the maximum allowed values (often 12 and 9, although some versions allow 15 and 12), may not be sufficient for heavy atom scatterers, and care should be taken in reducing these parameters, even for light atoms.
Several parameters restrict the number of paths that are calculated. These parameters can also be used to isolate contributions using EXTRACT if a full set of paths have been previously calculated.
PLMIN is the minimum path length ( usually 0 )
PLMAX is the maximum path length (default is 10)
MINANG is the minimum angle - scattering within a triplet of atoms, or a quadruplet derived from it, will not be calculated unless one of the bond angles exceeds this value.
OMIN is the minimum order - default is 1
OMAX is the maximum order - default 3, maximum 5 - use of a higher order will considerably slow down the calculations.
ATMAX determines the maximum number of scattering atoms in a path - default is 2. 3, 4 or 5 may be selected at high computational expense.
MINMAG determines a cut-off below which previously calculated paths will be ignored. Default is 0. This should be used with great care. If one of the above parameters is changed, or any bond length changes by more than a few percent, the path indices will change, causing the wrong paths to be omitted. Path indices may be inspected using EXTRACT. A safe way to use this, is to set MINMAG to 0, calculate a spectrum with small atom or low DLMAX and TLMAX, set MINMAG to a sensible cut-off, and recalculate the spectrum using a higher angular momentum.
MINDIST determines the minimum permitted distance in any MS path. When site vacancies are being used it can be used to avoid including a disordered site twice in the same path.
The command LIST MSPARS will give the current values of most multiple scattering parameters.
When using the SMALL_ATOM option (SET THEORY SMALL_ATOM), the precision may be selected using NUMAX, which determines the size of the scattering matrix. 0 uses the approximation of Gurman (1988), 1 and 2 use the Rehr and Albers (1990) method, where the value of NUMAX determines the matrix size.
7b. Theory parameters - energy parameters
7c. Theory parameters - whole-spectrum parameters
BOND has several uses in the program. It determines which bonds are included in plots produced by DRAW, and which distances and angles are included in the orresponding tables. It has a similar function in PLOT UNIT, and is also used to determine which shells are included in calculating bond valence sums. If BOND is zero, a value slightly greater than the first shell distance of the first cluster is assumed for DRAW and PLOT. When reading parameters in Brookhaven .pdb format, then if BOND is zero, it will be changed to a value slightly greater than the first shell distance.
OUTPUT has a default value of 0. Increasing its value will increase the amount of output to both the terminal and the log file. It is a good idea to use a value of at least 1 for long MS calculations, so the program does not appear to hang up.
9. Fourier transform parameters
RMAX effects the range in R-space of plots produced by PLOT FT, COPLOT and PLOT PATHS, but not FFILTER, which has other mechanisms for determining the range in R-space.
10a. Surface parameters - surface displacement parameters
The general surface displacement parameters DELTAX, DELTAY, DELTAZ and those indexed by atoms type, the surface layer displacement parameters - SDX, SDY, and SDZ, are used for correlated movement of surface atoms. They are specific to surfaces, are indifferent to the status of the CONSTRAINTS option and do not require the use of units. One other thing they have in common is that their origin is undefined. They have no absolute meaning and are provided as a convenient means of changing the fundamental coordinates x, y, z, r, θ, and φ.
DELTAX etc. will move all atoms other than those in the z >=0 adsorbate layer in the required direction (in the cartesian rather than the crystallographic cell). For example DELTAX or DELTAY will simulate movement of the adsorbate off a high symmetry site, while DELTAZ will alter the adsorbate-substrate distance. Note that because all atoms, including those in special positions, are moved, the use of these parameters is highly subject to the point symmetry of the cluster. Movement of DELTAZ is not permitted in point groups with a centre, a horizontal mirror plane, a horizontal diad or cubic symmetry. However, as these are improbably in a surface simulation these restrictions should not prove a problem. DELTAX and DELTAY however are incompatible with a principal axis higher than 1, and are effectively restricted to use with ci and cx. cx is a specially defined point group symbol representing cσ in an alternative orientation with the mirror plane parallel to x. As DELTAX etc. have no absolute origin as a reference point, only changes in its value are relevant, so atoms are moved by the difference between the new and old values.
SDX etc. operate in a manner analogous to DELTAX etc.. The differences are that SDX only moves atoms in the top substrate layer, or surface layer, and is indexed by atom type, so that only certain atoms are moved. The surface layer is defined to mean any atom with a z-coordinate between ZMIN and ZMAX angstroms. If the atoms to be moved are in a general position then there are no symmetry restrictions on which movements are allowed. If atoms are in a special position with respect to a point symmetry operator, such as lying on the principal axis, then symmetry restrictions will apply. These will not be checked when the change is made, as with DELTAX, and will result in errors the next time the coordinates of all the atoms are expanded using the point group symmetry. If it is necessary to distinguish crystallographically distinct sites occupied by the same atom type, it will be necessary to duplicate phaseshifts, and assign two different atom types to the same element.
The action resulting from such movements will clearly depend on which of the set of symmetry related atoms are given in the table of shell parameters. With DELTAX the symmetry rules ensured that all the atoms were translated in the same direction. With SDX however, the action of vertical mirror planes will cause contrary motion of individual atoms and non-unitary principal axes will cause rotations. Indeed, the principal use of these variables is to simulate surface reconstructions which involve rotations of atoms, in contrary directions for adjacent adsorbate sites. The effect of the symmetry operator will be different depending on which atom in a shell has been selected for each shell ( they will simulate the effect of different crystallographic space groups). In general, only one of the possible choices is correct, and this will not be the case which is usually selected. An example of a parameter table ( for reconstructed p4g N on Ni(100) is given below.
l car
E0 = 3.000 VPI = -4.000 AFAC= 0.800 EMIN= 3.000 EMAX=3000.000
RMIN= 0.100 RMAX= 10.000 WIND= 2.000 WP = 0.100 NS = 10.000
Shell 1 N1 = 4.000 T1 = 28(NI) X1 = 2.400 Y1 = 0.000 Z1 = -1.000
Shell 2 N2 = 4.000 T2 = 28(NI) X2 = -2.400 Y2 = 4.800 Z2 = -1.000
Shell 3 N3 = 4.000 T3 = 28(NI) X3 = -2.400 Y3 = -4.800 Z3 = -1.000
Shell 4 N4 = 4.000 T4 = 28(NI) X4 = -7.200 Y4 = 0.000 Z4 = -1.000
Shell 5 N5 = 4.000 T5 = 8 (O ) X5 = 0.000 Y5 = 4.800 Z5 = 0.000
Shell 6 N6 = 4.000 T6 = 8 (O ) X6 = 4.800 Y6 = 4.800 Z6 = 0.000
Shell 7 N7 = 1.000 T7 = 28(NI) X7 = 0.000 Y7 = 0.000 Z7 = -2.200
Shell 8 N8 = 4.000 T8 = 28(NI) X8 = 4.800 Y8 = 0.000 Z8 = -2.200
Shell 9 N9 = 4.000 T9 = 28(NI) X9 = 2.400 Y9 = 2.400 Z9 = -2.200
Shell 10 N10= 4.000 T10= 28(NI) X10 = 4.800 Y10 = 4.800 Z10 = -2.200
l frac
Cell parameters: 9.6000 9.6000 1.0000
Shell X Y Z
0 O 0 0.0000 0 0.0000 0 0.0000
1 NI 0 0.2500 0 0.0000 -1 0.0000
2 NI -1 0.7500 0 0.5000 -1 0.0000
3 NI -1 0.7500 -1 0.5000 -1 0.0000
4 NI -1 0.2500 0 0.0000 -1 0.0000
5 O 0 0.0000 0 0.5000 0 0.0000
6 O 0 0.5000 0 0.5000 0 0.0000
7 NI 0 0.0000 0 0.0000 -3 0.8000
8 NI 0 0.5000 0 0.0000 -3 0.8000
9 NI 0 0.2500 0 0.2500 -3 0.8000
10 NI 0 0.5000 0 0.5000 -3 0.8000
c sdy2 .02
l car
EF = 3.000 VPI = -4.000 AFAC= 0.800 EMIN= 3.000 EMAX=3000.000
RMIN= 0.100 RMAX= 10.000 WIND= 2.000 WP = 0.100 NS = 10.000
Shell 1 N1 = 4.000 T1 = 28(NI) X1 = 2.400 Y1 = 0.192 Z1 = -1.000
Shell 2 N2 = 4.000 T2 = 28(NI) X2 = -2.400 Y2 = 4.992 Z2 = -1.000
Shell 3 N3 = 4.000 T3 = 28(NI) X3 = -2.400 Y3 = -4.608 Z3 = -1.000
Shell 4 N4 = 4.000 T4 = 28(NI) X4 = -7.200 Y4 = 0.192 Z4 = -1.000
Shell 5 N5 = 4.000 T5 = 8 (O ) X5 = 0.000 Y5 = 4.800 Z5 = 0.000
Shell 6 N6 = 4.000 T6 = 8 (O ) X6 = 4.800 Y6 = 4.800 Z6 = 0.000
Shell 7 N7 = 1.000 T7 = 28(NI) X7 = 0.000 Y7 = 0.000 Z7 = -2.200
Shell 8 N8 = 4.000 T8 = 28(NI) X8 = 4.800 Y8 = 0.000 Z8 = -2.200
Shell 9 N9 = 4.000 T9 = 28(NI) X9 = 2.400 Y9 = 2.400 Z9 = -2.200
Shell 10 N10 = 4.000 T10= 28(NI) X10 = 4.800 Y10 = 4.800 Z10 = -2.200
l frac
Cell parameters: 9.6000 9.6000 1.0000
Shell X Y Z
0 O 0 0.0000 0 0.0000 0 0.0000
1 NI 0 0.2500 0 0.0200 -1 0.0000
2 NI -1 0.7500 0 0.5200 -1 0.0000
3 NI -1 0.7500 -1 0.5200 -1 0.0000
4 NI -1 0.2500 0 0.0200 -1 0.0000
5 O 0 0.0000 0 0.5000 0 0.0000
6 O 0 0.5000 0 0.5000 0 0.0000
7 NI 0 0.0000 0 0.0000 -3 0.8000
8 NI 0 0.5000 0 0.0000 -3 0.8000
9 NI 0 0.2500 0 0.2500 -3 0.8000
10 NI 0 0.5000 0 0.5000 -3 0.8000
If a new parameter file is read while DELTAX, SDX etc. are non-zero, then the starting point for any changes to the new data will be non-zero. As this is potentially confusing, these parameters are set to zero whenever a new set of parameters is read.
10b. Surface parameters - the surface relaxation parameter SURREL
SURREL alters the surface relaxation - that is, the distance between the top two substrate layers of a surface relative to the bulk interlayer spacing. The default value is 1. As with SDX etc. the surface layer is defined as that between ZMIN and ZMAX. When changing SURREL care should be taken to ensure that as a result of the change, atoms within the surface layer do not move out of it, and that atoms outside the surface layer do not move into it.
12. Restrained refinement parameters
13. Bond parameters
ALIAS is used to define commands.
Syntax: ALIAS alias_name [DEL]
ALIAS [LIST]
ALIAS DELETE [n]
Abbreviations: AL
This command allows users to define their own commands or abbreviations. For example:
ALIAS TEMPCOM
results in the prompt:
Enter definition of TEMPCOM:
You can then type in any command or list of commands e.g.:
gset dev 2;gset plotter ps_file
and for the rest of the session TEMPCOM will result in the execution of the list of commands.
If LIST is specified or else the command is entered without a keyword, then ALIAS lists the current aliases.
The number of aliases which may be defined is displayed using LIST DIMENSION. An alias may be up to 80 characters long. If the maximum number of aliases are already defined, the last n may be deleted using ALIAS DELETE n. If n is unspecified, the default is 1. DELETE may not be abbreviated, so that alias names such as DEL are allowed.
A specific alias may be deleted using:
ALIAS alias_name DEL
Aliases containing special characters such as *, % and ! should be enclosed in quotes:
ALIAS LL;'^ls -l'
Once defined, an alias may be used whenever a command name may be used.
An alias name must not conflict with an existing command or abbreviation. For example :
ALIAS R;C R1 2
Will have no effect, as R will always be interpreted as READ, for which it is a permitted abbreviation.
ANGLE is used to calculate an angle with vertex (0,0,0)
Syntax: ANGLE [cluster_number]
Abbreviations: AN
This command calculates the angle at (0,0,0) for two atoms. The program requests the spherical polar coordinates for two atoms. Normally the TH and PHI parameters for two shells are used, but any value or expression is valid.
ANGLE
results in the prompt:
Enter theta for atom A
TH1
......
Enter phi for atom B
PHI4
Will calculate the angle 1-0-4
CALC is used to calculate potentials and phaseshifts.
Syntax: CALC POTENTIALS n
CALC PHASESHIFTS n
Abbreviations: CA PO/PH
Option: atom number n
If the number n is defined then the calculation is performed for that atom number only (the atom numbers are those used in Tn parameters and elsewhere, usually 1 for the central atom, 2 for the first scattering atom etc.). If n is negative then the calculation is done for an atom of that type treated as a central excited atom. If n is omitted then the calculations are done for all defined atom types. Atom types are defined when the corresponding ATOMn variables are set to an atomic symbol or number.
e.g. C ATOM1 CU;C ATOM3 O;CA POT
Will calculate potentials for the Cu central atom (n=1) and the O scattering atom (n=3). The n=2 potential will be omitted unless ATOM2 has been previously defined.
Atom 1 is always an excited atom. If multiple edges are being fitted, additional excited atoms are required. These may be specified using:
CA POT -4
To make atom 4 an excited atom, or better, by initially defining the relevant atom variables as excited atoms.
C ATOM1 GA*;C ATOM4 SB*
For the Ga and Sb edges of GaSb. This procedure may be made mandatory in future releases. Atoms are also defined as excited atoms, if excited atom phaseshifts are read for that atom number. However they are defined, excited atoms are indicated by a * symbol when using LIST PHASE.
The potentials or phaseshifts are calculated using the approximations and methods outlined in the theory section.
Options which apply to potential calculations include those available through SET GROUND_STATE, SET EXCHANGE, SET COMMON ( see the help on SET ).
Options which apply to phaseshift calculations include those available through SET QUINN, SET RELCOR ( see the help on SET ).
Variables which apply to potential calculations for atom n, include ATOMn, MTRn, IONn, and CMAGn. Other variables affecting potential calculations, when using COMMON options, include V0, FE0 and RHO0.
The principal variables effecting phaseshift calculations are the core width parameters CW, CW1, CW2 etc. If CW is defined (i.e. if it is positive or zero), it will be used as the default core width for all phaseshift calculations. If CW1 etc. are defined (i.e. if they are positive or zero), they will be used as defaults for phaseshifts relating to experiment number 1 etc. These parameters should be set to -1 if not required, when program defaults will be used. The default values may be modified when CALC PH is executed.
CALC issues a number of prompts which are described in the sections Calculating Potentials and Calculating Phaseshifts of the program guide section, chapter 2. An example potential calculation is described in the examples section, chapter 3.
For Whole-spectrum fitting - using SET ATOMIC_ABS ON, slightly different rules apply - a potential will be calculated for an 'ATOM 0'. This duplicates the potential for the central atom, using an entirely real potential, as the treatment of inelastic effects is inappropriate for calculation of dipole transition rates. It is not necessary to change 'ATOM0' - there is no such variable. For this reason, the option to calculate the atomic absorption for a complex potential should be regarded as redundant.
CCHANGE is used to change multiple atomic coordinates using constraints, irrespective of the current status of the CONSTRAINTS option.
Syntax: CCHANGE variable_name new_value
Abbreviations: CC
Notes:
CCHANGE R1 is exactly equivalent to using
SET CONSTRAINTS ON;C R1;SET CONSTRAINTS OFF
All the atoms in the same unit as R1 will be moved so as to preserve the geometry of the unit. See documentation sections on units and constraints.
Note that if this method is used, subsequent refinements will not be constrained. The command is most useful in getting an approximate fit 'by hand' before using restrained refinement (q.v.).
CHANGE is used to alter the values of the parameters described in section 4.
Syntax: CHANGE variable_name new_value
CHANGE CENTRE [cluster_number]
Abbreviations: CH or C
Notes:
The effect is to change the value of variable_name to new_value.
