- Home
- Users & Science
- Scientific Documentation
- ESRF Highlights
- ESRF Highlights 2013
- Enabling technologies
- Segmenting datasets for radiation damage induced phasing
Segmenting datasets for radiation damage induced phasing
Radiation damage of biological material is a major problem in macromolecular crystallography experiments at synchrotron sources. Ionising radiation produces free radicals that destroy the crystalline order of the sample and damage, on a faster time scale, highly reactive centres. Data collection at cryogenic temperature extends the lifetime of measured samples, but intense and highly-focused beams still produce damage that cannot be ignored. Specific sites that exhibit higher radiation sensitivity can be damaged very quickly, making the interpretation of the electron density map more difficult, and weaken the anomalous signal in a dataset by depleting the electron density of heavier atoms (because of their bigger photoelectric cross-section). Some experiments have shown that specific radiation damage can be used to solve the three dimensional structure of novel macromolecules. These experiments are commonly known as radiation damage induced phasing (RIP) experiments.
 |
|
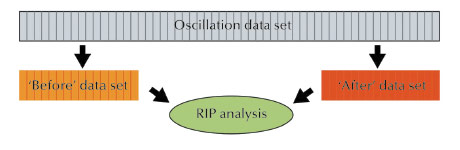
Fig. 147: Schematic diagram of the segmented RIP method. Images from a normally collected oscillation dataset are used to create two subsets, the ‘before’ (orange) and ‘after’ (red) datasets, which are then subjected to RIP analysis (green). |
Differences in intensities between two datasets collected at low X-ray dose, on the same crystal, interleaved by a high dose exposure, permit the location of the positions of the damaged sites that can eventually be used as “fake” heavy atoms to calculate ab initio experimental phases, as in an isomorphous replacement experiment. The damaged sites are usually heavy atoms and/or disulfide bonds [1-4]. Practical applications of this method are limited mainly by the difficulties of choosing the right X-ray dose: too high a dose can destroy the crystalline order and the diffracting power and too low a dose might not induce enough to damage the sites, and make their positions harder to find. Furthermore, the optimal dose varies from protein to protein and not every macromolecule is solvable by these means, as not all molecules present sites that can be used for RIP. In this work, we have further extended the way in which the experiment is conceived, by doing away with the collect-burn-collect protocol, and instead segmenting a large dataset into multiple subsets (Figure 147). The differences between two subsets, the first one beginning at frame 1, and the second one beginning with the last image in the dataset and extending as many images towards the first one, are then used to determine the position of damaged sites and to calculate experimental phases (Figure 148). Different subsets of images are automatically selected to constitute the second dataset, in an attempt to identify the optimal X-ray dose. In contrast to normal radiation damage induced phasing, no optimisation is required for the data collection protocol and no preliminary knowledge of the crystal composition is necessary.
 |
|
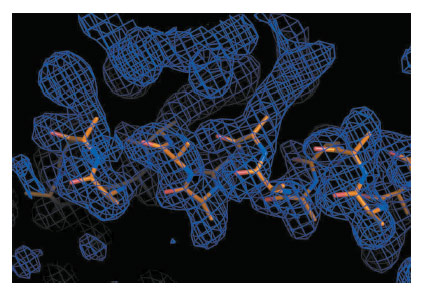
Fig. 148: Example of experimental electron density map calculated after Segmenting RIP, with a partial model built automatically. |
Segmenting RIP is started from the output of GrenADeS autoprocessing [5] and can be triggered automatically for each dataset collected on an ESRF MX beamline. The method is well suited to the trend towards faster detectors that result in datasets containing extremely large numbers of images, collected in a few minutes.
The implementation of Segmenting RIP at the ESRF extends further the concept of software automation that is provided to the user community and represents an excellent example of the synergistic interaction of the development of new tools and automation that takes place at the Structural Biology beamlines.
Principal publication and authors
D. de Sanctis (a) and M.H. Nanao (b,c), Acta Cryst. D68, 1152–1162 (2012).
(a) ESRF
(b) European Molecular Biology Laboratory, Grenoble (France)
(c) Unit of Virus Host–Cell Interactions, UJF-EMBL-CNRS, UMI 3265, Grenoble (France)
References
[1] G. Evans, M. Polentarutti, K. Djinovic Carugo and G. Bricogne, Acta Cryst. D59, 1429–1434 (2003).
[2] M.H. Nanao, G.M. Sheldrick and R.B.G. Ravelli, Acta Cryst. D61, 1227–1237 (2005).
[3] U.A. Ramagopal, Z. Dauter, R. Thirumuruhan, E. Fedorov and S.C. Almo, Acta Cryst. D61, 1289–1298 (2005).
[4] K. Futterer, R.G.B. Ravelli, S.A. White, A.J. Nicoll and R.K. Allemann, Acta Cryst. D64, 264–272 (2008).
[5] S. Monaco, E. Gordon, M.W. Bowler, S. Delagenière, M. Guijarro, D. Spruce, O. Svensson, S.M. McSweeney, A.A. McCarthy, G. Leonard and M.H. Nanao, J Appl Crystallogr. 46, 804–810 (2013).