Variable_name may be one of:
a) A parameter name, R1, EMIN etc. ( see the parameters section for a full list). Note that the program will not allow some parameters to be changed to values outside a specified range. Some parameters ( EMIN, EMAX, EF, KMIN, KMAX ) are interconnected and may change if you change the others (The relationship between KMIN and EMIN depends not only on EF, but also on the energy dependent self-energy, and may change when the theory is updated). New_value is usually a number, but in the case of parameters such as Tn, ATOMn new_value may be either an atomic number or chemical symbol:
C ATOM1 CU is equivalent to C ATOM1 29
New_value may always be an arithmetic expression including other parameters, as in:
C R2 R1*1.5
Parameters D1A, D1B, D2A and D2B are CHANGEd to the name of another parameter ( see d) below ).
b) A parameter list, R[1-3] etc. This format may be used with shell parameters N, T, R, A, UN, ANG, TH, PHI etc., with phaseshift parameters, MTR, ATOM etc., and with other indexed parameters such as memories, M1 etc.
Values may be changed for a range of shells. The format is similar to the Unix 'wildcard' format for filenames. The list is placed in square brackets [] after the parameter name. Thus 'CHANGE N[1-4] 7' will change N1, N2, N3 and N4 to 7. Several ranges may be specified in one set of [] by separating them with commas. Thus N[1,3-5,7] refers to N1, N3, N4, N5 and N7. Parameters may also be used inside the []. Thus N[1-NS] refers to all N values from 1 to the current value of NS. When used in this way CHANGE only reports the first and last values changed.
Identical rules apply to parameter lists in the command REFINE.
c) A shell symbol, S1, S2, S3, etc. New_value should be the number of the shell whose parameter values you wish to copy to the shell being changed. e.g.:
C S2 1
Will produce the result that shell 2 is identical to shell 1.
If new_value is unspecified, you will be prompted for new values for all the parameters of that shell. Enter '=' when sufficient parameters have been changed. This allows all the parameters for a shell to be entered using a single CHANGE command.
d) DF1 or DF2. The difference between parameters D1A and D1B or D2A and D2B will be set to new_value. D1A and D1B must previously have been set to the names of two parameters indexed by shell. If DF1 is set then during an REFINE or MAP the difference between the two parameters D1A and D1B will be maintained. Thus 'C D1A R1; C D1B R2; C DF1 1.0 ' will maintain a distance of 1.0 between shell 1 and shell 2. This can be useful if there are close peaks in an
EXAFS spectrum.
e) Table symbols Dn:m or Wn:m ( n and m integers ). These parameters are used in restrained refinement. 'C D1:0 3.4' for example sets the distance between shell 1 and the central atom to be 3.4; 'C D3:2 2.6' sets the distance from shell 3 to shell 2 to be 2.6. ( The larger shell index must come first. ) Wn:m is the weight given in restrained refinement to the distance Dn:m. (See the section on restrained refinement for more details.)
Special notation permits multiple changes:
C D:
Sets all restrained distances equal to the corresponding current inter-shell distance.
C W: 1
Sets all the weights to 1, unless the corresponding Dn:m is zero.
f) Table symbols Bn:m (n and m integers). These parameters are used in specifying non-standard bond distances in the commands DRAW and DISPLAY. n and m are the numbers associated with individual atoms - that is the numbers generated by the SYMMETRY command, not the shell numbers. One difficulty that may be encountered, is that the order of the atoms, and therefore the significance of the numbers, may change with small changes in the coordinates of the atoms. These parameters have not been tested since changes in the symmetry code and may not function correctly. They are due to be revised.
g) Table symbols An:m, Vn:m (n and m integers). These parameters are used in restrained refinements using angles. They are analogous to Dn:m and Wn:m used in distance restraints.
h) CENTRE. The effect of this is to re-centre the cluster specified by the numeric option. If the central atom has been moved, its new coordinates will be set to 0,0,0 and the coordinates of the other atoms in the cluster moved by the same amount. The move will be remembered when using WRITE PARAMETERS BR to produce a .pdb file. The default cluster number is 1.
CHANGE is affected by the SET CONSTRAINTS option (which may be OFF, ON or ALL).
Changes to torsion angles (using TORTA1 etc.) will only be effective if CONSTRAINTS is OFF.
Changes to SURREL, SURROT, SDX, TRANSX, ACELL etc. will ignore the status of the CONSTRAINTS option (in general it should be OFF if these options are used).
If RULES have been defined, related parameters will be update whenever a parameter is changed. The use of RULES will override any changes due to use of CONSTRAINTS. In general, RULES should only be used when CONSTRAINTS is OFF.
If a parameter is the subject rather than the object of a rule, CHANGE will have no effect.
COMPARE is used to compare a new experimental spectrum with the current experiment and/or theory.
Syntax: COMPARE EV/K/KEV option
Abbreviations: COM
Notes:
The name of the additional experimental file is requested. If the wavevector is being used as the X-axis then an energy shift may be specified to permit alignment of spectra. The energy shift is added to EF and the new value is used in calculating the wave vector for both the plot and the Fourier transform.
The keyword determines wether the spectrum is tabulated in eV, wavevector or keV. E0 is used with eV and keV (e.g. if absolute energy rather than energy relative to the edge is used). E0 is ignored for K. The energy zero may be added to the offset described above. The default keyword is EV.
option = 0 Only the experiments are displayed (default).
option = 1 The theory is displayed as well as the experiments.
CONFIG is used to change hardware dependent features such as fonts and line width. These are stored in the file excurve.cfg.
Syntax: CONFIG LIST/WRITE/FONTLIST/item
Abbreviations: CON
Notes:
LIST display items that may be altered (default)
FONTLIST display list of fonts (only a few of this list may be available on a particular device)
WRITE write a new excurve.cfg file
LWIDTH determines minimum line width for hard copy
TERMFONT font for windows terminal
PRINTFONT font for hard copy
TERMTEXTSIZE determines font size for terminal
PRINTTEXTSIZE determines font size for hard copy
XOFFSET hard copy left margin
YOFFSET hard copy bottom margin
PLOTTER determines the default plotter as in GSET PLOTTER
(COLOUR/MONO etc. for Windows, HP/PS etc. for Linux)
The font and textsize options are not yet implemented in the Linux version
COPLOT displays EXAFS and Fourier Transforms together.
Syntax: COPLOT option
Notes:
COPLOT produces EXAFS and Fourier transform plots. By default, these are in frames 1 and 2 (top and bottom left) of a 4-frame page for the first spectrum, frames 3 and 4 for te second. If the GSET option FRAMES is set to SELECT then you will be prompted for the frame numbers permitting other formats. If you wish to produce plots in other combinations then use GSET FRAMES SELECT and the PLOT command to produce the individual frames.
Option may be 0 (default), 1 or 2. 1 causes the previous generation of theory to be displayed alongside the current one. 2 displays 2 previous generations. Option 2 is not available if the number of experimental points exceeds half the dimension of the array (this is displayed by LIST DIMENSION). The display is also affected by the other GSET options as with PLOT.
DEBYE Calculates disorder using Debye theory, displays heat capacities, MSD's EXAFS Debye Waller, factors etc.
Debye-Waller parameters An are calculated which can then be used for fitting.
Syntax: DEBYE
Abbreviations: DEB or DE
Notes:
Uses the current values of parameters ACELL etc, Tn, TEMP, and point group, all of which must be defined.
Further information requested is the number of atoms per unit cell (e.g. 4 for FCC, 2 for BCC), and the Debye temperature.
DISPLAY is used to display interatomic distances and angles for a cluster whose point symmetry has been defined.
Syntax: DISPLAY ATOM/CLUSTER/SHELL [atom, cluster or shell number]
Abbreviations: DI
Notes:
This command displays a table of interatomic distances and angles. The positions used are calculated from the shell variables and the point symmetry of the cluster. The point group must be specified by using the SYMMETRY command. If this has not been done, or if a structural variable has been changed since the last use of SYMMETRY then SYMMETRY will be executed automatically (without output). ( see the section on the SYMMETRY command). SYMMETRY generates labels for every atom in the structure. The first column out output is the atom number (1,2,3....n). The second is a shell number with a symmetry index (e..g 1, 1a, 1b, 1c, 2, 2a, 3 ). The order is that required by XANES calculations.
DISPLAY with no parameters, displays distances and angles with the central atom as vertex, for all clusters.
DISPLAY CLUSTER n restricts the display to cluster n.
With CLUSTER the output is restricted to distances between each atom in the asymmetric unit and all other atoms.
DISPLAY SHELL n shows all interatomic distances and angles related to shell n.
DISPLAY ATOM n shows all interatomic distances and angles involving atom number n.
With ATOM and SHELL keywords, all relevant distances are displayed. If n is omitted, all distances in the structure are displayed.
For ATOM , atoms are labelled with numbers generated by the SYMMETRY command (the 'ID' column).
For the other keywords, the atoms are labeled by shell number.
Information generated by this command may be written to a file using the PRINT GEOM command.
DRAW will display a simple plot of the clusters surrounding each central atom, or of individual units. The point symmetry of each cluster must be defined.
Syntax: DRAW keyword [cluster/unit number]
Abbreviations: DR
Keywords: CLUSTER (default) draw the structure on new axes
PATH draw the atoms of a multiple scattering path
UNIT draw an excurve 'unit' (part of the cluster defined by common 'unit numbers')
NOCLEAR overlay the structure on an existing display
POSITIVE use absolute values of the coordinates of the point of projection
Notes:
This command displays a projection of the three dimensional atomic positions. These positions should have been calculated from the shell variables by using the SYMMETRY command. If this has not been done, or if a structural variable has been changed since the last use of SYMMETRY then SYMMETRY will be executed automatically ( see command documentation for symmetry ). The information generated by SYMMETRY can be saved to a file using the PRINT ATOM command.
Atoms are labelled with their chemical symbol, provided ATOM variables as well the atom types T are defined. By default, the label also includes the shell numbers, but by selecting GSET LINE SYMBOL, the numbers generated by the SYMMETRY command are used. GSET LABEL LARGE affects the size of the labels. GSET LABEL OFF removes them. GSET AXES OFF removes the axes.
The appearance of the plot is controlled by the variables BOND, SIZE, VX, VY, VZ. These can be viewed with LIST and changed with CHANGE. A line representing the bond between two atoms is drawn if the distance between those two atoms is less than BOND. If BOND has its default value of 0, then a distance slightly greater than R1 is assumed. The atoms are represented by circles of radius SIZE*MTRn where MTRn is the muffin-tin radius for that atom type. If SIZE is given a value of 0 then a predefined minimum is assumed.
The structure is drawn as seen looking from a viewing position specified by three cartesian coordinates towards the origin of the coordinates. If the SET option VIEW is set to AUTO, the projection position is calculated automatically so as to give a minimum overlap. If VIEW is set to current, then three variables, VX, VY and VZ are updated using the last viewing position and are then used by DRAW. VX, VY and VZ may thereafter be changed manually if required. The default positions of 3.5, 2.5, 1.5 usually gives a useful view. VX, VY, and VZ must be positive and the sum of their squares must exceed 1.1. If VX+VY+VZ=0, then a minimum overlap position will be generated even though CURRENT has been selected.
In addition to bonds drawn due to the value of BOND, specific bonds may be included by setting the variables Bn:m
to a non-zero value. n and m here are atom numbers as generated by SYMMETRY not shell numbers. n must exceed m. m is zero for the central atom.
With DRAW UNIT and DRAW PATH, if automatic viewing is in use, the viewing position will be selected for the whole cluster. Although this means that it is not optimised for a particular unit, it allows additional units to be added one by one using, for example:
DR NOC 2
To add unit 2 to the picture. NOCLEAR assumes the CLUSTER, PATH or UNIT option is in accord with previous usage.
If a cluster was drawn at the last call, the new cluster will be overlaid. This makes it possible to see how a cluster has changed during refinement.
DR CL; ( ... refine the structure ... ) ; DR NOC
POSITIVE is the same as CLUSTER, but the minimum overlap projection always has VX, VY and VZ > 0.
Terminates the program.
Syntax: END
Abbreviations: none
Notes:
The program saves parameter values. Use RECOVER the next time the program is used in order to restore them, before, using the command CHANGE, which will overwrite the saved values.
EXPAND creates a parameter table containing 1 atom per shell, of c1 symmetry, from a table of higher symmetry. It can be used when it is necessary to reduce the symmetry of an idealised model of the structure. It can also be used when it is necessary to include some inter-ligand as well as intra-ligand MS paths when using the MS UNIT rather than MS ALL option.
Syntax: EXPAND
Abbreviations: EXP
Notes:
The cluster will be truncated if the total number of atoms exceeds the number of shells allowed by the program ( see LIST DIMENSION ).
EXTRACT creates a file containing either the whole sum or a partial sum of multiple scattering paths. In addition it generates a table of scattering paths, including their lengths, multiplicities and magnitudes. The latter function is likely to be used more often than the former, which was the original purpose of the command.
Syntax: EXTRACT [PAUSE/NOPAUSE/QUIET]
Abbreviations: EXT
Notes:
The command provides information on MS paths that have already been calculated (subject to filter conditions). A list of all the paths calculated will be displayed. The effect of additional filter conditions on both the number of paths, and the resultant spectrum, may then be evaluated. The path sum defined by the current filter conditions, as well as the path information, is written to a file whose name is requested.
QUIET disables terminal output, writing path information only to the disc file.
PAUSE pauses between each line of output.
NOPAUSE is the default option.
This command does not automatically update the theory. It uses the last set of paths calculated, further reduced in number by any change in filter conditions.
Isolate contributions of specific shells to the experimental EXAFS spectrum using Fourier filtering.
Syntax: FFILTER [SHORT/MEDIUM/LONG] [+/-spectrum_number]
Abbreviation: FF
Notes:
This command allows parts of the Fourier transform of the experimental data to be back-transformed. In this way it is possible to see which shells in the structure caused which parts of the EXAFS spectrum. After receiving this command the program displays the current experimental spectrum and its Fourier transform.
The user is prompted:
ENTER RMIN and ENTER RMAX
in order to determine which part of the displayed spectrum is to be back-transformed (the window). The program then prompts:
Enter MOVE, NEXT OR CONTINUE
MOVE allows the current window to be changed. NEXT allows a new window to be defined. CONTINUE goes on to the next stage. You can define up to three windows at a time. Lines indicating the extent of the windows appear on the Fourier transform plot. After a CONTINUE the back-transform of the last window is displayed.
The range in r-space is calculated automatically and depends on the point spacing Δk. This is dependent on the length of the spectrum in use, but may be altered using the option number described below.
The next prompt is:
Enter shell radius or CONTINUE
If you enter a number which indicates the actual radius of the shell that this window is trying to isolate then a graph showing the backscattering magnitude and phase is displayed. CONTINUE will go to the next stage. NOTE: the last window entered is processed first.
The next prompt is:
Enter PRINT, REPLACE, NEXT or END
PRINT will write the new spectrum to a file. You will be prompted for a filename. If the phase and backscattering magnitude have been calculated these are also written out. REPLACE will replace the current experimental spectrum with the new spectrum for all future plotting etc. NEXT will go on to the process another window, if there is one. END will exit from the FFILTER command.
SHORT, MEDIUM and LONG control how many points are used in the back-transformed spectrum ( and consequently how much R space is visible in the original spectrum ). SHORT (default) uses 200, LONG uses 500 points. If the range in r-space is initially too short, MEDIUM or LONG should be used as required.
For a multi-spectrum fit, the spectrum_number should be specified. If the spectrum number is -ve, each plot will be in full-page format, rather that 4 per page.
Note that the parameters RMIN and RMAX are not used by the FFILTER command. As no window function or phase correction is used in the Fourier transform, FTSET options referring to these are ignored. The weighting used by the Fourier transform is given by FTSET option FTWEIGHT. The GSET options are used to determine the format of the plots.
except for WEIGHTING, FFILTER uses a k**2 weighting at present. Previously a k**3 weighting has been used.
EF and EMIN are reset for the duration of the command, so the spectrum plotted will not be directly comparable with that displayed by the command PLOT.
Calculates and displays the updated fit-index.
Syntax: FIT BOND_VALENCE/EXAFS/FT/STATS/QUIET/ weighting
Abbreviations: FI
Notes:
FIT Recalculates the theory (if required) and quantifies the fit between experiment and theory. The fit is described in terms of the fit-index, chi^2 and R-factor which are defined under refinement, in the theory section, chapter 1. The discrepancies between ideal and actual parameter values are also displayed if restrained refinement is in use. The relative weighting of theory, distance restraints, and angle restraints are given by WEX, WD and WA. If multiple spectra are being used, their relative contributions to the fit-index and R-factor are determined by WEX1, WEX2 etc. (multiplied by WEX). These weighting are displayed by the command.
If the keyword FT is specified, then the Fourier transforms are also recalculated. If weight is specified then it is used as the k-weighting in the calculations rather than the value currently specified by SET WEIGHTING, which is the default.
Keyword STATS gives statistics referring to specific shells.
The calculation of chi2 relies on the number of independent points in the spectrum which is normally calculated automatically by the program using 1/2πΔkΔr, but may alternatively be specified using the parameter NIND. It also uses Np, the number of refined parameters. This must be set by the user, using the parameter NPARS, to the total number of parameters refined at any time during the analysis.
The fit-index includes a term due to the soft constraints used by he program, especially that due to A parameter values which fall beyond DWMIN or DWMAX. The term is given by:
Σi constant x ((DWMIN-Ai)2 + (Ai-DWMAX)2 ) for all terms A < DWMIN or A > DWMAX
There are no such terms in the R-factor or chi2.
With keyword BOND\-VALENCE the bond valence sum for each cluster is displayed, using the parameter BOND to define the number of bonded atoms. A default value of atomic valence is assumed, which may be overwritten using ION parameters for each atom type. Note however that these should be reset before further potential calculations are performed.
Controls options concerned with Fourier transforms.
Syntax: FTSET keyword value
FTSET LIST
FTSET
Abbreviations: FT
Notes:
The options which control the way Fourier transforms are calculated are set by this command.
The first format changes the setting of keyword to option and returns to the normal EXCURVE command level.
The second lists the current values of all the options and stays within the FTSET environment.
The third format enters the FTSET environment.
For historical reasons these parameters also control some features of radial distribution calculations.
The keywords are:
This determines how the phase of the Fourier transform is calculated.
First shell : the phase is calculated from the first shell backscattering factor.
Second : the phase is calculated from the second shell
None : no phase correction
Controls the weighting applied to the spectrum before it is transformed.
None : no weighting
K : k-weighting
K2 : k2 weighting
K3 : k3 weighting
K/fpim : weighted by k/back-scattering magnitude
Controls the type of window.
Hanning
Gaussian
Kaiser/Bessel
Blackmann/Harris
The parameter WP determines the characteristics of the window, in a manner depending on the choice of window function.
One : no multiplication of transform
R-squared : transform is multiplied by R**2
Controls the position of the centre of the window which is Fourier
transformed. Note that the window type is controlled using the
parameters WIND and WP. The default window is a Gaussian with
width 0.1
Auto : centre of window is midpoint of spectrum
K : centre uses the parameter CENT (in wave numbers)
Energy : centre uses the parameter CENT (in energy)
Controls the type of radial distribution function calculated and displayed by PLOT RDF.
Atomic : atomic radial distribution function
Electron : electron radial distribution function (requires that ATOM parameters are defined).
Atomic/r2 : 1/R2 times atomic rdf.
Determines whether the imaginary part of the Fourier transform is displayed as well as its modulus.
Modulus : only the modulus of the complex FT is displayed.
Sine+modulus : both the modulus and the imaginary part are displayed, revealing errors in the phase as well as the magnitude of the EXAFS function.
QUIT
Leave FTRANS command.
LIST
Display option table.
RESET
sets all options to 1
The values of WP are set using the CHANGE command. The values
allowed depend on the setting of FTWIND and are:
1 uses a Blackmann-Harris window, there are two sets of pre-defined coefficients defining a narrow or wide window. If WP = 1.0 the narrow window is used, if WP = 1.5 the wide window is used.
of window )
2 (default) uses a Gaussian window, the exponential power is -WP * ( square of K - K of centre of window )
3 uses a Hanning window, weight is ( cos( (K-Centre) * PI / (Kmax-Kmin)) ) ** WP
4 uses a Kaiser-Bessel window (no parameters).
Generates additional clusters from an atom in an existing cluster.
Syntax: GENERATE {LINKED/UNLINKED} new_cluster_number
The program will prompt for an atom in an existing structure, to be used as the centre of the new one.
Abbreviations: GE
Notes:
Given an existing cluster, the program generates a new cluster centred upon one of the atoms in the first cluster. The clusters may be linked or unlinked. With a linked cluster, any changes in one of the clusters results in changes in the others. This case is used when two central atoms (of either similar or different type) are within close proximity (say, 5 to 6 Angstroms). Unlinked cluster are used when the clusters are far apart, or else when they are near, but the inter-cluster distance itself is the variable of interest. GENERATE simplifies the task of setting up variables for a new cluster, but does not do all the work. Normally the atom types (Tn) must be changed for the new central atom (in general a scattering atom in the original cluster) and for the old central atom (now a scattering atom). The occupation numbers N may also need to be changed. Indeed, the command is not reliable except in the case that all the occupation numbers are 0 (point group C1). A minimum requirement is that the point group of the central atoms is the same. Even then, it will be necessary to adjust the occupation numbers if they are not equal to one. The only occasion when the occupation numbers will be correct with higher point symmetries is when the central atom of the first cluster is also the central atom of the second. This may arise when the command is used as a starting point for generating unlinked clusters about different atoms in a close-packed lattice. Note however, that this may often be done by using mixed sites for both absorbing and scattering atoms.
The default value of LINK variables is 0, and it is not possible to link a shell to shell 0 (the central atom of the first cluster). Shell 0 may however itself be changed, and the other clusters will be updated accordingly. This restriction may be removed in later versions.
If linked clusters are used, the links may be inspected using LIST LINKS. In recent versions links may also me inspected or CHANGEd individually, by means of the variables LINKn which specify the shell which will be updated when shell n is changed. Note that it is normal to change the first cluster which will then update the others, but in later versions any cyclical change may be performed ( i.e., in a 3 cluster structure, changing cluster 2 will update cluster 3 which will in turn update
cluster 1).
Note that link information is lost when NS is reduced, so it may not be reduced temporarily if links are in use. This was done in recent versions to avoid unwanted changes due to 'hidden' links.
Controls the options which affect the appearance of graphical output.
Syntax: GSET keyword option
GSET LIST
GSET
Abbreviations: G, GS
Notes:
This command selects options which control the appearance of graphical output. If GSET alone is entered on the command line the command issues prompts for further input. At this stage you can enter a keyword and a new option, LIST (to list current options), RESET to change all options to the first on the list or QUIT to leave the GSET command. GSET LIST displays the current values and remains within the command. GSET keyword option changes the option associated with keyword and returns immediately to the EXCURVE command prompt. The keywords and options are as follows (all keywords and options may be abbreviated so long as they uniquely define the entry required.
A number corresponding to the list position may be also be entered ( e.g. G LA 1 will select the first option, which is ON).
ON
OFF
BOTTOM_ONLY
graphs have axes (default)
graphs have no axes
x-axes are only displayed on the lower graph of each column.
(default)
hard copy only
FIXED
SELECT
graphics output uses the whole screen, for single frame plots, and frames in sequential order for multi-frame plots (coplot, plot spectra etc.) (default)
allows the position of frames to be selected by the user. You will be prompted for a frame number for each plot. Frame 0 is the whole screen, 1,2,3 and 4 are for quarter screen, 5-19 for 5x3 frames.
ON
OFF
LARGE
axis and curve-identification labels are visible (default)
labels do not appear
large labels are used in DRAW (otherwise this is the same as ON)
NORMAL
THICK
single thickness lines are used in plots (default)
data points are marked with symbols for the first curve of any plot. This also controls whether shell or atom indices are used in DRAW.
double thickness lines are used
ON
OFF
hard copy plots use the same option values as terminal plots (default)
hard copy plots have all possible annotation irrespective of plot option values
MONO
COLOUR.
selects any HP device (pen plotter, Deskjet 1200/1600, or Laser HP mode)
selects a POST-SCRIPT laser printer
selects a disc file, to contain HP output for subsequent submission to a printer, or for input to other software.
Selects a POST-SCRIPT file.
With all FILE options, the file contains a single plot with a unique filename.
selects a Windows monochrome printer.
selects a Windows colour printer - generates grey-scaling on a b/w device.
ON
OFF
MAX
"Enter Command" prompt appears after plotting
"Enter Command" prompt does not appear until carriage return typed
No suppression of messages during command files.
NONE
K, K2, K3 etc.
The keyword PWEIGHT is used to alter the k-weighting of EXAFS plots independently of the SET WEIGHTING option. Any subsequent use of SET WEIGHTING will modify PWEIGHT.
No weighting is used
A kn weighting is used
EMIN/MAX
SELECT
the x axis limits are determined by EMIN/EMAX, KMIN/KMAX or the spectrum length (default)
prompts are issued for axis limits every time a spectrum is plotted
uses EMIN/EMAX etc. for the x-axis, YMIN/YMAX for the y-axis
ALL
EXPERIMENT
experiment and theory are plotted (default)
experiment only is plotted.
theory only is plotted.
difference between theory and experiment plotted
LONG
MEDIUM
SHORT
NONE
parameters and filenames displayed on graphs
only parameters are displayed
only shell parameters displayed
no parameter table is displayed
Single screen device with alpha and T4010 modes (Selanar, PCs running TELNET).
Two logical screen device with alpha and T4010 modes (Pericom, PCs running KERMIT).
Single screen device with T4010 mode only.
XTERM, normally for a terminal without full X capabilities.
Colour X-window display, for terminal with full X capabilities. Requires that the
UNIX environment variable DISPLAY is set to the current client window, e.g. by setenv DISPLAY mscsv1.dl.ac.uk:0.
AUTO
CURRENT
DRAW uses a minimum overlap projection (default)
VX, VY and VZ are set to the current projection position. Thesevariables are used for subsequent use of DRAW.
K
ENERGY
EXAFS backscattering and windows are plotted against wave vector (default)
EXAFS, backscattering and windows are plotted against energy
Syntax: HELP command/topic
Abbreviations: H
Notes:
The command does not currently exist
A command list can be obtained at the ENTER COMMAND: prompt using ?.
Interactive help on individual commands can be obtained using command ?.
Further information can be obtained from the WWW version of the manual.
This command idealises the geometry of amino acid residues in protein structures using data taken from the constraints file used by the program PROLSQ.
Syntax: IDEALISE [ROTATE/NOROTATE] [unit_number]
Abbreviation: ID
Notes:
If a unit number is specified, only that unit is idealised. Otherwise, all units are idealised. If ROTATE is specified, the unit is rotated to a standard orientation in the coordinate frame. NOROTATE is the default keyword.
Idealisation assumes that all four main-chain atoms (N, O, C, CA) remain unchanged and that the torsion angles TORA and TORB remain unchanged. These two assumptions mean that the cumulative effect of small changes in interatomic distances, angles and torsion angles may result in very large movements of the atoms coordinated to the metal atom. The metal-ligand distance, the principal angle, and the torsion angle TORC can be optimised, without changing other distances or angles, using either the command spin or map. In each case TORA and TORB are varied. It is also possible to perform a general rotation about any bond, although unlike changing a torsion angle this will also result in changes to the main-chain atoms. A rotation can be achieved using a ROT parameter, after setting VECA and VECB to the correct shells numbers.
Syntax: INFO keyword
Abbreviation: INF
Notes:
Displays data used by the program as tables.
Keywords available are:
ENERGIES POINT_GROUPS
K-EDGES K-WIDTHS
L1-EDGES L3-EDGES
L3-WIDTHS M5-EDGES
No defaults
INQUIRE displays information about current program values.
Syntax: INQUIRE [NPT]
Abbreviation: IN
Keyword NPT limits the display to the value of NPT the number of points on which the theory is currently calculated.
LIST displays parameters values, parameter tables, program options, etc..
Syntax: LIST keyword [logfile]
Abbreviation: L
Notes:
This command displays values of parameters, the status of set, gset, and ftset options and other such things. The information is displayed on the terminal unless logfile is specified, when it goes to exout98.lis.
The options available, which determine the information displayed are:
NSHELL (default) information about the first NS shells.
ALL information about all shells, whether currently in use or not.
parameter_name The value of the parameter. Indexed parameters for several shells, atom types, etc. can be listed at once using the [] notation described under the CHANGE command. e.g.
L MTR[1-4]
will list the Muffin-tin radii for the first four atom-types.
Doubly-indexed parameters, such as the distance restraint parameters (d3:0 etc.) may be included.
A:,B:,D:,W:,V: give a complete table of the relevant doubly-indexed parameters A1:0 etc. as follows:
A: actual and ideal angles as used in restrained refinement.
D: actual and ideal distances as used in restrained refinement.
W: distance weightings corresponding to each D: value
V: angle weightings corresponding to each A: value.
BACK parameters associated with the atomic-background contribution to the total absorption spectrum. Only valid if ATOMIC_ABSORPTION is selected using the SET command.
CART cartesian coordinates of atoms
CLUSMAP displays the relationship of defined clusters to experiments that have been read-in.
DIMENSION program dimension limits.
FILES names of all files currently in use.
FRAC fractional coordinates of atoms in terms of the current 'unit cell' (usually not the true crystallographic unit cell - see notes on ACELL etc.).
FTOPTS current settings of all the options controlled by the FTSET command.
GSETOPTS current settings of all the options controlled by the GSET command.
MSPARS multiple scattering parameters.
RADIAL spherical polar coordinates of atoms relative to central atom, rather than origin of coordinate system. The too systems will differ only if the central atom position is not at (0,0,0). If this is the case the option will normally therefore also display the absolute coordinates R, TH and PHI. This keyword has been changed in recent versions from RELATIVE to avoid conflicts with the set option RELCOR etc. RADIAL may now be more simply abbreviated to RAD or R.
SETOPTS current values of all the options controlled by the SET command. Each individual option may also be listed (e.g. L WAVEVEC).
SPARS polarisation dependent parameters.
UPARS unit and angle parameters.
Calculates and displays the variation in fit-index as a function of two parameters.
Syntax: MAP keyword [number_of_maps/map_number]
Abbreviation: MA
Notes:
The MAP command calculates a set of contours showing the variation of the fit-index with two parameters. The minimum fit-index values of the two parameters are displayed on the plot.
The keywords are:
CALC The two parameters are requested in a similar way to that used by the REFINE command. The calculation is performed interactively. The option to perform the calculation as a batch job is not currently available.
READ Reads in the output from a batch map calculation. Interactive calculation always writes to a file called MAP which is overwritten each time a map is calculated. MAP may be renamed using ^mv if required. If a map has been calculated interactively, READ may not be required if that is the map required.
LIST Displays the values of fit-index and parameters in a table.
PLOT Draws a contour map of the fit-index. A large X marks the minimum.
TRACE Extracts information from background-processed refinement log files, and marks the path taken by the refinement on an existing contour map.
The default keyword is CALC.
Several maps may be processed at once: the numeric option indicates either how many maps are to be calculated or which map of a multiple set is to be read in or displayed. The maximum number is 9 and the default is 1.
Any parameters which are controlled by the CHANGE command can be used as one of the two map parameters. The resolution of the map is controlled by the SET option MAPSIZE. If MAPSIZE is SMALL, additional values will be interpolated, resulting in a poorer result. LARGE is recommended except when large number of multiple scattering paths are required.
example:
to produce a table of the fit-index for values of R1 between 1.0 and 1.5 and R3 between 1.5 and 2.0 follow these steps: (user input in capitals):
R1;1.0;1.5
R3;1.5;2.0
Continue
then when the calculation is finished:
MAP READ
MAP
MAP LIST
displays values of the fit-index
MAP PLOT
draws a contour map
PLANE defines the orientation of a plane defined by three selected atoms (in terms of a vector normal to the plane), the angle between the plane and a reference axis, and a torsion angle 0-1-2-3, where 0 is the central atom.
Syntax: PLANE
Abbreviation: PLA
Notes: There are no keywords or other options, the three atoms defining the plane, and the x, y and z coordinates of the reference vector are requested by the command.
Used to produce graphical displays of data.
Syntax: PLOT keyword option
Abbreviations: PL, P
Notes:
The format of the graphs is controlled mostly by the parameters which can be set using the GSET command. The keywords control the content of the plots. These keywords are:
ATOMIC Plot the atomic contribution to thew whole spectrum (when ATOMIC_ABS is in use,see SET).
BSF The magnitude and phase of backscattering factor are plotted. The shell specified by the PS parameter of the FTSET command is used to calculate the backscattering factor.
BSMAG As for BSF, but plots the backscattering magnitude only.
BSPHASE As above, with backscattering phase only.
CAP Plots the central atom phase.
EXAFS The current EXAFS experiment and theory are plotted. Option = 1 displays the previous theory as well, option = 2 the previous two theory spectra. The default PLOT option is PLOT EXAFS 0.
FTRANS The Fourier transforms of the current experiment and theory are plotted. The calculation of the Fourier transform depends on the settings of the options of the FTSET command. Option = 1 and option = 2 have the same effect as with EXAFS.
PATHS Plots individual scattering paths and their FTs after using EXACT MS theory. Unlike most other PLOT commands it does not update the theory. By default, 15 frames per page are displayed. P PATHS 1 displays 4 frames per page. GSET FRAMES SELECT may be used for other options.
PFT Plots EXAFS multiplied by window and FT weighting term. This shows what is actually being Fourier transformed.
PHASES Plots the first 7 of a set of phaseshifts (the L=0 to 6 terms). If an option number is specified, the phaseshift corresponding to that atom number are displayed on one page. If no atom number is supplied, all the phaseshifts are plotted 4 frames to a page. The page layout may of course be modified using GSET FRAMES SELECT.
RDF Plots either an atomic or electronic radial distribution function depending on the value of the FTSET parameter TR. The number of points between RMIN and RMAX used to calculate is determined by the FTSET parameter ND. If option = 1 the contributions of individual shells are plotted.
REF The real and imaginary parts of the photoelectron self-energy are displayed.
REFP As above, but on the experimental, not the phaseshift grid.
SHELLS The theoretical contribution of each shell is plotted separately. After each shell is plotted you must enter a carriage return in order to display the next one.
SIGMA Plots the experimental weighting function (if any), which can be used in place of kn in EXAFS refinement.
TPHASE Plots total phase = 2 * scattering phase + central atom phase.
UNIT Draws a projection of the currently defined units onto the x-y plane, displays a table of unit parameters, and allows some editing of the unit coordinates.
WFX The weighted and windowed EXAFS function used in Fourier transforms.
WINDOW Plots the window being used for Fourier transforms.
The command COPLOT is similar, but displays both the EXAFS and the Fourier transform on one page.
Generates files containing various type of data.
Syntax: PRINT keyword [column_combination]
Abbreviation: PR
Notes:
Column_combination is only relevant for the SPECTRUM option. The keywords are associated with different types as data as follows:
SPECTRUM (The default keyword) - any of a variety of spectra are printed in columns according to the value of column_combination. The first digit of column_combination specifies the contents of column one of the file. 1 gives energy, 2 gives wave number and 3 gives distance. The remaining digits ( up to three are allowed ) specify the contents of columns 2 to 4 as follows:
0 omit
1 exafs experiment
2 exafs theory
3 fourier transform experiment
4 fourier transform theory
5 sine transform experiment
6 sine transform theory
9 phase of backscattering factor
If column_combination is initially omitted then you will be prompted for the contents of
each column. In this mode an additional option is available:
10 difference between theory and experiment
The SPECTRUM option updates the fit-index before data is printed
ATOMIC_COORDS prints the coordinates of all atoms, as generated by SYMMETRY
BACK produces the 'atomic background' and 'XAFS' contribution derived from the total absorption spectrum
GEOMETRY prints out all current interatomic distances and angles
PARAMETERS the current theory parameters are printed. Options for producing parameters in different format are discussed below.
PHASESHIFTS prints any phaseshifts. You will be prompted for filenamesfor the phaseshifts of each atom.
POTENTIAL muffin-tin potentials.
REF energy dependent self-energy function.
PATHS individual multiple scattering terms, binary format, for use with READ PATHS.
TMATRIX prints the current central atom phase and l=0 to 2 diagonal elements of the central atomT-matrix.
VARIABLES prints out the current values of all the internal variables
The command requests a filename, suggesting an automatically generated name, which includes a number that is incremented each time a file is generated. A prompt will be issued before an existing file is overwritten.
Most of the files contain a header containing the fit-index etc. If this is the case, the theory is updated before the file is generated.
The options associated with PRINT PARAMS allow parameter files to be be produced in several format. By default, the parameters are in EXCURVE format. Alternatives are CERIUS EXPLORER (.car) format, produced using:
PRINT PAR CAR
Files in BROOKHAVEN (.pdb) format are produced using:
PRINT PAR BR
More details of these formats are given in the section on Brookhaven files in part 2, and in the command documentation for READ.
Read data from files specified by the user.
Syntax: READ keyword [spectrum_number]
Abbreviation: R
The keywords available are:
ALL reads EXPERIMENT, PARAMETERS and PHASESHIFTS, i.e. all the files needed to perform an EXAFS calculation. See a description of each of the keywords below.
EXPERIMENT reads a file containing experimental data. This option issues prompts for the following information:
filename the name of the file
point frequency 1 to read every line of the file, 2 every second line, and so on
column combination e.g. 32 means read energy from column 3, EXAFS from column 2
edge details of the edge - element + initial state with the format:
CU K
I L2 etc.
The values entered will be used in theory, potential, and phaseshift calculations.
sequence-number in polarisation set
is used only for polarisation dependent spectra (otherwise 0). If experiments 4, 5 and 6 were 3 polarisation dependent spectra, differing only in their orientation with respect to the beam. The sequence-numbers would be 1 for experiment 4, 2 for 5 etc. If three polarisation dependent spectra each referred to a different sample, or to a different excited atom, the response would be 1 in each case.
number of clusters for this experiment
is used to relate the clusters in the structural model of the phase or phases being analyzed to experimental spectra. The sum of all the clusters for all the experimental spectra read must equal the number of clusters represented in the parameter table. If a spectrum is to share the clusters of the previous one, the response should be 0. This will be the case if a). the central atom is specified in the parameter table as a mixed site, and successive spectra have central atoms that are components of the mixed site or b). if the spectra are part of a polarisation set. Thus if the reply to the previous prompt was 2 or more, the replay to this must be 0. If the structural model is changed during the refinement, it will be necessary to re-read the spectra.
The numeric option is the number of the experiment (default 1).
Thus to read a second experiment:
R EX 2
If several experiments have been read, but only the first is currently required, the parameter NSPEC may be reduced to 1, and reset later to restore all the spectra.
If several experimental files are read (for example, spectra measured using different polarisation vectors), it is very important that they share an identical energy grid.
ME matrix elements for XANES calculations, weighting of multiple final states or whole-spectrum fitting are read.
MS read a file of data produced by a multiple scattering batch calculation. When a file is read the XADD flag is automatically set ON. See the XADD help for more details of this flag.
PARAMETERS reads a file containing parameters. Parameter files can produced using PRINT PARAMS or generated from scratch using the same format. If shell parameters are to be added to an existing list, they should be read using R PAR ADD (make sure that the total number of shells does not exceed the dimension limit given at the start of the program).
In addition to the usual EXCURVE format, parameters may be read in BROOKHAVEN protein database format (.pdb files) - R PAR BR. Or in CERIUS EXPLORER format (.car files) - R PAR CAR. If a .pdb file is read, ligands which are within MAXRAD Å of the specified central atom will be read. MAXRAD may be changed from its default value of 5 if required. When a .car file is read, the central atom is requested. An atom label (e.g.FE2) within the .car file is requested. If an exafs experimental file has been read in, the atom type should match the edge specified.
R PAR ADD, will instead of replacing the coordinates, add extra shells, generating a new cluster for the additional shells.
In order to read several cluster from a .pdb or .car file, read one, PRINT it, read another, then read the first using R PAR ADD. Alternatively (for nearby clusters), read a cluster using a high value of MAXRAD, then use GENERATE to produce multiple clusters.
PHASESHIFTS reads phaseshift files. The program prompts for the number of files to read and then for Central Atom File, First Atom File and so on. All files must contain the real and imaginary parts of both the phaseshifts and the photoelectron self-energy. Older files must be converted.
If several file-names have a common prefix then only the tail need be entered, starting with a '.'. This actually applies to all file-names read by the program, although only with phase-shifts is it likely that many will share a prefix.
POTENTIAL reads a file containing a potential tabulated on a logarithmic energy grid (Rydbergs). The program will first prompt for an atom number (answering, for example, 1 means that atoms of ATOM1 type will use this potential), and then for an input file. The format for potential files is described under PRINT. After reading the file, prompts for values of V0 and MTR (muffin tin radius ) are given. In both cases the default values are those read from the file.
SIGMA read a file specifying the values of SIGMA (see the refinement section of the manual).
SPECTRA read the spectra of individual MS paths as produced by the command PRINT SPECTRA or by MS refinements run in the background
Recovers the latest parameter values, from the last time the program was used in the current directory.
Syntax: RECOVER
Abbreviation: REC
Notes:
The parameters must be recovered before CHANGE or REFINE is used, before a parameter file is read, or parameters are changed in any other way. Either of these procedures will overwrite the copy of the parameters on disc.
A least squares refinement command, with some other functions:
(a) calculates spectra at equally incremented values of one or more parameter
(b) refines parameters by least squares
Syntax: REFINE keyword step_parameter
Abbreviation: REF
The keywords are divided into two groups according to whether they are associated with
functions (a) or (b).
For (a) step_parameter is used as the number of steps.
For (b) it relates to the step size - high number, small steps in the parameters being refined, small number, large steps. The default is 100.
The type (a) keywords are:
GRID Calculated a grid, in any number of dimensions, of the fit-index associated with equally spaced values of a number of parameters. All information is requested through prompts (the step parameter may optionally be specified on the command line)
TABLE Generates a table for the reverse Monte-Carlo program of McGreevy et al.
The type (b) options are:
prompts the user to specify the parameters to be refined. This is
the default keyword. The example below shows how a table of parameters
is constructed.
uses the parameter list from the last refinement. The STEP size may be changed, and the parameter list may be edited.
perform only sufficient steps (number of variables+1) to calculate statistical errors.
A
All the A parameters (values which are initially identical will be grouped together)
MTR
Muffin-tin radii of atoms used in potential calculations
N
N parameters for each shell
PHI
PHI parameters for each shell
R
R parameters for each shell
TH
TH parameters for each shell
X
X parameters for each shell
Y
Y parameters for each shell
Z
Z parameters for each shell
EF
EF, or EFn values for each experiment, if these are non-zero
POST
Post-edge polynomial correction to the atomic contribution (with ATABS ON). Up to NPOST terms are included
RESET
Resets internal refinement variables
Notes
Keywords may be combined using + signs. E.g. REF N+A+R
For some keywords, parameters that are initially zero will be ignored. Parameters that are the subject of RULES may not be refined and will automatically be excluded from lists generated by the program.
The weighting used by the refinement is that same as that used by FIT, and can be altered using the SET option WEIGHTING.
The command uses numerical estimates of the derivatives whose values can depend on the step parameter. Convergence is often poor and refinements using a range of values of the step parameter are often required. A useful method of improving convergence is to refine different sub-sets of the variables required separately.
As with the command CHANGE, a list of parameters may be used, e.g. A[1-3,5].
An example of the LIST form is given below:
REFINE 150 (no keyword, so LIST assumed, step parameter set to 150)
Enter parameter name ot "=" to skip
EF
Enter parameter name ot "=" to skip
R1
Enter parameter name ot "=" to skip
=
Least squares refinement using k**3 weighting
Initial parameters
1 EF 17.240 0.00272 17.51203
2 R1 1.500 0.00002 1.50159
Enter : CONTINUE to refine interactively
a number to edit, add to, or delete from the list
3
we forgot R2, 3 means that we want to add parameter
number 3 to the list.
Enter parameter name ot "=" to skip
R2
Least squares refinement using k**3 weighting
Initial parameters
1 R1 1.500 0.00002 1.50159
2 E0 17.240 0.00272 17.51203
3 R2 2.922 0.00003 2.92553
Enter: CONTINUE to refine interactively
number to edit, add to or delete from the list
C
(for continue)
R1 = 1.50000 E0 = 17.23991 R2 = 2.92244
Call 1 F= 351.5847 R= 177.353 F(min)= 351.585 R(ex)= 177.353 R(dis)= 0.000
Select Next,Exit,Complete,Stats or Plot
N 33 (or just 33)
Perform 33 steps. The values of R1, R2 and EF will be refined until the program reaches an apparent minimum in the fit-index, or until the specified number of steps has been carried out. Refinement uses Harwell routine VA05A, using numerical estimates of the derivatives.
The other options at this point are:
Exit: leave the refine command immediately
Complete: carry on without further prompts, except for the name of the statistics file
Stats: terminate the command after calculating sufficient terms for statistical errors.
Plot: toggle simultaneous plotting on/off
In this case the refinement should end with:
R1 = 1.95281 R2 = 2.87818 E0 = 21.12186
Call 50 F= 19.9091 R= 44.305 F(min)= 19.909 R(ex)= 44.305 R(dis)= 0.000
Minimum Predicted, Accuracy = 0.2817E-02 Cond = 2 At Call 50
Requests a filename for the correlation matrix and statistical errors, calculated from the last estimate of the partial derivatives before the predicted minimum.
If only terminal and logfile output is required, enter '='.
Rotate a cluster or a unit about an axis.
Syntax: ROTATE UNIT/CLUSTER/AOMS_IN_UNIT [cluster or unit number]
Abbreviation: RO
Notes:
Rotate allows a cluster, a unit or atoms within a unit to be rotated. The latter option corresponds to one of the torsion angles within the unit being changed.
Rotation of a cluster requires the direction of an axis which passes through the origin of the coordinate system to be defined. For a unit, a point through which the axis passes must also be defined. In all cases a rotation angle must be provided.
The cluster or unit number is entered as part of te command. If no cluster/unit number is supplied then 1 is assumed.
Prompts are issued for all other information. There are several means by which the axis and an intersection point may be defined (1 to 7 below).
1. The axis is defined by means of the three angles (degrees) that it makes with the orthogonal axes of the coordinate system, i.e. the angles the axis makes with each of the x- y- and z- axes. The default corresponds to a cube diagonal (useful for orienting a cubic point group so the principal axis is a triad). The axis stored by the last use of PLANE may also be used. In this case, the effect is that the cluster is rotated so that the pre-defined plane is normal to the principal axis which was also selected in PLANE.
For UNIT, the intersection point is requested as three coordinates (x,y,z). Note that as always, parameters or expressions may be used, e.g. X1;Y1;Z1 to enter the coordinates of the first shell atom.
2. is quicker but less flexible. The axis direction is given as a miller index of the coordinate system (e.g. 111 corresponds to the default values of 1). The intersection point must be an atom, specified by its shell number. It is not yet possible to specify an index in terms of the crystallographic cell defined by ACELL etc.
The other options require that the axis is defined by means of two atom positions. For a cluster, one of the positions must be the central atom of the cluster, which means that the last three options are not suitable for this case.
3. requests the two atom (shell numbers).
4. uses VECA and VECB, which must be CHANGEd to suitable shell numbers, as when refining a rotation.
5. uses the central two atoms of thoses defining the torsion angle TORA. This option is most appropriate with the ATOMS_IN_UNIT option.
6. Uses the atoms defining TORB.
7. Uses the atoms defining TORC.
In each case a rotation about the axis is then requested. Note that the sign of this angle is important, and depends on factors of 180 degrees in the definition of the axis.
The effect of the rotation may be removed using UNDO.
Generate an algebraic rule to link parameters. This provides a means of constraining parameters that are not possible with the SET option CONSTRAINTS.
Syntax: RULE {parameter/DELETE} {DELETE}
Abbreviation: RU
Notes:
The parameter specified is defined in terms of a simple expression, which is requested. The rule must link parameters by the four operators +,-,* and /. Brackets and functions are not permitted. e.g.
RULE R2
R1*2
Will ensure that R2 is always twice the value of R1. R2 will be updated whenever R1 is changed either explicitly using the CHANGE command, or implicitly using REFINE. It will not be updated if new values are read from a parameter file however.
Constants may be stored in the memory parameters M1, M2, etc.
If RULE is entered without parameters, the currently defined rules are displayed.
There are two ways to delete rules
RULE DELETE will delete all rules
RULE parameter DELETE will delete the rule for parameter only
Save parameters and fitting statistics for use in tables and statistical tests.
Syntax: SAVE
SAVE RESET
Abbreviation: SA
Notes:
Saves the values of all current parameters internally. These values can later be displayed in a table. See the TABLE command for an example table construction. Saved sets of parameters are also used by the STATS command.
RESET removes all sets that have been previously saved.
This command is used to set values of the following variables which control the method of calculation or appearance of output or other such things. Compare with the commands FTSET and GSET which are used to set the values of variables controlling fourier transforms and graphs respectively, and the command CHANGE which is used to set the values of numerical variables.
Syntax: SET keyword option
SET LIST
SET RESET
Abbreviation: S
Notes:
LIST generates a list of keywords and options, including the current settings.
RESET sets all the options to the first in the table (not very helpful, as these are often not the defaults).
The allowed keywords are as follows:
OFF the atomic contribution is not included in the theory
ON the theory includes the atomic contribution - dipole matrix elements must have been calculated or read, and the experimental fermi energy must have been aligned exactly to provide a reference zero using the E0 variable, before the experimental spectrum is read.
OFF no automatic atomic correction
ON post-edge polynomial corrections to the atomic background are adjusted
OFF no domain averaging for surface calculations.
ON the theory for the first two spectra are averaged, in order to take into account the random orientation of surface domains.
NONE no iterative MTR adjustment
V adjust MTRs to give common V0
CHARGE determines the Norman radius (charge on the Wigner-Seitz sphere is always equal to ION)
If NONE is selected, the parameters V0, RHO0 and FE0 will be recalculated each time the potentials are calculated. Otherwise, the phaseshifts will be calculated to common value of one of these parameters.
ON movement of one of the atoms in a unit will cause the
others to move as if the whole unit had been moved as a rigid body.
OFF atoms in structural units can move independently.
ALL causes movement of the whole cluster if any atoms coordinate is moved.
ON produces extra information about the internal state of the program which is probably not useful to ordinary users
APPROX uses Debye-Waller factors in 4 cumulants for disorder
EXACT uses numerical integration assuming a 4-term cumulant expansion of the pair-distribution function. Applies to single-scattering only.
HEDIN-LUNDQVIST includes the HEDIN-LUNDQVIST excited state exchange term
XALPHA use a constant complex self-energy
ALL - both the L-1 and L+1 final states are calculated. Cross terms are not yet included. As the cross terms are usually more important than the minor channel, the use of this option is not recommended. The atomic contribution for both states must have been calculated for this option to work.
L-1 - only the minor channel is calculated.
L+1 (default) - only the dominant L+1 channel is calculated. This option should be used for data analysis.
XALPHA use X-α exchange and correlation energy for ground state potentials, using current values of ALFz
VON_BARTH. Use von Barth and Hedin exchange. See theory section EXCHANGE for more information.
the two options SMALL or LARGE control the resolution of output produced by the MAP command.
OFF (default) no multiple scattering is calculated
ON multiple scattering contributions are calculated for orders of paths between OMIN
and OMAX for atoms in units only.
ALL calculates all MS paths not just those associated with units - requires that the point symmetry of each cluster is defined.
OFF (default) the calculations are performed assuming a polycrystalline or amorphous sample.
ON performs polarisation dependent single crystal calculations or surface, using a small atom polarisation factor. In this mode the beam direction and polarisation parameters BEW, BETH and BEPHI must be set. Domain averaging will be used if the first two experiments have the same name.
EXACT performs full curved wave polarisation dependent calculations.
OFF excludes any contribution to inelastic scattering below the plasmon threshold (plasmon pole approximation)
ON calculates low-energy inelastic contributions according to the QUINN model
OFF (default) no correction for 3D motion
ON 3D motion correction (experimental at present)
OFF no scalar relativistic corrections to the phaseshifts (atomic charge densities are always calculated relativistically)
ON include scalar relativistic terms in the phaseshifts
ON Weight spectra by 1/sigma, where sigma is determined experimentally.
OFF Weight spectra by kn.
ON / OFF selects two algorithms during program development - not intended for users
SMALL_ATOM theory to be used in EXAFS multiple scattering calculations. Includes the Rehr and Albers option - changing NUMAX to 0 selects the small-atom approximation, 1 gives the Rehr and Albers approximation using a reduced matrix size, 3 selects for full-matrix Rehr and Albers theory.
AVERAGE use average of energy dependent self-energy of central and scattering atom for each shell.
CENTRAL use central atom self-energy throughout.
SCATTER use appropriate scattering atom self-energy.
Selects the choice of weighting function for EXAFS fits (using FIT), least squares refinement using REFINE and output generated by the PRINT command. Options available are:
NONE
K
K2
K3
K4
Setting the WEIGHTING will also modify the GSET option PWEIGHT, and the FTSET
option FTWEIGHT. The latter two may be changed individually after WEIGHTING has been altered.
is a title which will appear on graphical output. Note that setting TITLE is slightly
different from setting the other variables in that you must wait for a prompt rather than being able to enter the title on the command line.
: enters the command and prompts for further input
SET keyword option-name : example SET DISORDER DTEMP
SET keyword option-number : example SET DISORDER 3
SET LIST : lists the current options and then prompts for further input
SET RESET : sets all the variables to their FIRST value ( NOT their default )
Syntax: SITE name
Abbreviation: SIT
Notes:
If name is specified, a mixed site is defined, with a pseudo-element symbol name. This should be 1 or 2 characters in length, and should not conflict with any actual element. The program will request the details of up to three components, each characterised by an element and a proportion. Data should be entered in the form:
Enter component A and proportion:
CU .33
Enter component B and proportion:
ZN .67
Enter component C and proportion:
=
Either 1, 2 or 3 components may be entered. Use = to avoid unnecessary entries. If the total is less than 1, the site is assumed to contain vacancies. If the total is more than 1 you are a complete pratt.
If name is not specified, a list of all defined sites will be given.
The list will include the numbers associated with each site. When the site is defined, the proportions are stored in variables PERCAn, PERCBn, PERCCn where n is the number associated with each site. The site names may be used in most places where element symbols are used, e.g. in defining the variables ATOMn and Tn. Z values associated with pseudo-element symbols are -n. These may also be used instead of Z values in some instances.
Once the site is defined, the composition can be modified by changing the relevant PERC values. These may also be refined. If PERCAn is changed, PERCBn will be adjusted so as to preserve the previous sum of PERCAn+PERCBn. If a cell STOICHIOMETRY has been defined, using the command of that name, the PERC values for other atoms may be adjusted so as to preserve the stoichiometry.
The fixed stoichiometry procedure is not available in EXCURVE
The central atom of a cluster may be a mixed site. In this way one cluster may be used to fit more than one spectrum, where the central atoms are different.
Sorts all shells into order of increasing R value. Optionally delete empty shells or shells with atoms of a certain type.
Syntax: SORT {DELETE atom_type}
Abbreviation: SO
Notes: The command will always sort shells in order of R and adjust related variables, such as the unit parameter PIV. If DELETE is specified, shells with an occupation number of 0 will be deleted. If an atom type (a number > 0) is also specified, shells of that atom type will also be deleted. The command may be used, for example, to remove H atoms from the structure.
A very useful command for compressing MS path-lists.
Syntax: SPARE
Abbreviation: SP
Notes:
Controls statistical tests.
Syntax: STATS
Abbreviation: ST
Notes: this command is used to test the significance of a shell. To do this it compares the values of fit-index obtained by performing calculations with two different parameter sets. The NS values in these parameter sets must differ by one. The command prompts for both sets of parameters. You can use the current parameters, a set stored using the SAVE command, a set read from a parameter file or a set read from a multiple scattering file. In the last case the data from the file
is also used. A list of any sets of parameters which have been saved is displayed.
After two sets of parameters have been calculated the program will report whether or not the extra shell is significant.
The statistical basis of this test is invalid - use changes in CHI^2 instead. See the manual section on statistics.
Generates shell parameters for simple structures.
Syntax: STRUCTURE structure_type
Abbreviation: STR
Structure_type can be one of: FCC, HCP, BCC, diamond, halite, fluorite, CsCl, sphalerite - these names can be abbreviated to three letters. If a point group is given then it must represent the symmetry of the central atom and not of the molecule as a whole.
In order to generate the shell parameters, R1, T1 and NS need to be defined ( and T2 if the structure needs more than one type of atom ). The other shell parameters will be generated automatically. The point group is set automatically. e.g. to Oh for FCC. Normally a structure is defined only once. If parameters are changed, and the atomic coordinates need to be regenerated the command SYMMETRY is used (SYM OH etc.).
Any subsequent use of STRUCTURE will overwrite the shell coordinates.
Syntax: SYMMETRY [point_group]
Abbreviation: SYM, SY
Notes:
This command uses the shell variables N, T, R, UN, ANG, TH and PHI to calculate the actual atomic positions for a specified point group. The command is essential for full multiple scattering calculation, for XANES calculations, and for the commands DRAW and DISPLAY, all of which require atomic coordinates. The command will not generate or change any of the shell parameters. Once the point groups have been defined for each cluster, the command will be executed automatically whenever updated atomic coordinates are required.
The command can be issued in two ways:
SYMMETRY
Causes a prompt for a point group for each structure. Replying ? to the prompt will display a list of allowed
point groups and structures. Point group names are in the Schoenflies form. INDEX SYMMETRY gives a list of all possible point groups including the equivalent International symbol for each.
SYMMETRY point_group
The usual usage
All of the structural parameters for NS shells must be defined and they must comply with the rules set out below.
a) a shell must contain a complete set of atoms related by the operations of the point group. The maximum number of atoms in a shell is thus equal to the order of the group. If a shell is to contain less atoms they must be at special positions with respect to one of the symmetry operators. Thus, for example, the point group Oh has order 48 but it is possible to have 8 atoms in a shell if they occupy positions on the triad axes ( at the corners of a cube with the origin in the centre ). A table of valid occupation numbers is given below. Further information can be found in, for instance, International Tables for X-Ray
Crystallography, Vol A, starting at p745.
b) if shells are NOT assigned to units then the R, TH and PHI variables for each shell should, in general, be defined before
this command is executed. If the occupation number of a shell is equal to the order of the point group then the given values
of TH and PHI will be used for the position of the first atom and the other positions will be generated with the symmetry
operations. If the occupation number is such that the positions the atoms must occupy are uniquely defined then the given values of TH and PHI will be ignored. If the occupation number means that the atom positions are defined, but not uniquely, then the given values of TH and PHI will be used to make a choice of atom positions. If the values of TH and PHI do not help in the choice of positions then a prompt for more information will be made.
c) if shells ARE assigned to units then only the relative positions of the atoms in the units need to be defined before
this command is executed. If the atoms are all in general positions then a position in theta and phi will be requested
for each unit. If the atoms are special positions but not uniquely defined then further information will be requested.
Note that if one atom in a unit is in a special position then all the atoms in that unit must be in a similar special
position. Requests for rotations expect angles measured from the current position of the unit whereas requests for
coordinates expect positions measured from the origin.
d) The command will always generate atomic coordinates in a standard orientation, with he principal rotation axis parallel to the z axis. When there are two-fold axes normal to the principal axis one of them will be parallel to the x axis. Ideally the shell coordinates should also correspond to this orientation, but the program will attempt to generate the cluster using a number of settings for most point groups (setting 0 corresponds to the standard setting). It is also possible to specify the setting explicitly, using for example:
SYM Cs:1
If the mirror plane is normal to the y rather than the z axis.
For complex structures a trial and error procedure using SYMMETRY, DRAW and DISPLAY is recommended.
( minimum abbreviation TA )
This command is used to display a table of sets of parameter values and to save them in a file for later use. All parameter values are saved using the SAVE command. As well as the parameters normally available to the LIST and CHANGE commands TABLEs may also include the following values:
REX - the exafs part of the R-factor
RD - the distances part of the R-factor
NPT - the number of points
CHI - the chi2 value
Tables may have up to 100 columns and 100 rows
The example below constructs a table which shows the effect of two values of R1 on the value of R(exafs) (prompts from EXCURVE are not shown):
SAVE ( saves current values of everything )
C R1 1.5 ( change R1 to a different value )
SAVE ( save new values of everything )
TABLE ( start table construction )
EXAMPLE.TABLE ( file to save to )
This is an Example Table ( title of table )
First Set ( first column title )
Second Set ( second column title )
=
R1 ( parameter list )
R2
A1
A2
REX
=
Displays the date and time.
Syntax: TIME
Abbreviation: T
Notes: The CPU time and total time are in seconds.
Undo the effect of the last CHANGE
Syntax: UNDO
Abbreviation: UND
Notes:
The parameter list will be restored to its state before the last CHANGE. This will include parameters modified indirectly as a result of the CHANGE. Note that REFINE and some other commands will execute a CHANGE.
For manipulating theoretical contributions calculated in batch jobs.
Syntax: XADD option
Abbreviation: XAD
Notes: Spectra calculated by batch jobs (for example multiple scattering contributions) can be added to the normal theoretical spectrum with this command. The XADD spectrum is read in using XADD READ and stored within the program. If the XADD addition flag is ON then the XADD spectrum is added to all theoretical spectra before they are plotted or written. If the addition flag is OFF then the XADD spectrum is not added.
The following options are available:
READ reads a file into the XADD spectrum. Note that spectra read in must have the same EMIN, EMAX and NPT as the current theory. Also sets the addition flag to ON.
ON sets the XADD addition flag to ON. The XADD spectrum will be added to any theory calculations.
OFF sets the XADD addition flag to OFF.
STATUS shows the current value of the addition flag and contents of the XADD spectrum.
ADD another file is read in and the contents added to the XADD spectrum. EMIN, EMAX and NPT must be compatible again.
MULT multiplies the XADD spectrum by a constant or function. This option prompts for values of CONSTANT, ALPHA and BETA in turn. The XADD spectrum is multiplied by:
CONSTANT * EXP( ALPHA * K**2 ) * EXP( BETA / K )
obviously CONSTANT, ALPHA and BETA can be set to 1.0, 0.0 and 0.0 respectively to avoid using that part of the function.
Perform XANES calculations and display results.
Syntax: XANES option
Abbreviation: XAN
Notes:
The program used for calculation is basically the same as the program described in P.J. Durham, J.B. Pendry and C.H. Hodges, Computer Physics Communications, v29, 1981. It has been modified to allow complex phaseshifts and (for K-edges only) , polarisation dependence.
Options:
LIST (L) Show the current data that would be used for a XANES calculation. This is based on the current structural model for experiment 1 and current phaseshifts..
PLOT (P) Displays output from XANES calculations and if required data produced by the Excurve PLOT command.
If PLOT is followed by ABSOLUTE ( which may be abbreviated to A ) then the actual ordinate values from the file will be used. Otherwise all spectra will be normalised to have Y values in the range 0 to 1.
Filenames and details are requested. A number of plots may be displayed simultaneously. Further details are given below.
CALC (C) Performs a XANES calculation, by default using the current data, as in LIST, or optionally using existing files produced by CALC or WRITE.. The program will prompt for a filename for the dipole matrix elements. In order to use existing files the syntax is:
XANES CALC exanesa1
,for example
WRITE (W) Produces data files which may subsequently be used by XANES CALC. See notes on filenames below.
Further Notes:
Filenames
The XANES command uses files to communicate with the XANES program. These files will have automatically generated names, though a prompt for an alternative first part of the name is given.
The root part of the default filenames will be 'exanesa1' for the first calculation, then 'exanesa2' and so on. If files already exist the first available suffix will be used. Four files are used for each XANES calculation:
exanesa1.con
input file for XANES program containing parameters which control the type of calculation, energy range, number of points etc.
exanesa1.ato
input file for XANES program containing details of the structure and phaseshifts
exanesa1.dat
contains the calculated spectrum both with and without the atomic contribution
The .dat file begins with title data and the input structural data but in eV and Angstroms (the original program used atomic units). This is followed by the calculated spectrum in seven columns:
column 1 energy in eV
column 2 spherically averaged XANES ( i.e. one third of m=0 contribution plus two thirds of m=1 contribution )
column 3 the m = 0 XANES
column 4 the m = +/- 1 XANES
column 5 spherically averaged EXAFS - the contribution of the atomic matrix elements is removed. This may be read into
EXCURVE as an experimental spectrum to be compared with theory calculated by the EXAFS algorithm
column 6 the m = 0 EXAFS
column 7 the m = +/- EXAFS
Note the relationship:
XANES = 2k |m|**2 ( 1 + EXAFS )
in which k is the photoelectron wavevector and m is the atomic dipole matrix element.
exanesa1.log
The log file, which should be checked for error messages.
XANES PARAMETERS
VPI the constant imaginary part of the interatomic potential which is used to represent photoelectron lifetime.
Present for historical reasons - not required with complex phaseshifts
XMIN lowest energy required in the XANES spectrum ( eV ), the default value is the origin of the phaseshifts and matrix
elements
XMAX highest energy required in the XANES spectrum ( eV ), the default value is the lesser of 120eV or the upper limit
of phaseshift energies
XNP number of energy points in spectrum, default 50
XOPT determines how the photoelectron lifetimes are calculated. If XOPT is 0 ( the default ) then the imaginary part of the self energy is used in the same way as the original version of the XANES program. The wavevector and phaseshifts are treated as complex when calculating the scattering tau matrix but real when adding the atomic contributions. If XOPT is 1 then complex phaseshifts and wavevectors will be used throughout. This makes the calculation less susceptible to some problems caused by the central atom phaseshifts but will cause the results to be slightly different from those calculated by other versions of the XANES program.
XLMAX is the maximum l value to be used in XANES calculations. The default value is 3
XLOUT is the maximum l value to be used in the single centre expansions within the XANES calculation. The default
value is 10.
OUTPUT determines how much information about the calculation is displayed in the .log file. The default value of 0 is
recommended. Increasing the value produces increasingly more output.
Example Calculation.
The commands which need to be issued are given in capital letters, notes in lower case program responses in bold.
C ATOM[1-2] SI this changes atom types 1and 2 to silicon
CALC POT Calculate potentials - default responses will suffice
CALC PHASE
Do You Require the Atomic Absorption [no]
Y
Filename : [exmtxa1.dat]
<RETURN>
STR DIAMOND calculates actual positions from the first shell variables and NS assuming Td diamond structure
XANES CALC perform XANES calculation for this structure
Enter Filename for Matrix Elements [current data/exmtxa1]
<return> current information for matrix elements
Filename : [exanesa8]:
<return> exanesa1 as filename root
Enter title:
SILICON title, the calculation will then begin,>
Using the PLOT sub-command
filename ? . Any file containing a spectrum can be entered. If the name is prefixed with “+” then this spectrum will be added to the previous one. “ =” will terminate the PLOT subcommand
Cols ? [12] is for the column combination to be used. The default, 12, will read energy and averaged XANES. 15 will read averaged EXAFS. (See .dat file description above).
Ev, Relative, Hartrees, K [eV]
This selects the scale for the x axis. The options are:
eV plots the spectrum as read, presumed to be eV
Relative plots the spectrum with a scale in eV relative to its edge, assumed to be where the spectrum has its maximum derivative.
Hartrees the input spectrum is assumed to have an energy scale in Hartrees. This is converted to eV before plotting.
K the input spectrum is against eV, the displayed spectrum is against wavevector.
If the option is followed by a number then this will be taken as an x-axis shift ( in eV ) for eV, Relative and Hartrees and as the origin of wavevectors for K. Note that the EXCURVE variables XEn and E0, which have similar functions are ignored. EF is however used in the calculations.
Abs/Der? [a] This selects whether absorbance or the derivative of absorbance ( with respect to the current x axis ) is plotted. The choice here may also be followed by a number. This is taken to be a weighting term. When the ABSOLUTE option has been selected ( see above ) the spectrum will be multiplied by this factor before being displayed. If ABSOLUTE has not been selected then this factor has no effect on the appearance of individual spectra but will act as a weighting factor when spectra are added together.
Enter Add, Delete, Edge, Multiply, Replot, SHift, SMooth Continue, or Quit [Replot + Continue]
The options may be abbreviated to the part in capital letters. Note that only the final version of any particular spectrum will appear in any hard copy graphical output which is produced.
ADD number
number is added to the y-values of the spectrum. Note that spectra are not renormalised after these graphical operations so this option can be used to position spectra whether or not ABSOLUTE was specified.
DELETE
The current spectrum is removed.
EDGE number
The effect is to convolute the spectrum with a fermi function so as to simulate an absorption edge at energy = number. The option also prompts for an effective temperature.
MULTIPLY number
The y-values of the spectrum will be multiplied by number.
REPLOT
This option replots all the spectra which have been entered so far. If the ABSOLUTE option was used then the graph will be rescaled to accommodate all the spectra. If it was not then the original y scale of 0 to 1 will be used even if some of the spectra have been modified with ADD and MULTIPLY.
SHIFT number
The spectrum is shifted along the x-axis by number current x-axis units.
SMOOTH number
The default value of number is 1 x-axis unit. The spectrum is smoothed by convolution with a triangle of base ( 2 * number ). This is roughly equivalent to imposing an experimental resolution of number.
CONTINUE
Specifies that manipulation of this spectrum is complete.
QUIT
Leaves the PLOT subcommand.
Point Group Tables
Operators Valid Occupation Numbers
GROUP Tri Diad Vert Mir Axis General Tri Diad Vert Mir Axis
C1 n n n n 1 1
Cs n n n y 1 2 1
Ci n n n n -2 2
C2 n n n n 2 2 1
C3 n n n n 3 3 1
C4 n n n n 4 4 1
C5 n n n n 5 5 1
C6 n n n n 6 6 1
C7 n n n n 7 7 1
C8 n n n n 8 8 1
D2 n y n n 2 4 2 2
D3 n y n n 3 6 3 2
D4 n y n n 4 8 4 2
D5 n y n n 5 10 5 2
D6 n y n n 6 12 6 2
C2v n n y n 2 4 2 1
C3v n n y n 3 6 3 1
C4v n n y n 4 8 4 1
C5v n n y n 5 10 5 1
C6v n n y n 6 12 6 1
C2h n n n y 2 4 2 2
C3h n n n y 3 6 3 2
C4h n n n y 4 8 4 2
C5h n n n y 5 10 5 2
C6h n n n y 6 12 6 2
D2h n y y y 2 8 2 4 4 2
D3h n y y y 3 12 3 6 6 2
D4h n y y y 4 16 4 8 8 2
D5h n y y y 5 20 5 10 10 2
D6h n y y y 6 24 6 12 12 2
D8h n y y y 8 32 8 16 16 2
D2d n y y n -4 8 4 4 2
D3d n y y n -6 12 6 6 2
D4d n y y n -8 16 8 8 2
D5d n y y n -10 20 10 10 2
D6d n y y n -12 24 12 12 2
S4 n n n n -4 4 2
S6 n n n n -6 6 2
S8 n n n n -8 8 2
T y y n n 2 12 4 6
Th y y y y 2 24 8 12 12 6
Td y n y n -4 24 4 12 6
O y y n n 4 24 8 12 6
Oh y y n y 4 48 8 12 24 24 6
C0v n n n n 1 1
D0h n n n y 1 2
Abbreviations:
Tri set of 4 triad axes associated with cubic point groups.
Diad set of diad axes normal to the principal axis.
Vert set of reflection planes parallel to the principal axis.
Mir a reflection plane normal to the principal axis.
Axis principal axis.
Operators:
y indicates the presence of the operator.
n indicates the absence of the operator.
if the order of the principal axis is -ve, a rotation/reflection ( NOT rotation/inversion ) axis isimplied.
Valid Occupation numbers:
general the maximum occupation number for a shell, associated with a general position.
other occupation numbers are associated with special positions on the relevant symmetry operator.
Parameter list with default values
EQ 0.0000E+00 -0.9900E+02 0.9999E+04 1 9
PPPP 0.0000E+00 -0.9990E+03 0.9900E+02 1 71
QQQQ 0.8000E+00 -0.9990E+03 0.9999E+04 0 0
RRRR 0.3000E+01 0.3000E+01 0.9999E+04 0 0
SSSS 0.3000E+04 -0.9990E+03 0.3000E+04 0 0
LMAX 0.2500E+02 0.2000E+01 0.2500E+02 0 7
DLMAX 0.2500E+02 0.1000E+01 0.2500E+02 0 67
TLMAX 0.1200E+02 0.1000E+01 0.1200E+02 0 67
WP 0.1000E+00 -0.9990E+03 0.9900E+02 0 0
NS 0.1000E+01 0.1000E+01 1.3400E+02 0 71
N0 0.4000E+01 0.0000E+00 0.9990E+03 0 67
T0 0.1000E+01 0.1000E+01 0.9990E+03 0 71
R0 0.2400E+01 -0.1000E+01 0.9990E+03 2 67
A0 0.1000E-01 -0.9900E+02 0.9990E+03 3 66
B0 0.0000E+00 -0.9900E+02 0.9990E+03 9 66
C0 0.0000E+00 -0.9900E+02 0.9990E+03 10 66
D0 0.0000E+00 -0.9900E+02 0.9990E+03 2 66
UN0 0.0000E+00 0.0000E+00 0.2500E+02 0 67
ANG0 0.0000E+00 -0.1800E+03 0.3600E+03 4 67
TH0 0.9000E+02 0.0000E+00 0.7200E+03 4 67
PHI0 0.0000E+00 -0.7200E+03 0.7200E+03 4 67
X0 0.2400E+01 -0.9990E+03 0.9990E+03 2 67
Y0 0.0000E+00 -0.9990E+03 0.9990E+03 2 67
Z0 0.0000E+00 -0.9990E+03 0.9990E+03 2 67
ROT0 0.0000E+00 -0.3600E+03 0.3600E+03 4 67
PIV0 0.0000E+00 0.0000E+00 0.9990E+03 0 0
PLA0 0.0000E+00 0.0000E+00 0.9990E+03 0 0
PLB0 0.0000E+00 0.0000E+00 0.9990E+03 0 0
TORA0 0.0000E+00 -0.3600E+03 0.3600E+03 4 67
TORB0 0.0000E+00 -0.3600E+03 0.3600E+03 4 67
TORC0 0.0000E+00 -0.3600E+03 0.3600E+03 4 67
PAT0 0.0000E+00 -0.3600E+03 0.3600E+03 0 0
PANG0 0.0000E+00 -0.3600E+03 0.3600E+03 4 67
UOC0 0.0000E+00 -0.3600E+03 0.3600E+03 0 67
TWST0 0.0000E+00 -0.3600E+03 0.3600E+03 4 67
TILT0 0.0000E+00 -0.3600E+03 0.3600E+03 4 67
CLUS0 0.1000E+01 -0.9900E+02 0.9900E+02 0 71
LINK0 0.0000E+00 -0.9900E+03 0.9900E+03 0 0
POSX0 0.1000E+01 -0.9900E+02 0.9900E+02 0 0
POSY0 0.1000E+01 -0.9900E+02 0.9900E+02 0 0
POSZ0 0.1000E+01 -0.9900E+02 0.9900E+02 0 0
BI0 0.1000E+01 -0.9900E+02 0.9900E+02 0 0
POST00 1.0000E+00 -0.9999E+04 0.9999E+04 0 2
POST10 0.0000E+00 -0.9999E+04 0.9999E+04 0 2
POST20 0.0000E+00 -0.9999E+04 0.9999E+04 0 2
POST30 0.0000E+00 -0.9999E+04 0.9999E+04 0 2
POST40 0.0000E+00 -0.9999E+04 0.9999E+04 0 2
POST50 0.0000E+00 -0.9999E+04 0.9999E+04 0 2
POST60 0.0000E+00 -0.9999E+04 0.9999E+04 0 2
POST70 0.0000E+00 -0.9999E+04 0.9999E+04 0 2
POST80 0.0000E+00 -0.9999E+04 0.9999E+04 0 2
PRE00 0.0000E+00 -0.9999E+04 0.9999E+04 0 2
PRE10 0.0000E+00 -0.9999E+04 0.9999E+04 0 2
PRE20 0.0000E+00 -0.9999E+04 0.9999E+04 0 2
AFAC0 0.1000E+01 -0.9999E+04 0.9999E+04 0 66
WEX0 1.0000E+00 -0.9999E+04 0.9999E+04 0 2
EF0 0.0000E+00 -0.9999E+04 0.9999E+04 1 79
BETH0 0.0000E+00 -0.9999E+04 0.9999E+04 4 67
BEPHI0 0.0000E+00 -0.9999E+04 0.9999E+04 4 67
BEW0 0.9000E+02 -0.9999E+04 0.9999E+04 4 67
EMIN0 0.3000E+01 0.3000E+01 0.9999E+04 1 79
EMAX0 0.3000E+04 -0.9990E+03 0.3000E+04 1 79
CW0 -0.1000E+01 -0.1000E+33 0.1000E+33 1 16
VPI0 0.0000E+00 -0.9990E+03 0.9900E+02 1 71
MOPG0 0.0000E+00 -0.9990E+03 0.9900E+02 1 71
XE1 0.0000E+00 -0.9900E+02 0.9900E+02 1 71
XV1 0.0000E+00 -0.9900E+02 0.9000E+01 1 0
ATOM1 0.0000E+00 -0.1000E+02 0.1030E+03 0 16
ION1 0.0000E+00 -0.1000E+01 0.1000E+01 0 16
MTR1 0.1000E+00 0.1000E+00 0.1000E+02 2 16
ALF1 0.6667E+00 0.0000E+00 0.2000E+01 0 16
CMAG1 0.0000E+00 -0.9900E+02 0.9900E+02 0 16
SDX1 0.0000E+00 -0.9900E+02 0.9900E+02 0 67
SDY1 0.0000E+00 -0.9900E+02 0.9900E+02 0 67
SDZ1 0.0000E+00 -0.9900E+02 0.9900E+02 0 67
PERCA1 0.0000E+00 -0.9900E+02 0.9900E+02 0 71
PERCB1 0.0000E+00 -0.9900E+02 0.9900E+02 0 71
PERCC1 0.0000E+00 -0.9900E+02 0.9900E+02 0 71
KMIN 0.8874E+00 0.0000E+00 0.9900E+02 5 79
KMAX 0.2806E+02 0.0000E+00 0.9900E+02 5 79
PMIN 0.0000E+00 -0.9999E+04 0.9999E+04 0 0
PMAX 0.1000E+01 0.0000E+00 0.9999E+04 0 0
OMIN 0.1000E+01 0.1000E+01 0.5000E+01 0 1091
OMAX 0.3000E+01 0.1000E+01 0.5000E+01 0 1091
RMIN 0.1000E+00 0.0000E+00 0.9999E+04 2 0
RMAX 0.1000E+02 0.1000E+00 0.9999E+04 2 0
WIND 0.2000E+01 0.1000E+01 0.4000E+01 0 0
SPARE12 0.0000E+00 0.0000E+00 0.1800E+03 0 0
SPARE13 0.0000E+00 0.0000E+00 0.1800E+03 0 0
SPARE14 0.0000E+00 0.0000E+00 0.1800E+03 0 0
SPARE15 0.0000E+00 0.0000E+00 0.3600E+03 0 0
SPARE16 0.0000E+00 0.0000E+00 0.3600E+03 0 0
SPARE17 0.0000E+00 0.0000E+00 0.3600E+03 0 0
SPARE18 0.0000E+00 0.0000E+00 0.3600E+03 0 0
SPARE19 0.0000E+00 0.0000E+00 0.3600E+03 0 0
SPARE20 0.0000E+00 0.0000E+00 0.3600E+03 0 0
DWMIN -0.1000E+01 -0.9900E+02 0.9900E+02 0 0
DWMAX 0.1000E+01 0.0000E+00 0.9900E+02 0 0
XMIN -0.1000E+01 -0.9990E+03 0.9990E+03 0 0
XMAX 0.1000E+01 -0.9990E+03 0.9990E+03 0 0
YMIN -0.1000E+01 -0.9990E+03 0.9990E+03 0 0
YMAX 0.1000E+01 -0.9990E+03 0.9990E+03 0 0
TMIN -0.1000E+04 -0.9990E+03 0.9990E+03 0 0
TMAX 0.1000E+04 -0.9990E+03 0.9990E+03 0 0
D1A 0.1100E+02 0.1100E+02 0.7780E+03 0 0
D1B 0.1100E+02 0.1100E+02 0.7780E+03 0 0
D2A 0.1100E+02 0.1100E+02 0.7780E+03 0 0
D2B 0.1100E+02 0.1100E+02 0.7780E+03 0 0
OUTPUT 0.0000E+00 0.0000E+00 0.1000E+02 0 0
BOND 0.0000E+00 -0.9990E+03 0.9990E+03 2 0
SIZE 0.3000E+00 -0.9990E+03 0.9990E+03 0 0
VX 0.0000E+00 -0.9990E+03 0.9990E+03 2 0
VY 0.0000E+00 -0.9990E+03 0.9990E+03 2 0
VZ 0.0000E+00 -0.9990E+03 0.9990E+03 2 0
M1 0.0000E+00 -0.1000E+31 0.1000E+31 0 0
M2 0.0000E+00 -0.1000E+31 0.1000E+31 0 0
M3 0.0000E+00 -0.1000E+31 0.1000E+31 0 0
M4 0.0000E+00 -0.1000E+31 0.1000E+31 0 0
M5 0.0000E+00 -0.1000E+31 0.1000E+31 0 0
M6 0.0000E+00 -0.1000E+31 0.1000E+31 0 0
M7 0.0000E+00 -0.1000E+31 0.1000E+31 0 0
M8 0.0000E+00 -0.1000E+31 0.1000E+31 0 0
M9 0.0000E+00 -0.1000E+31 0.1000E+31 0 0
M0 0.0000E+00 -0.1000E+31 0.1000E+31 0 0
XEMIN 0.0000E+00 -0.1000E+02 0.9999E+04 1 0
XEMAX 0.1200E+03 0.0000E+00 0.9999E+04 1 0
XNP 0.5000E+02 0.3000E+01 0.2000E+03 0 0
XLMAX 0.3000E+01 0.1000E+01 0.6000E+01 0 0
XLOUT 0.1000E+02 0.1000E+01 0.2400E+02 0 0
XOPT 0.1000E+01 0.0000E+00 0.1000E+01 0 0
MINDIST 0.7500E+00 0.0000E+00 0.2000E+01 2 67
WD 0.0000E+00 0.0000E+00 0.1000E+03 0 0
WA 0.0000E+00 0.0000E+00 0.1000E+01 0 0
R4W 0.0000E+00 0.0000E+00 0.1000E+01 0 0
MINANG -0.1000E+01 -0.3600E+03 0.9999E+04 4 67
PLMIN 0.0000E+00 0.0000E+00 0.9999E+04 2 1091
PLMAX 0.1000E+02 0.0000E+00 0.9999E+04 2 1091
NGRID 0.3490E+03 0.0000E+00 0.9999E+04 0 16
DFAC 0.1000E+03 0.0000E+00 0.9999E+04 0 0
NPARS 0.1000E+01 0.1000E+01 0.9999E+04 0 0
SNOISE 0.0000E+00 -0.9900E+10 0.9900E+10 0 0
MINMAG 0.0000E+00 -0.9900E+10 0.9900E+10 0 67
SPARE9 -0.1000E+01 -0.1000E+33 0.1000E+33 1 16
SPARE8 -0.1000E+01 -0.1000E+33 0.1000E+33 0 0
WNOISE -0.1000E+01 -0.1000E+33 0.1000E+33 0 2
NPEAKS 0.1000E+03 -0.1000E+33 0.1000E+33 0 0
SPARE11 0.9700E+02 -0.1000E+33 0.1000E+33 0 71
ATMAX 0.2000E+01 2.0000E+00 5.0000E+00 0 67
VECA 0.0000E+00 0.0000E+01 0.6400E+02 0 0
VECB 0.0000E+00 -0.5000E+01 0.6400E+02 0 0
E0 0.0000E+00 -0.9990E+06 0.9990E+06 1 0
BFAC 0.1000E+01 -0.9900E+02 0.9999E+04 0 2
NIND 0.0000E+00 -0.9900E+02 0.9999E+04 0 0
V0 0.0000E+00 -0.9900E+02 0.9999E+04 1 16
FE0 0.0000E+00 -0.9900E+02 0.9999E+04 1 16
RHO0 0.0000E+00 -0.9900E+02 0.9999E+04 0 16
ZMIN 0.1000E+01 -0.9999E+30 0.9999E+30 0 0
ZMAX 0.1000E+01 -0.9999E+30 0.9999E+30 0 0
SURREL 0.1000E+01 0.5000E+00 0.9999E+04 0 67
DELTAX 0.0000E+00 -0.9900E+02 0.9999E+04 0 67
DELTAY 0.0000E+00 -0.9900E+02 0.9999E+04 0 67
DELTAZ 0.0000E+00 -0.9900E+02 0.9999E+04 0 67
SURROT 0.0000E+00 -0.9900E+04 0.9999E+04 4 67
ACELL 0.1000E+01 -0.9900E+02 0.9999E+04 2 67
BCELL 0.1000E+01 -0.9900E+02 0.9999E+04 2 67
CCELL 0.1000E+01 -0.9900E+02 0.9999E+04 2 67
ALPHA 0.9000E+02 -0.9900E+02 0.9999E+04 4 67
BETA 0.9000E+02 -0.9900E+02 0.9999E+04 4 67
GAMMA 0.9000E+02 -0.9999E+02 0.9999E+04 4 67
AH 0.9000E+02 -0.9999E+04 0.9999E+04 4 6
BH 0.9000E+02 -0.9999E+04 0.9999E+04 4 6
CH 0.9000E+02 -0.9999E+04 0.9999E+04 4 6
BACK0 0.0000E+00 -0.9999E+04 0.9999E+04 0 2
BACK1 0.0000E+00 -0.9999E+04 0.9999E+04 0 2
BACK2 0.0000E+00 -0.9999E+04 0.9999E+04 0 2
BACK3 0.0000E+00 -0.9999E+04 0.9999E+04 0 2
BACK4 0.0000E+00 -0.9999E+04 0.9999E+04 0 2
BACK5 0.0000E+00 -0.9999E+04 0.9999E+04 0 2
BACK6 0.0000E+00 -0.9999E+04 0.9999E+04 0 2
BACK7 0.0000E+00 -0.9999E+04 0.9999E+04 0 2
BACK8 0.0000E+00 -0.9999E+04 0.9999E+04 0 2
OFFSET 0.1000E-01 -0.9999E+04 0.9999E+04 0 2
CENT 0.0000E+00 -0.9999E+04 0.9999E+04 0 0
NPOST 0.3000E+01 -0.9999E+04 0.9999E+04 0 2
DE0 0.0000E+00 -0.9999E+04 0.9999E+04 1 2
NSPEC 0.1000E+01 0.1000E+01 0.2000E+02 0 71
MAXRAD 0.5000E+01 -0.9900E+02 0.9999E+04 4 0
SPARE1 0.0000E+00 -0.9900E+02 0.9999E+04 0 0
SPARE2 0.0000E+00 -0.9900E+02 0.9999E+04 0 0
SPARE3 0.0000E+00 -0.9900E+02 0.9999E+04 0 0
TEMP 0.2980E+03 -0.9900E+02 0.9999E+04 0 2
CLE 0.0000E+00 -0.9900E+02 0.9999E+04 0 2
NUMAX 0.2000E+01 0.0000E+00 0.2000E+01 0 67
K edges
1 H 13.6 2 HE 24.6 3 LI 54.8 4 BE 111.0 5 B 188.0
6 C 284.0 7 N 402.0 8 O 532.0 9 F 685.0 10 NE 867.0
11 NA 1072.0 12 MG 1305.0 13 AL 1559.0 14 SI 1838.0 15 P 2145.0
16 S 2472.0 17 CL 2822.0 18 AR 3202.0 19 K 3607.0 20 CA 4038.0
21 SC 4493.0 22 TI 4966.0 23 V 5465.0 24 CR 5989.0 25 MN 6539.0
26 FE 7112.0 27 CO 7709.0 28 NI 8333.0 29 CU 8979.0 30 ZN 9659.0
31 GA 10367.0 32 GE 11103.0 33 AS 11867.0 34 SE 12658.0 35 BR 13474.0
36 KR 14325.0 37 RB 15199.0 38 SR 16104.0 39 Y 17038.0 40 ZR 17998.0
41 NB 18986.0 42 MO 20000.0 43 TC 21044.0 44 RU 22117.0 45 RH 23219.0
46 PD 24350.0 47 AG 25514.0 48 CD 26711.0 49 IN 27940.0 50 SN 29200.0
51 SB 30491.0 52 TH 31813.0 53 I 33169.0 54 XE 34561.0 55 CS 35985.0
56 BA 37440.0 57 LA 38925.0 58 CE 40443.0 59 PR 41990.0 60 ND 43569.0
61 PM 45184.0 62 SM 46834.0 63 EU 48519.0 64 GD 50239.0 65 TB 51996.0
66 DY 53789.0 67 HO 55618.0 68 ER 57486.0 69 TM 59390.0 70 YB 61332.0
71 LU 63314.0 72 HF 65351.0 73 TA 67416.0 74 W 69525.0 75 RE 71676.0
76 OS 73870.0 77 IR 76111.0 78 PT 78395.0 79 AU 80724.0 80 HG 83102.0
81 TL 85530.0 82 PB 88004.0 83 BI 90525.0 84 PO 93105.0 85 AT 95730.0
86 RN 98404.0 87 FR 101137.0 88 RA 103921.0 89 AC 106755.0 90 TH 109650.0
91 PA 112601.0 92 U 115606.0 93 NP 118678.0 94 PU 121818.0 95 AM 125027.0
96 CM 128220.0 97 BK 131590.0 98 CF 135960.0 99 ES 39490.0 100 FM 143090.0
101 MD 146780.0 102 NO 150540.0 103 LR 154380.0
L3 edges
1 H .0 2 HE .0 3 LI .0 4 BE .0 5 B .0
6 C .0 7 N .0 8 O .0 9 F 8.6 10 NE 18.3
11 NA 31.1 12 MG 51.4 13 AL 73.1 14 SI 99.2 15 P 132.0
16 S 165.0 17 CL 200.0 18 AR 245.0 19 K 294.0 20 CA 346.0
21 SC 402.0 22 TI 456.0 23 V 513.0 24 CR 575.0 25 MN 640.0
26 FE 708.0 27 CO 779.0 28 NI 855.0 29 CU 931.0 30 ZN 1020.0
31 GA 1115.0 32 GE 1217.0 33 AS 1323.0 34 SE 1436.0 35 BR 1550.0
36 KR 1675.0 37 RB 1804.0 38 SR 1940.0 39 Y 2080.0 40 ZR 2222.0
41 NB 2371.0 42 MO 2520.0 43 TC 2677.0 44 RU 2838.0 45 RH 3004.0
46 PD 3173.0 47 AG 3351.0 48 CD 3538.0 49 IN 3730.0 50 SN 3939.0
51 SB 4132.0 52 THE 4341.0 53 I 4557.0 54 XE 4782.0 55 CS 5011.0
56 BA 5247.0 57 LA 5483.0 58 CE 5723.0 59 PR 5964.0 60 ND 6208.0
61 PM 6459.0 62 SM 6716.0 63 EU 6977.0 64 GD 7243.0 65 TB 7514.0
66 DY 7790.0 67 HO 8071.0 68 ER 8358.0 69 TM 8648.0 70 YB 8944.0
71 LU 9244.0 72 HF 9561.0 73 TA 9881.0 74 W 10206.0 75 RE 10535.0
76 OS 10870.0 77 IR 11215.0 78 PT 11563.0 79 AU 11918.0 80 HG 12284.0
81 TL 12658.0 82 PB 13035.0 83 BI 13419.0 84 PO 13813.0 85 AT 14213.0
86 RN 14619.0 87 FR 15031.0 88 RA 15444.0 89 AC 15871.0 90 TH 16300.0
91 PA 16733.0 92 U 17166.0 93 NP 17610.0 94 PU 18057.0 95 AM 18504.0
96 CM 18930.0 97 BK 19452.0 98 CF 19930.0 99 ES 20410.0 100 FM 20900.0
101 MD 21390.0 102 NO 21880.0 103 LR 22360.0
Ankudinov,A.L.,Zabinsky,S.I. and Rehr,J.J.(1996). Single configuration Dirac-Fock code. Computer Phys.Commun.
Barth,U.von and Grossman,G.(1979). The effect of the core hole on X-ray emission spectra in simple metals. Solid State Commun. 32,645.
Barth,U.von and Grossman,G.(1982). Dynamical effects in X-ray spectra and the final state rule. Phys.Rev.B. 25,5150.
Barth,U.von and Hedin,L.(1972). A local exchange-correlation potential for the spin polarised case: I. J.Phys.C. 5,1629.
Barton,J.J. and Shirley,D.A.(1985). Small-atom approximations for photoelectron scattering in the intermediate-energy range. Phys.Rev.B. 32,1906.
Barton,J.J. and Shirley,D.A.(1985). Curved wave front corrections for photoelectron scattering. Phys.Rev.B. 32,1892.
Beattie,I.R.,Binsted,N.,Levason,W.,Ogden,J.S.,Spicer,M.D.and Young.N.J.(1990). EXAFS, matrix isolation and high temperature chemistry. High Temp Sci. 26,71.
Benfatto,M.,Natoli,C.R.,Bianconi,A.,Garcia,J.,Marcelli,A.,Fanfoni,M. and Davoli,I.(1986). Multiple-scattering regime and higher-order correlations in x-ray absorption spectra of liquid solitions. Phys.Rev.B. 34,5774.
Benfatto,M.,Natoli,C.R.,Brouder,C.,Pettifer,R.F. and Ruiz Lopez,M.F.(1989). Polarized curved-wave extended x-ray absorption fine structure: Theory and application. Phys.Rev.B. 39,1936.
Beni,G. and Platzman,P.M.(1976). Temperature and polarization dependence of extended x-ray absorption fine structure spectra. Phys.Rev.B. 14,1514.
Bethe,H.A. and Salpeter,E.E.(1957). Quantum mechanics of one- and two-electron atoms. Springer-Verlag.
Bianconi,A.(1980). Surface X-ray absorption spectroscopy: surface EXAFS and surface XANES. Appl.of Surface Sci. 6,392.
Biedenharn,L.C. and Loucks,J.D.(1981). Angular momentum in quantum physics. Addison-Wesley.
Binsted,N. and Hasnain,S.S.(1996). State of the art calculations of the total absorption spectrum. J.Synch.Rad. 3,185.
Binsted,N. and Norman,D.(1993). Relaxations and reconstructions derived from SEXAFS spectra of adsorbates on single crystal surfaces. in X.Xie,S.Y.Tong, and van Hove (eds.), the structure of surfaces IV,World Scientific. 118.
Binsted,N. and Norman,D.(1994). Multiple-scattering effects in polarization-dependent surface XAFS. Phys.Rev.B. 49,15531.
Binsted,N. and Norman,D.(1993). Multiple scattering effects in polarisation-dependent surface EXAFS. Jpn.J.Appl.Phys. 32,343.
Binsted,N.,Cook,S.L.,Evans,J.,Greaves,G.N.and Price,R.J.(1987). EXAFS and near-edge structure in the cobalt K-edge absorption spectra of metal carbonyl complexes. J.Am.Chem.Soc 109,3669.
Binsted,N.,Dann,S.E.,Pack,M.J. and Weller,M.T.(1998). The structure of gallobicchulite. A combined EXAFS/powder neutron diffraction refinement. Acta Cryst. B.
Binsted,N.,Norman.D. and Thornton,G.(1995). Analysis of EXAFS spectra of transition-metal sulphides and sulphur on nickel surfaces. Phys.Rev.B 51,7905.
Binsted,N.,Pack,M.J.,Weller,M.T. and Evans,J.(1996). Combined EXAFS and powder diffraction analysis. J.Am.Chem.Soc. 118,10200.
Binsted,N.,Strange,R.W. and Hasnain,S.S.(1992). Constrained and restrained refinement in EXAFS data analysis with curved wave theory. Biochemistry 31,12117.
Binsted,N.,Weller,M.T. and Evans,J.(1995). Combined EXAFS and powder diffraction analysis. Physica B. 208,129.
Blake,R.J.(1993). Relativistic effects in EXAFS and XANES. DL/SCI/P 729.
Bohmer,W. and Rabe,P.(1979). Temperature dependence of the mean square relative displacements of nearest-neighbour atoms derived from EXAFS spectra. J.Phys.C. 12,2465.
Boland,J.J. and Baldeschwieler,J.D.(1984). The effect of thermal vibrations on extended x-ray absorption fine structure.I. J.Chem.Phys. 80,3005.
Boland,J.J. and Baldeschwieler,J.D.(1984). The effect of thermal vibrations on extended x-ray absorption fine structure II. J.Chem.Phys. 81,1145.
Boland,J.J.,Crane,S.E. and Baldeschwieler,J.D.(1982). Theory of extended x-ray absorption fine structure: single and multiple scattering formalisms. J.Chem.Phys. 77,142.
Boland,J.J.,Halaka,F.G. and Baldeschwieler,J.D.(1986). A novel scheme for structure determination in EXAFS. Chem.Phys.Letts. 129,1.
Boland,J.J.,Halaka,G. and Baldeschwieler,J.D.(1983). Data analysis in the extended x-ray-absorption fine structure: determination of the background absorption and the threshold energy. Phys.Rev.B. 28,2921.
Brade,R.M. and Yates,B.(1969). Thermodynamic calculations of the Debye-Waller frequency integrals of alkaline earth fluorides. Phys.Stat.Sol. 1969,551.
Breinig,M,Chen,M.H.,Ice,G.E.,Parente,F and Crasemann,B.(1980). Atomic inner-shell level energies determined by absorption spectrometry with synchrotron radiation. Phys.Rev.A. 22,520.
Brink,D.M. and Satchler,G.R.(1968). Angular momentum. OUP (Oxford).
Brouder,C,Ruiz Lopez,M.F.,Pettifer,R.F.,Benfatto,M. and Natoili,C.R.(1989). Systematic approach to the calculation of the polarization dependent (and polarization averaged) general term of the curved-wave multiple-scattering series in the x-ray-absorption
Case,D.A.(1982). Electronic structure calculations using the Xalpha method. Ann.Rev.Phys.Chem. 33,151.
Chou,S.-H., Rehr,J.J., Stern,E.A. and Davidson,E.R.(1987). Ab initio calculation of extended x-ray absorption fine structure in Br2. Phys.Rev.B. 35,2604.
Clarke,L.J.(1984). Surface crystallography. John Wiley and Sons, Chichester.
Cooper,J.W.(1988). Near-threshold K-shell absorption cross section of argon: relaxation and correlation effects. Phys.Rev.A. 38,3417.
Cromer,D.T.(1983). Calculation of anomalous scattering factors at arbitrary wavelengths. J.Appl.Cryst. 16,437.
Cromer,D.T. and Lieberman,D.A.(1970). Relativistic calculation of anomalous scattering factors for X-rays. J.Chem.Phys. 53,1891.
Cromer,D.T. and Lieberman,D.A.(1981). Anomalous dispersion calculations near to and on the long-wavelength side of an absorption edge. Acta Cryst.A. 37,267.
Dalba,G.,Fornasini,P. and Rocca,F.(1993). Cumulant analysis of the EXAFS of beta-AgI. Phys.Rev.B. 47,8502.
Dalba,G.,Fornasini,P.,Rocca,F. and Mobilio,S.(1990). Correlation effects in the EXAFS Debye-Waller factors of AgI. Phys.Rev.B. 41,9668.
Dehmer,J.L. and Dill,D.(1975). Shape resonances in K-shell photoionization of diatomic molecules. Phys.Rev.Letts. 35,213.
DeWames,R.E.,Wolfram,T. and Lehman,G.W.(1963). Temperature dependence of the Debye-Waller factor for copper and aluminium. Phys.Rev. 131,528.
Di Cicco,A.(1995). EXAFS multiple-scattering data-analysis: GNXAS methodology and applications. Physica B. 208,125.
Dill,D and Dehmer,J.L.(1974). Electron-molecule scattering and molecular photoionisation using the multiple-scattering method. J.Chem.Phys. 61,692.
Doniach,S. and Sondheimer,E.H.(1974). Green's functions for solid state physicists. W.A.Benjamin.
Durham,P.J.,Gyorffy,B.L. and Pindor,A.J.(1980). On the fundamental equations of the Korringa-Kohn-Rostoker (KKR) version of the coherent potential approximation (CPA). J.Phys.F. 10,661.
Durham,P.J.,Pendry,J.B. and Hodges,C.H.(1981). XANES: Determination of bond angles and multi-atom correlations in ordered and disordered systems. Solid State Com.
Durham,P.J.,Pendry,J.B. and Hodges,C.H.(1982). Calculation of X-ray absorption near edge structure, XANES. Comput.Phys.Commun. 25,193.
Dyall,K.G. and Larkins,F.P.(1982). Satellite structure in atomic spectra I. Theoretical framework and application to exited states of singly ionised rare gases. J.Phys.B. 15,203.
Edmonds,A.R.(1960). Angular momentum in quantum physics. Princeton Univ. Press.
Edwards,A.B.,Tildesley,D.J. and Binsted,N.(1997). Cumulant expansion analysis of thermal disorder in FCC copper metal by molecular dynamics simulation. Molecular Physics 91,357.
Fano,U. and Cooper,J.W.(1986). Spectral distribution of atomic oscillator strengths. Rev.Mod.Phys. 40,441.
Filipponi,A.(1994). Radial distribution function probed by x-ray absorption spectroscopy. J.Phys.: Cond.mat.
Filipponi,A.(1995). Double-electron excitation effects above inner shell X-ray absorption edges. Physica B. 208,29.
Filipponi,A.,Bernieri,E.,and Mobilio,S.(1988). Multielectron excitations in X-ray absorption spectra of alpha-Si:H. Phys.Rev.B. 38,3298.
Filipponi,A.,Di Cicco,A.,Sessa,V.,Dossi,C. and Psaro,R.(1991). Multiple-scattering analysis of the X-ray absorption spectrum of Os3(CO)12 carbonyl cluster. Chem.Phys.Letts. 184,485.
Foulis,D.L.,Pettifer,R.F. and Sherwood,P.(1994). Full-potential XANES calculations for HCl using SCF electron densities. sub Elsevier Science.
Foulis,D.L.,Pettifer,R.F. and Sherwood,P.(1995). The removal of the muffin-tin approximation and use of self-consistent-field electron densities for calculating the k-edge XANES of chlorine. Europhysics letts. 29,647.
Foulis,D.L.,Pettifer,R.F.,Natoli,C.R. and Benfatto,M.(1990). Full-potential scattered-wave calculations for molecules and clusters: fundamental tests of the method. Phys.Rev.A. 41,6922.
Fox,L. and Goodwin,E.T.(1949). Some new methods for the numerical integration of ordinary differential equations. Trans.Cam.Phil.Soc. 45,373.
Fox,R and Gurman,S.J.(1981). Theoretical calculations of EXAFS data from specular reflectivity. Phys.Chem.Glasses 22,32.
Frenkel,A.I. and Rehr,J.J.(1993). Thermal expansion and XAFS cumulants. Phys.Rev.B. 48,585.
Frenkel,A.I.,Voronel,A.,Katzir,A.,Newville,M. and Stern,E.A.(1995). Buckled crystalline structure of disordered mixed salts. Physica B.
Freund,J.(1991). On the determination of interatomic potential anharmonicities from EXAFS measurements. Phys.Lett.A. 157,256.
Freund,J.,Ingalls,R. and Crozier,E.D.(1989). Extended x-ray absorption fine structure study of Cu under high pressure. Phys.Rev.B. 39,12537.
Fritsche,V.(1992). Shortcomings of separable representations of the Green function in multiple-scattering theory for photoelectron diffraction and EXAFS. J.Elec.Spec. and Rel.Phen. 58,299.
Fritzsce,V.(1990). A new spherical-wave approximation for photoelectron diffraction, EXAFS and MEED. J.Phys.:Condens.Matter. 2,1413.
Fujikawa,T.(1993). Basic features of the short-range-order multiple scattering XANES theory. J.Phys.Soc.Japan. 62,2155.
Fujikawa,T. and Miyanaga,T.(1993). Quantum statistical approach to Debye-Waller factors in EXAFS, EELS and ARXPS. I. Anharmonic contribution in plane-wave approximation. J.Phys.Soc.Jap. 62,4108.
Fujikawa,T.,Yikegaki,T. and Usami,S.(1993). Theory of EELFS compared with EXAFS for catalysis study. Catalysis Letts. 20,149.
Gupta,O.P.(1984). Phonon frequencies and Debye Temperature of copper. J.Phys.F. 14,2899.
Gurman,S.J.(1988). The small atom approximation in EXAFS and surface EXAFS. J.Phys.C. 21,3699.
Gurman,S.J.(1983). An even simpler method for calculating x-ray absorption parameters. J.Phys.C. 16,2987.
Gurman,S.J.(1980). Notes on the EXAFS programs at Daresbury Laboratory. DL Technical Memorandum DL/SCI/TM 21T.
Gurman,S.J.,Binsted,N. and Ross,I.(1984). A rapid, exact curved-wave theory for EXAFS calculations. J.Phys.C. 17,143.
Gurman,S.J.,Binsted,N. and Ross,I.(1986). A rapid,exact,curved-wave theory for EXAFS calculations; II. The multiple-scattering contributions. J.Phys.C. 19,1845.
Hayes,T.M. and Boyce,J.B.(). Extended X-ray absorption fine structure. Solid State Phys. 37,173.
Hedin,L. and Lundquist,S.(1969). Solid State Physics 23,1.
Hohenberg,P. and Kohn,W.(1964). Inhomogeneous electron gas. Phys.Rev.B. 136,864.
Holland,B.W.,Pendry,J.B.,Pettifer,R.F. and Bordas,J.(1978). Atomic origin of structure in EXAFS experiments. J.Phys.C. 11,633.
Keski-Rahkonen,O and Krause,M.O.(1974). Total and partial atomic-level widths. Atomic Data Nuclear Data Tables 14,139.
Knapp,G.S.,Pan,H.K. and Tranquada,J.M.(1985). EXAFS Einstein frequency and moments of the phonon spectrum: An experimental and theoretical study. Phys.Rev.B. 32,2006.
Koelling,D.D. and Harmon,B.N.(1977). A technique for relativistic spin-polarised calculations. J.Phys.C. 10,3107.
Koningsberger,D.C. and Prins,R.(eds.)(1991). Principles, applications, techniques of EXAFS, SEXAFS and XANES. Wiley New York,.
Kotani,A.(1993). Many body effects in the X-ray absorption edge of rare-earth systems. Jpn.J.Appl.Phys. 32,3.
Krause,M.O.(1979). Atomic radiative and radiationless yields for K and L shells. J.Phys.Chem.Ref.Data. 8,307.
Krause,M.O. and Oliver,J.H.(1979). Natural widths of atomic K and L levels, Kalpha X-ray lines and several KLL auger lines. J.Phys.Chem.Ref.Data. 8,329.
Lee,P.A. and Beni,G.(1977). New method for the calculation of atomic phaseshifts: application to extended x-ray absorption fine structure (EXAFS) in molecules and crystals. Phys.Rev.B. 15,2862.
Lee,P.A. and Pendry,J.B.(1975). Theory of the extended x-ray absorption fine structure. Phys.Rev.B. 11,2795.
Loucks,T.L.(1967). Augmented Plane Wave Method, W.A.Benjamin (NY).
Lytle,F.W., Sayers,D.E. and Stern,E.A.(1989). Report of the International Workshop on Standards and Criteria in X-ray Absorption Spectroscopy. Physica B. 158,701.
Mattheis,L.F.(1964). Energy bands for solid argon. Phys.Rev. 133,184.
Miyanaga,T and Fujikawa,T.(1993). Theory of Debye-Waller factors II.Quantum statistical approach to Debye-Waller factor in EXAFS,EELS and ARXPS II. Application to one-dimensional models. Preprint.
Monti,F.(1996). Characterisation of static disorder by cumulant analysis of EXAFS: an investigation on a two-gaussian distribution. J.Synch.Rad.
Muller,J.E. and Wilkins,J.W.(1984). Band-structure approach to the x-ray spectra of metals. Phys.Rev.B. 29,4331.
Mustre de Leon,J. Yacoby,Y., Stern,E.A. and Rehr,J.J.(1990). Analysis of experimental extended x-ray-absorption fine-structure (EXAFS) data using calculated curved-wave, multiple-scattering EXAFS spectra. Phys.Rev.B. 42,10843.
Mustre de Leon,J. Yacoby,Y., Stern,E.A., Rehr,J.J. and Dell'ariccia,M.(1989). A flexible method for fitting EXAFS spectra - application to structural phase transitions. Physica B. 158,263.
Mustre de Leon,J.,Rehr,J.J.,Zabinsky,S.I. and Albers,R.C.(1991). Ab initio curved-wave x-ray-absorption fine structure. Phys.Rev.B. 44,4146.
Natoli,C.R., in Bianconi,A., Incoccia,L. and Stipcich,S.(eds.).(1983). EXAFS and near edge structure. Springer series in chemical physics. 27,43.
Natoli,C.R.,Benfatto,M.,Brouder,C.,Ruiz-Lopez,M.F. and Foulis,D.L.(1990). Multichannel multiple-scattering theory with general potentials. Phys.Rev.B. 42,1944.
Norman,D.(1986). X-ray absorption spectroscopy (EXAFS and XANES) at surfaces. J.Phys.C. 19,3273.
Pendry,J.B.(1974). Low energy electron diffraction. Academic Press London.
Pickering,I.J.,Sansone,M.,Marsch,J. and George,G.N.(1993). Diffraction anomalous fine structure: a new technique for probing local atomic environment. J.Am.Chem.Soc. 115,6302.
Pickering,I.J.,Sansone,M.,Marsch,J. and George,G.N.(1993). Site-specific X-ray absorption spectroscopy using DIFFRAXAFS. Jpn.J.Appl.Phys. 32-2,206.
Quinn,J.J.(1962). Range of excited electrons in metals. Phys.Rev. 126,1453.
Rehr,J.J.(). Inelastic Effects in X-ray Spectroscopies; Theory vs. Experiment.
Rehr,J.J.(1993). Recent developments in multiple scattering calculations of XAFS and XANES. Jpn.J.Appl.Phys. 32,8.
Rehr,J.J. and Albers,R.C.(1990). Scattering-matrix formulation of curved-wave multiple-scattering theory: application to XAFS. Phys.Rev.B. 41,8139.
Rehr,J.J.,Albers,R.C. and Zabinsky,S.I.(1992). High-order multiple-scattering calculations of X-ray Absorption Fine Structure. Phys.Rev.Letts. 69,3397.
Rehr,J.J.,Booth,C.H.,Bridges,F. and Zabinsky,S.I.(1994). X-ray absorption fine structure in embedded atoms. Phys.Rev.B. 49,12347.
Rehr,J.J.,Mustre de Leon,J.,Zabinsky,S.I. and Albers,R.C.(1991). Theoretical X-ray absorption fine structure standards. J.Am.Chem.Soc. 113,5135.
Rehr,J.J.,Zabinsky,S.I.,Ankudinov,A. and Albers,R.C.(1995). Atomic XAFS and XANES. Physica B. 208,23.
Rennert,P.(1992). Curved-wave corrections to the electron scattering Debye-Waller factor. J.Phys.Condens.Matter. 4,4315.
Rennert,P.(1993). Calculation of the XAFS debye-waller factor with spherical-wave corrections. Jpn.J.Appl.Phys. 32,79.
Sakthivel,A. and Young,R.A.(1991). User's guide to programs DBWS-9006 and DBWS-9006PC.
Schiff,L.I.(1968). Quantum mechanics. McGraw-Hill New York,.
Sevillano,E.,Meuth,H. and Rehr,J.J.(1979). EXAFS Debye-Waller factors. I. Monatomic crystals. Phys.Rev.B. 20,4908.
Sham,L.J. and Kohn,W.(1966). One-particle properties of an inhomogeneous interacting electron gas. Phys.Rev. 145,561.
Singh,N. and Sharma,P.K.(1971). Debye-Waller factors of cubic metals. Phys.Rev.B. 3,1141.
Stohr,J and Bauchspiess,K.R.(1991). Unified picture of near-edge and extended X-ray absorption fine structure in low-Z molecules. Phys.Rev.Lett. 67,3376.
Stohr,J.(1992). NEXAFS spectroscopy. Springer series in surface sciences, Springer-Verlag, Berlin. 25.
Takahashi,M.,Emura,S.,Ito,Y.,Mukoyama,T.,Yoshikado,S. and Omote,K.(1995). The contribution of the shakeup and shakeoff effects to XAFS. Physica B. 208,75.
Thole,B.T. and Van der Laan,G.(1988). Branching ratio in x-ray absorption spectroscopy. Phys.Rev.B. 38,3158.
UKAEA Harwell (1987). Harwell Subroutine Library: a catalogue of subroutines. Harwell Report AERE R 9185 (HMSO)
Vacinova,J.,Hodeau,J.L.,Wolfers,P.,Lauriat,J.P. and ElKaim,E.(1995). Use of anomalous diffraction, DAFS and DANES techniques for site-selective spectroscopy of complex oxides. J.Sync.Rad. 2,236.
Van der Laan,G.(1990). Polaronic satellites in x-ray-absorption spectra. Phys.Rev.B. 41,12366.
Van der Laan,G.(1991). M2,3 absorption spectroscopy of 3d transition-metal compounds,. J.Phys.Condens.Matter. 3,7443.
Van der Laan,G. and Kirkman,I.W.(1992). The 2p absorption spectra of 3d transition metal compounds in tetradehral and octohedral symmetry. J.Phys.Condens.Matter. 4,4189.
Van der Laan,G. and Thole,B.T.(1988). Local probe for spin-orbit interaction. Phys.Rev.Lett. 60,1977.
Vvedensky,D.D.,Saldin,D.K. and Pendry,J.B.(1986). An update of DLXANES, the calculation of X-ray absorption near-edge structure. Comp.Phys.Comm. 40,421.
Wille,L.T.,Durham,P.J. and Sterne,P.B.(1986). X-ray absorption in ionic materials. Journal de Physique,Colloque C87 47,43.
Zare,R.N.(1988). Angular momentum. John Wiley (New York).
Zhang,K.,Stern,E.A. and Rehr,J.J.(1984). Adiabatic versus sudden turn-on of multielectron effects in X-ray absorption. Springer Proc. Phys. 2,18.
Absolute coordinates (definition)
Fourier transform parameters 1 2
Relative coordinates (definition)